3TJN
 
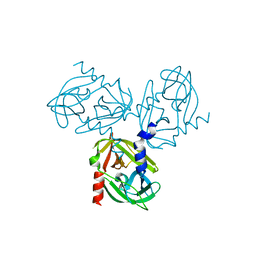 | | HtrA1 catalytic domain, apo form | | Descriptor: | Serine protease HTRA1 | | Authors: | Eigenbrot, C, Ultsch, M. | | Deposit date: | 2011-08-24 | | Release date: | 2012-05-16 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural and Functional Analysis of HtrA1 and Its Subdomains.
Structure, 20, 2012
|
|
3TJO
 
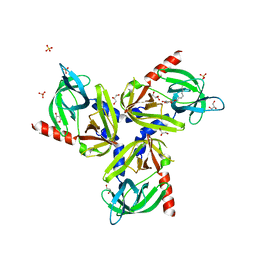 | |
3TJQ
 
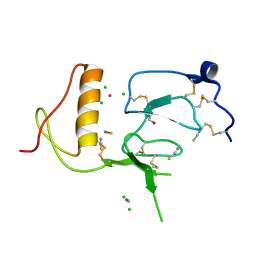 | | N-domain of HtrA1 | | Descriptor: | CHLORIDE ION, GLYCEROL, PLATINUM (II) ION, ... | | Authors: | Eigenbrot, C, Ultsch, M. | | Deposit date: | 2011-08-24 | | Release date: | 2012-05-16 | | Last modified: | 2012-06-27 | | Method: | X-RAY DIFFRACTION (2.001 Å) | | Cite: | Structural and Functional Analysis of HtrA1 and Its Subdomains.
Structure, 20, 2012
|
|
2YQ7
 
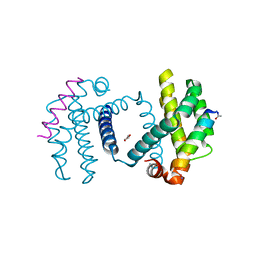 | | Structure of Bcl-xL bound to BimLOCK | | Descriptor: | BCL-2-LIKE PROTEIN 1, BCL-2-LIKE PROTEIN 11, GLYCEROL | | Authors: | Smith, B.J, Czabotar, P.E. | | Deposit date: | 2012-11-06 | | Release date: | 2012-11-28 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.901 Å) | | Cite: | Stabilizing the Pro-Apoptotic Bimbh3 Helix (Bimsahb) Does not Necessarily Enhance Affinity or Biological Activity.
Acs Chem.Biol., 8, 2013
|
|
2YQ6
 
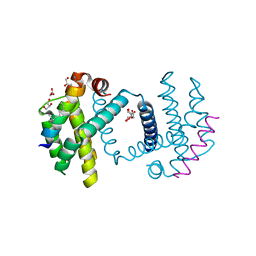 | | Structure of Bcl-xL bound to BimSAHB | | Descriptor: | BCL-2-LIKE PROTEIN 1, BIM BETA 5, GLYCEROL | | Authors: | Smith, B.J, Czabotar, P.E. | | Deposit date: | 2012-11-06 | | Release date: | 2012-11-28 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.799 Å) | | Cite: | Stabilizing the Pro-Apoptotic Bimbh3 Helix (Bimsahb) Does not Necessarily Enhance Affinity or Biological Activity.
Acs Chem.Biol., 8, 2013
|
|
3K48
 
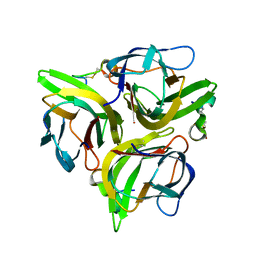 | | Crystal structure of APRIL bound to a peptide | | Descriptor: | Tumor necrosis factor ligand superfamily member 13, peptide | | Authors: | Hymowitz, S.G. | | Deposit date: | 2009-10-05 | | Release date: | 2009-11-24 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Multiple novel classes of APRIL-specific receptor-blocking peptides isolated by phage display.
J.Mol.Biol., 396, 2010
|
|
4O3T
 
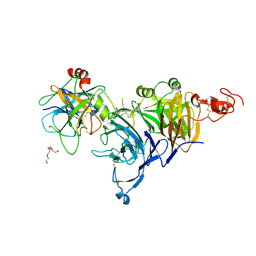 | | Zymogen HGF-beta/MET with Zymogen Activator Peptide ZAP.14 | | Descriptor: | Hepatocyte growth factor, Hepatocyte growth factor receptor, PENTAETHYLENE GLYCOL, ... | | Authors: | Eigenbrot, C, Landgraf, K.E, Steffek, M. | | Deposit date: | 2013-12-18 | | Release date: | 2014-06-04 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.99 Å) | | Cite: | An allosteric switch for pro-HGF/Met signaling using zymogen activator peptides.
Nat.Chem.Biol., 10, 2014
|
|
4O3U
 
