1ERN
 | |
1EBA
 
 | |
1EBP
 
 | |
2AVI
 
 | |
6R6E
 
 | |
6R3D
 
 | |
6R6X
 
 | |
6R8J
 
 | |
4S2Z
 
 | |
4S34
 
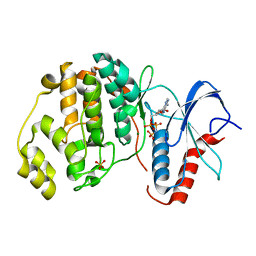 | | ERK2 (I84A) in complex with AMP-PNP | | Descriptor: | MAGNESIUM ION, Mitogen-activated protein kinase 1, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, ... | | Authors: | Livnah, O, Karamansha, Y, Tzarum, N. | | Deposit date: | 2015-01-26 | | Release date: | 2016-01-27 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Multiple mechanisms render Erk proteins MEK-independent
To be Published
|
|
4S32
 
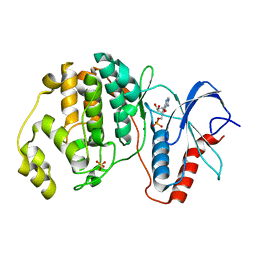 | |
4S30
 
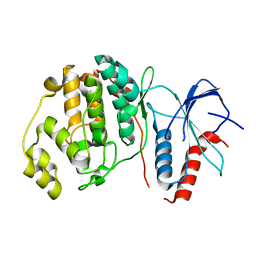 | |
4S31
 
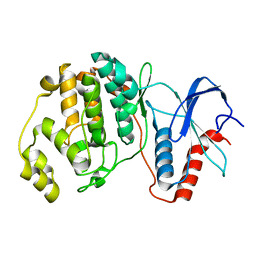 | |
4S33
 
 | | ERK2 R65S mutant complexed with AMP-PNP | | Descriptor: | MAGNESIUM ION, Mitogen-activated protein kinase 1, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, ... | | Authors: | Livnah, O, Karamansha, Y, Tzarum, N. | | Deposit date: | 2015-01-26 | | Release date: | 2016-01-27 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Multiple mechanisms render Erk proteins MEK-independent
To be Published
|
|
7OA0
 
 | | Crystal structure of di-phosphorylated human CLK1 in complex with 5-(6,7-dichloro-1H-indol-3-yl)pyrimidin-4-amine | | Descriptor: | 5-[6,7-bis(chloranyl)-1-methyl-indol-3-yl]pyrimidin-4-amine, Dual specificity protein kinase CLK1 | | Authors: | Livnah, O, Domovich, Y, Bracher, F, Aiger, C. | | Deposit date: | 2021-04-18 | | Release date: | 2022-05-04 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Development of novel CLK1 inhibitors as potential drugs for treatment of Chikungunya virus infections
To Be Published
|
|
7O9Y
 
 | |
4GEO
 
 | | P38a MAP kinase DEF-pocket penta mutant (M194A, L195A, H228A, I229A, Y258A) | | Descriptor: | Mitogen-activated protein kinase 14, octyl beta-D-glucopyranoside | | Authors: | Livnah, O, Tzarum, N. | | Deposit date: | 2012-08-02 | | Release date: | 2013-05-22 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.66 Å) | | Cite: | P38a MAP kinase DEF-pocket penta mutant (M194A, L195A, H228A, I229A, Y258A)
To be Published
|
|
4GGZ
 | | The structure of bradavidin2-biotin complex | | Descriptor: | BIOTIN, Bradavidin 2 | | Authors: | Livnah, O, Meir, A. | | Deposit date: | 2012-08-07 | | Release date: | 2013-06-19 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | The highly dynamic oligomeric structure of bradavidin II is unique among avidin proteins.
Protein Sci., 22, 2013
|
|
4GGR
 
 | | The structure of apo bradavidin2 (Form A) | | Descriptor: | Bradavidin 2 | | Authors: | Livnah, O, Meir, A. | | Deposit date: | 2012-08-07 | | Release date: | 2013-06-19 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The highly dynamic oligomeric structure of bradavidin II is unique among avidin proteins.
Protein Sci., 22, 2013
|
|
4GGT
 
 | | Structure of apo Bradavidin2 (Form B) | | Descriptor: | Bradavidin 2 | | Authors: | Livnah, O, Meir, A. | | Deposit date: | 2012-08-07 | | Release date: | 2013-06-19 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.693 Å) | | Cite: | The highly dynamic oligomeric structure of bradavidin II is unique among avidin proteins.
Protein Sci., 22, 2013
|
|
6RFO
 
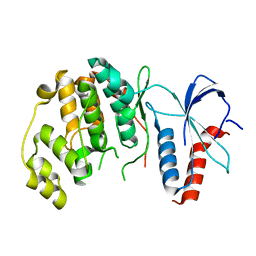 | | ERK2 MAP kinase with the activation loop of p38alpha | | Descriptor: | Mitogen-activated protein kinase 1,Mitogen-activated protein kinase 14,Mitogen-activated protein kinase 1 | | Authors: | Livnah, O, Eitan-Wexler, M. | | Deposit date: | 2019-04-15 | | Release date: | 2020-05-27 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The bacterial metalloprotease NleD selectively cleaves mitogen-activated protein kinases that have high flexibility in their activation loop.
J.Biol.Chem., 295, 2020
|
|
6HDV
 
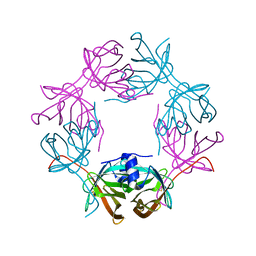 | |
6HDT
 
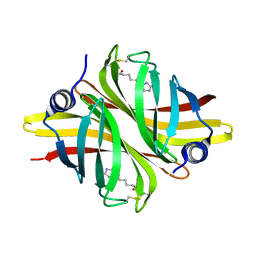 | |
6HDS
 | | Crystal Structure of apo short afifavidin | | Descriptor: | short afifavidin | | Authors: | Livnah, O, Avraham, O. | | Deposit date: | 2018-08-19 | | Release date: | 2018-11-14 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | Crystal structure of afifavidin reveals common features of molecular assemblage in the bacterial dimeric avidins.
FEBS J., 285, 2018
|
|
6RTQ
 
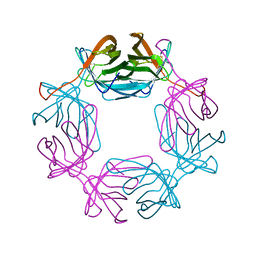 | |
