7M60
 
 | | Structure of Mouse Importin alpha MLH1-S467A NLS Peptide Complex | | Descriptor: | DNA mismatch repair protein Mlh1 NLS peptide, Importin subunit alpha-1 | | Authors: | De Oliveira, H.C, Da Silva, T.D, Fukuda, C.A, Fontes, M.R.M. | | Deposit date: | 2021-03-25 | | Release date: | 2021-07-07 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural and calorimetric studies reveal specific determinants for the binding of a high-affinity NLS to mammalian importin-alpha.
Biochem.J., 478, 2021
|
|
7TMX
 
 | |
7TMY
 
 | |
6WBA
 
 | | Structure of Mouse Importin alpha MLH1- R470A NLS Peptide Complex | | Descriptor: | DNA mismatch repair protein Mlh1, Importin subunit alpha-1 | | Authors: | de Barros, A.C, da Silva, T.D, Oliveira, H.C, Fukuda, C.A, Fontes, M.R.M. | | Deposit date: | 2020-03-26 | | Release date: | 2021-07-07 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.151 Å) | | Cite: | Structural and calorimetric studies reveal specific determinants for the binding of a high-affinity NLS to mammalian importin-alpha.
Biochem.J., 478, 2021
|
|
6WBB
 
 | | Structure of Mouse Importin alpha - MLH1-E475A NLS peptide complex | | Descriptor: | DNA mismatch repair protein Mlh1, Importin subunit alpha-1 | | Authors: | De Barros, A.C, Da Silva, T.D, Oliveira, H.C, Fukuda, C.A, Fontes, M.R.M. | | Deposit date: | 2020-03-26 | | Release date: | 2021-07-07 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.663 Å) | | Cite: | Structural and calorimetric studies reveal specific determinants for the binding of a high-affinity NLS to mammalian importin-alpha.
Biochem.J., 478, 2021
|
|
6WBC
 
 | | Structure of Mouse Importin alpha- MLH1-R472K NLS Peptide Complex | | Descriptor: | DNA mismatch repair protein Mlh1, Importin subunit alpha-1 | | Authors: | de Barros, A.C, da Silva, T.D, Oliveira, H.C, Fukuda, C.A, Fontes, M.R.M. | | Deposit date: | 2020-03-26 | | Release date: | 2021-07-07 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Structural and calorimetric studies reveal specific determinants for the binding of a high-affinity NLS to mammalian importin-alpha.
Biochem.J., 478, 2021
|
|
6P6E
 
 | | Structure of Mouse Importin alpha - PAC3 NLS peptide complex | | Descriptor: | Importin subunit alpha-1, PAC3 NLS | | Authors: | Bernardes, N.E, Silva, T.D, Fukuda, C.A, Oliveira, H.C, Fontes, M.R.M. | | Deposit date: | 2019-06-03 | | Release date: | 2020-01-29 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Comparative study of the interactions between fungal transcription factor nuclear localization sequences with mammalian and fungal importin-alpha.
Sci Rep, 10, 2020
|
|
6P6A
 
 | | Structure of Mouse Importin alpha - NIT2 NLS peptide complex | | Descriptor: | Importin subunit alpha-1, Nitrogen catabolic enzyme regulatory protein | | Authors: | Bernardes, N.E, Fukuda, C.A, Silva, T.D, Oliveira, H.C, Fontes, M.R.M. | | Deposit date: | 2019-06-03 | | Release date: | 2020-01-29 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.151 Å) | | Cite: | Comparative study of the interactions between fungal transcription factor nuclear localization sequences with mammalian and fungal importin-alpha.
Sci Rep, 10, 2020
|
|
6EOA
 
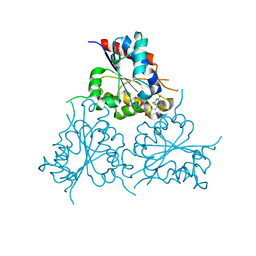 | | Crystal Structure of HAL3 from Cryptococcus neoformans | | Descriptor: | FLAVIN MONONUCLEOTIDE, Phosphopantothenoylcysteine decarboxylase | | Authors: | Reverter, D, Zhang, C, Molero, C, Arino, J. | | Deposit date: | 2017-10-09 | | Release date: | 2018-10-31 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.18 Å) | | Cite: | Characterization of the atypical Ppz/Hal3 phosphatase system from the pathogenic fungus Cryptococcus neoformans.
Mol.Microbiol., 111, 2019
|
|
5CAB
 
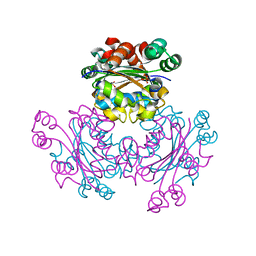 | | Structure of Leishmania nucleoside diphostate kinase mutant Del5-Cterm | | Descriptor: | Nucleoside diphosphate kinase, SULFATE ION | | Authors: | Vieira, P.S, de Giuseppe, P.O, de Oliveira, A.H.C, Murakami, M.T. | | Deposit date: | 2015-06-29 | | Release date: | 2015-10-14 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.953 Å) | | Cite: | The role of the C-terminus and Kpn loop in the quaternary structure stability of nucleoside diphosphate kinase from Leishmania parasites.
J.Struct.Biol., 192, 2015
|
|
5C7P
 
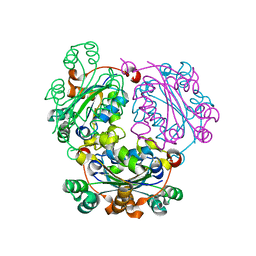 | |
5CAA
 
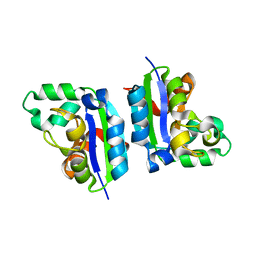 | |
4KPC
 
 | | Crystal structure of the nucleoside diphosphate kinase b from Leishmania braziliensis | | Descriptor: | Nucleoside diphosphate kinase b, PHOSPHATE ION | | Authors: | Vieira, P.S, Giuseppe, P.O, Santos, C.R, Cunha, E.M.F, de Oliveira, A.H.C, Murakami, M.T. | | Deposit date: | 2013-05-13 | | Release date: | 2014-07-09 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure and biophysical characterization of the nucleoside diphosphate kinase from Leishmania braziliensis.
Bmc Struct.Biol., 15, 2015
|
|