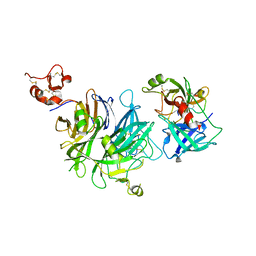 | |
3UW4
 
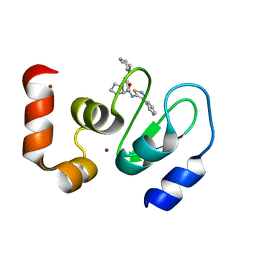 | | Crystal structure of cIAP1 BIR3 bound to GDC0152 | | Descriptor: | Baculoviral IAP repeat-containing protein 2, Baculoviral IAP repeat-containing protein 4, GDC0152, ... | | Authors: | Maurer, B, Hymowitz, S. | | Deposit date: | 2011-11-30 | | Release date: | 2012-02-22 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Discovery of a Potent Small-Molecule Antagonist of Inhibitor of Apoptosis (IAP) Proteins and Clinical Candidate for the Treatment of Cancer (GDC-0152).
J.Med.Chem., 55, 2012
|
|
3UW5
 
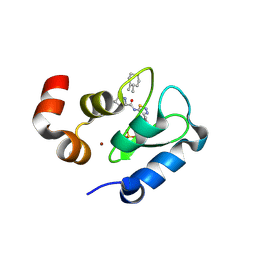 | | Crystal structure of the BIR domain of MLIAP bound to GDC0152 | | Descriptor: | Baculoviral IAP repeat-containing protein 7, Baculoviral IAP repeat-containing protein 4, GDC-0152, ... | | Authors: | Maurer, B, Hymowitz, S.G. | | Deposit date: | 2011-11-30 | | Release date: | 2012-02-22 | | Last modified: | 2017-08-02 | | Method: | X-RAY DIFFRACTION (1.71 Å) | | Cite: | Discovery of a Potent Small-Molecule Antagonist of Inhibitor of Apoptosis (IAP) Proteins and Clinical Candidate for the Treatment of Cancer (GDC-0152).
J.Med.Chem., 55, 2012
|
|
4NMX
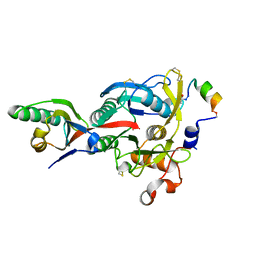 
 | |
5WWU
 
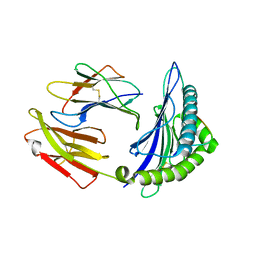 | | Crystal Structure of HLA-A*2402 in complex with 2009 pandemic influenza A(H1N1) virus and avian influenza A(H5N1) virus-derived peptide H1-25 | | Descriptor: | Beta-2-microglobulin, HLA class I histocompatibility antigen, A-24 alpha chain, ... | | Authors: | Zhao, M, Liu, K, Chai, Y, Qi, J, Liu, J, Gao, G.F. | | Deposit date: | 2017-01-05 | | Release date: | 2018-01-17 | | Last modified: | 2019-07-31 | | Method: | X-RAY DIFFRACTION (2.794 Å) | | Cite: | Heterosubtypic Protections against Human-Infecting Avian Influenza Viruses Correlate to Biased Cross-T-Cell Responses.
Mbio, 9, 2018
|
|
5WXD
 
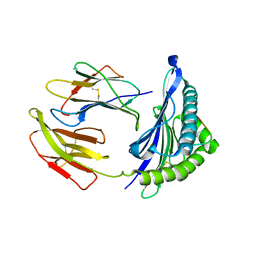 | | Crystal Structure of HLA-A*2402 in complex with avian influenza A(H7N9) virus-derived peptide H7-25 (data set 1) | | Descriptor: | Beta-2-microglobulin, H7-25, HLA class I histocompatibility antigen, ... | | Authors: | Zhao, M, Liu, K, Chai, Y, Qi, J, Liu, J, Gao, G.F. | | Deposit date: | 2017-01-07 | | Release date: | 2018-01-17 | | Last modified: | 2019-07-31 | | Method: | X-RAY DIFFRACTION (3.295 Å) | | Cite: | Heterosubtypic Protections against Human-Infecting Avian Influenza Viruses Correlate to Biased Cross-T-Cell Responses.
Mbio, 9, 2018
|
|
5B7Q
 
