4UU4
 
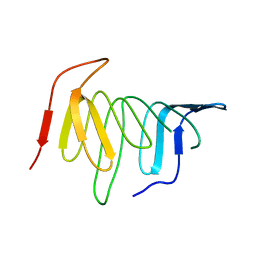 | | Crystal structure of LptH, the LptA homologous periplasmic component of the conserved lipopolysaccharide transport device from Pseudomonas aeruginosa | | Descriptor: | PERIPLASMIC LIPOPOLYSACCHARIDE TRANSPORT PROTEIN LPTH | | Authors: | Bollati, M, Villa, R, Gourlay, L.J, Barbiroli, A, Deho, G, Benedet, M, Polissi, A, Martorana, A, Sperandeo, P, Bolognesi, M, Nardini, M. | | Deposit date: | 2014-07-24 | | Release date: | 2015-03-25 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.751 Å) | | Cite: | Crystal Structure of Lpth, the Periplasmic Component of the Lipopolysaccharide Transport Machinery from Pseudomonas Aeruginosa.
FEBS J., 282, 2015
|
|
7P2B
 
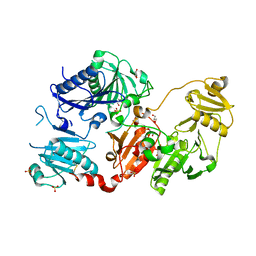 | |
3V5B
 
 | | Structure of Coil 2b of human lamin | | Descriptor: | Prelamin-A/C | | Authors: | Bollati, M, Bolognesi, M. | | Deposit date: | 2011-12-16 | | Release date: | 2012-02-22 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structures of the lamin A/C R335W and E347K mutants: Implications for dilated cardiolaminopathies.
Biochem.Biophys.Res.Commun., 418, 2012
|
|
3V4Q
 
 | | Structure of R335W mutant of human Lamin | | Descriptor: | Prelamin-A/C | | Authors: | Bollati, M, Bolognesi, M. | | Deposit date: | 2011-12-15 | | Release date: | 2012-02-29 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3.06 Å) | | Cite: | Structures of the lamin A/C R335W and E347K mutants: Implications for dilated cardiolaminopathies.
Biochem.Biophys.Res.Commun., 418, 2012
|
|
3V4W
 
 | | Structure of E347K mutant of Lamin | | Descriptor: | Prelamin-A/C | | Authors: | Bollati, M, Bolognesi, M. | | Deposit date: | 2011-12-15 | | Release date: | 2012-02-22 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3.7 Å) | | Cite: | Structures of the lamin A/C R335W and E347K mutants: Implications for dilated cardiolaminopathies.
Biochem.Biophys.Res.Commun., 418, 2012
|
|
3ELU
 
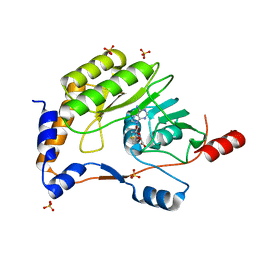 | | Wesselsbron virus Methyltransferase in complex with AdoMet | | Descriptor: | Methyltransferase, S-ADENOSYLMETHIONINE, SULFATE ION | | Authors: | Bollati, M, Ricagno, S, Milani, M, Mastrangelo, E, Bolognesi, M. | | Deposit date: | 2008-09-23 | | Release date: | 2008-11-18 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Recognition of RNA Cap in the Wesselsbron Virus NS5 Methyltransferase Domain: Implications for RNA-Capping Mechanisms in Flavivirus
J.Mol.Biol., 385, 2009
|
|
3EMD
 
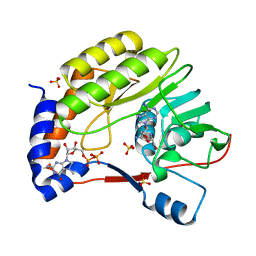 | | Wesselsbron virus Methyltransferase in complex with Sinefungin and 7MeGpppA | | Descriptor: | P1-7-METHYLGUANOSINE-P3-ADENOSINE-5',5'-TRIPHOSPHATE, SINEFUNGIN, SULFATE ION, ... | | Authors: | Bollati, M, Milani, M, Ricagno, S, Mastrangelo, E, Bolognesi, M. | | Deposit date: | 2008-09-24 | | Release date: | 2008-11-18 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Recognition of RNA Cap in the Wesselsbron Virus NS5 Methyltransferase Domain: Implications for RNA-Capping Mechanisms in Flavivirus
J.Mol.Biol., 385, 2009
|
|
3EMB
 
 | | Wesselsbron virus Methyltransferase in complex with AdoMet and 7MeGpppG | | Descriptor: | 7-METHYL-GUANOSINE-5'-TRIPHOSPHATE-5'-GUANOSINE, Methyltransferase, S-ADENOSYLMETHIONINE, ... | | Authors: | Bollati, M, Ricagno, S, Milani, M, Mastrangelo, E, Bolognesi, M. | | Deposit date: | 2008-09-24 | | Release date: | 2008-11-18 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Recognition of RNA Cap in the Wesselsbron Virus NS5 Methyltransferase Domain: Implications for RNA-Capping Mechanisms in Flavivirus
J.Mol.Biol., 385, 2009
|
|
3ELY
 
