8TT7
 
 | |
8PNV
 
 | | Cryo-EM structure of styrene oxide isomerase | | Descriptor: | Nanobody, PROTOPORPHYRIN IX CONTAINING FE, Styrene oxide isomerase | | Authors: | Khanppnavar, B, Korkhov, B, Li, X. | | Deposit date: | 2023-07-02 | | Release date: | 2024-04-03 | | Last modified: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (2.048 Å) | | Cite: | Structural basis of the Meinwald rearrangement catalysed by styrene oxide isomerase.
Nat.Chem., 2024
|
|
8PNU
 
 | |
1VDF
 
 | |
4TO7
 
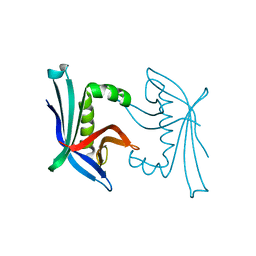 | |
5IIR
 
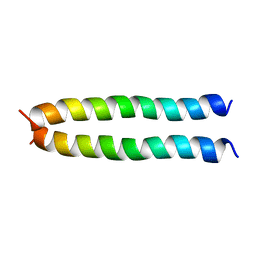 | |
5IIV
 
 | | GCN4p pH 1.5 | | Descriptor: | General control protein GCN4 | | Authors: | Brady, M.R, Kaplan, A.R, Alexandrescu, A.T. | | Deposit date: | 2016-03-01 | | Release date: | 2017-03-15 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Nuclear Magnetic Resonance Structures of GCN4p Are Largely Conserved When Ion Pairs Are Disrupted at Acidic pH but Show a Relaxation of the Coiled Coil Superhelix.
Biochemistry, 56, 2017
|
|
5IEW
 
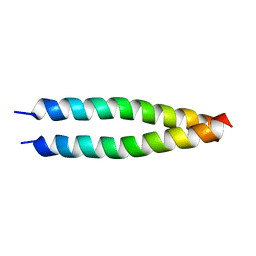 | |
5ITZ
 
 | |
5MOY
 
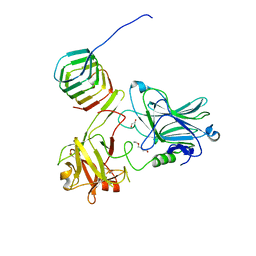 | |
5N74
 
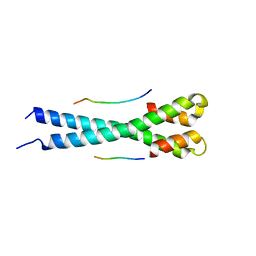 | | Microtubule end binding protein complex | | Descriptor: | Karyogamy protein KAR9, Microtubule-associated protein RP/EB family member 1 | | Authors: | Kumar, A, Steinmetz, M. | | Deposit date: | 2017-02-18 | | Release date: | 2017-06-14 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Short Linear Sequence Motif LxxPTPh Targets Diverse Proteins to Growing Microtubule Ends.
Structure, 25, 2017
|
|
5O5K
 
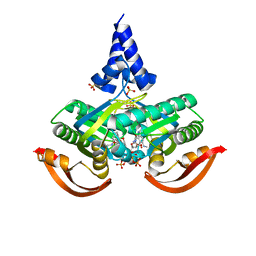 | | X-ray structure of a bacterial adenylyl cyclase soluble domain | | Descriptor: | 3'-O-(N-METHYLANTHRANILOYL)-GUANOSINE-5'-TRIPHOSPHATE, Adenylate cyclase, MANGANESE (II) ION, ... | | Authors: | Vercellino, I, Korkhov, V.M. | | Deposit date: | 2017-06-02 | | Release date: | 2017-11-01 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | Role of the nucleotidyl cyclase helical domain in catalytically active dimer formation.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
2O7H
 
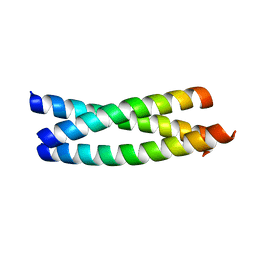 | |
5O5L
 | |
1JC7
 
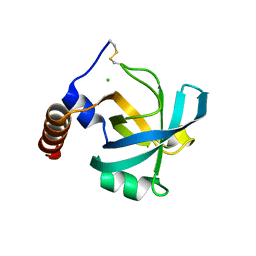 | |
2OVN
 | |
1JB3
 
 | |
4TQ7
 
 | |
4TTZ
 
 | |
4TTW
 
 | |
4TTY
 
 | |
4TPZ
 
 | |
4TTX
 
 | |
4U2I
 
 | |
4U2J
 
 | |