 | |
5B7N
 
 | |
5WXC
 
 | | Crystal Structure of HLA-A*2402 in complex with avian influenza A(H7N9) virus-derived peptide H7-25 (data set 2) | | Descriptor: | Beta-2-microglobulin, H7-25-F, HLA class I histocompatibility antigen, ... | | Authors: | Zhao, M, Liu, K, Chai, Y, Qi, J, Liu, J, Gao, G.F. | | Deposit date: | 2017-01-07 | | Release date: | 2018-01-17 | | Last modified: | 2019-07-31 | | Method: | X-RAY DIFFRACTION (2.295 Å) | | Cite: | Heterosubtypic Protections against Human-Infecting Avian Influenza Viruses Correlate to Biased Cross-T-Cell Responses.
Mbio, 9, 2018
|
|
5Y9Q
 
 | |
5B7P
 
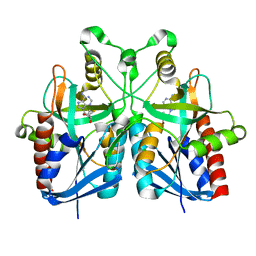 | |
5B7G
 
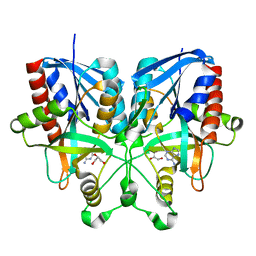 | |
1GNB
 
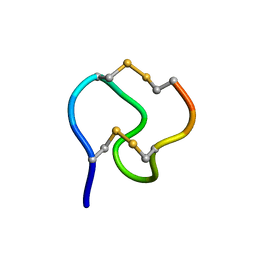 | |
1GNA
 
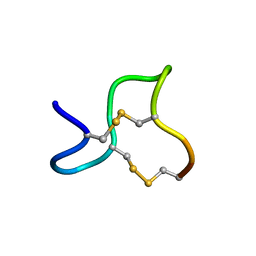 | |
6K28
 
 | | Crystal structure of the 5-(Hydroxyethyl)-methylthiazole Kinase ThiM from Klebsiella pneumonia in complex with TZE | | Descriptor: | 2-(4-METHYL-THIAZOL-5-YL)-ETHANOL, Hydroxyethylthiazole kinase, MAGNESIUM ION | | Authors: | Chen, Y, Wang, L, Shang, F, Lan, J, Liu, W, Xu, Y. | | Deposit date: | 2019-05-13 | | Release date: | 2019-07-10 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.002 Å) | | Cite: | Structural insight of the 5-(Hydroxyethyl)-methylthiazole kinase ThiM involving vitamin B1 biosynthetic pathway from the Klebsiella pneumoniae.
Biochem.Biophys.Res.Commun., 518, 2019
|
|
6JYY
 
 | | Crystal structure of the 5-(Hydroxyethyl)-methylthiazole Kinase ThiM from Klebsiella pneumonia | | Descriptor: | Hydroxyethylthiazole kinase | | Authors: | Chen, Y, Wang, L, Shang, F, Lan, J, Liu, W, Xu, Y. | | Deposit date: | 2019-04-29 | | Release date: | 2019-06-26 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural insight of the 5-(Hydroxyethyl)-methylthiazole kinase ThiM involving vitamin B1 biosynthetic pathway from the Klebsiella pneumoniae.
Biochem.Biophys.Res.Commun., 518, 2019
|
|
6K63
 
 | | The crystal structure of cytidine deaminase from Klebsiella pneumoniae | | Descriptor: | 1,4-DIETHYLENE DIOXIDE, Cytidine deaminase, ZINC ION | | Authors: | Liu, W, Shang, F, Lan, J, Chen, Y, Wang, L, Xu, Y. | | Deposit date: | 2019-06-01 | | Release date: | 2019-07-31 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.073 Å) | | Cite: | Biochemical and structural analysis of the Klebsiella pneumoniae cytidine deaminase CDA.
Biochem.Biophys.Res.Commun., 519, 2019
|
|
6K2L
 
 | | Crystal structure of the Siderophore-interacting protein SipS from Aeromonas hydrophila | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, Siderophore-interacting protein | | Authors: | Shang, F, Lan, J, Liu, W, Xu, Y. | | Deposit date: | 2019-05-14 | | Release date: | 2019-06-12 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of the Siderophore-interacting protein SIP from Aeromonas hydrophila.
Biochem.Biophys.Res.Commun., 519, 2019
|
|