 | | Wesselsbron virus Methyltransferase in complex with AdoHcy | | Descriptor: | DI(HYDROXYETHYL)ETHER, Methyltransferase, S-ADENOSYL-L-HOMOCYSTEINE, ... | | Authors: | Bollati, M, Milani, M, Ricagno, S, Mastrangelo, E, Bolognesi, M. | | Deposit date: | 2008-09-23 | | Release date: | 2008-11-18 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Recognition of RNA Cap in the Wesselsbron Virus NS5 Methyltransferase Domain: Implications for RNA-Capping Mechanisms in Flavivirus
J.Mol.Biol., 385, 2009
|
|
3ELW
 
 | | Wesselsbron virus Methyltransferase in complex with AdoMet and GpppG | | Descriptor: | DIGUANOSINE-5'-TRIPHOSPHATE, Methyltransferase, S-ADENOSYLMETHIONINE, ... | | Authors: | Bollati, M, Ricagno, S, Milani, M, Mastrangelo, E, Bolognesi, M. | | Deposit date: | 2008-09-23 | | Release date: | 2008-11-18 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Recognition of RNA Cap in the Wesselsbron Virus NS5 Methyltransferase Domain: Implications for RNA-Capping Mechanisms in Flavivirus
J.Mol.Biol., 385, 2009
|
|
3GCZ
 
 | | Yokose virus Methyltransferase in complex with AdoMet | | Descriptor: | GLYCEROL, Polyprotein, S-ADENOSYLMETHIONINE, ... | | Authors: | Bollati, M, Milani, M, Mastrangelo, E, Bolognesi, M, Structural Proteomics in Europe (SPINE) | | Deposit date: | 2009-02-23 | | Release date: | 2009-03-24 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal structure of a methyltransferase from a no-known-vector Flavivirus
Biochem.Biophys.Res.Commun., 382, 2009
|
|
3ELD
 
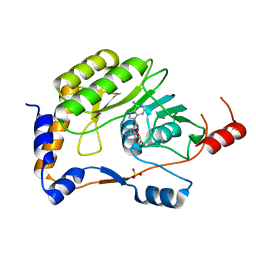 | | Wesselsbron methyltransferase in complex with Sinefungin | | Descriptor: | Methyltransferase, SINEFUNGIN, SULFATE ION | | Authors: | Bollati, M, Milani, M, Mastrangelo, E, Ricagno, S, Bolognesi, M. | | Deposit date: | 2008-09-22 | | Release date: | 2008-11-18 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Recognition of RNA Cap in the Wesselsbron Virus NS5 Methyltransferase Domain: Implications for RNA-Capping Mechanisms in Flavivirus
J.Mol.Biol., 385, 2009
|
|
2OXT
 
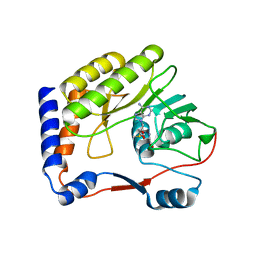 | | Crystal structure of Meaban virus nucleoside-2'-O-methyltransferase | | Descriptor: | NUCLEOSIDE-2'-O-METHYLTRANSFERASE, S-ADENOSYLMETHIONINE | | Authors: | Mastrangelo, E, Milani, M, Bollati, M, Bolognesi, M. | | Deposit date: | 2007-02-21 | | Release date: | 2007-05-15 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structural bases for substrate recognition and activity in Meaban virus nucleoside-2'-O-methyltransferase
Protein Sci., 16, 2007
|
|
2W2G
 
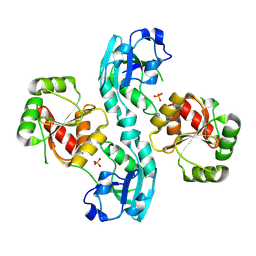 | | Human SARS coronavirus unique domain | | Descriptor: | NON-STRUCTURAL PROTEIN 3, SULFATE ION | | Authors: | Tan, J, Vonrhein, C, Smart, O.S, Bricogne, G, Bollati, M, Kusov, Y, Hansen, G, Mesters, J.R, Schmidt, C.L, Hilgenfeld, R. | | Deposit date: | 2008-10-30 | | Release date: | 2009-05-26 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.22 Å) | | Cite: | The Sars-Unique Domain (Sud) of Sars Coronavirus Contains Two Macrodomains that Bind G-Quadruplexes.
Plos Pathog., 5, 2009
|
|
2WCT
 
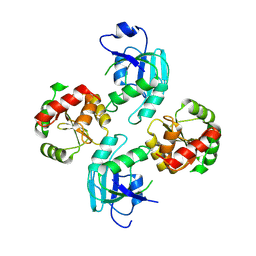 | | human SARS coronavirus unique domain (triclinic form) | | Descriptor: | NON-STRUCTURAL PROTEIN 3 | | Authors: | Tan, J, Vonrhein, C, Smart, O.S, Bricogne, G, Bollati, M, Hansen, G, Mesters, J.R, Hilgenfeld, R. | | Deposit date: | 2009-03-16 | | Release date: | 2009-05-26 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.79 Å) | | Cite: | The Sars-Unique Domain (Sud) of Sars Coronavirus Contains Two Macrodomains that Bind G-Quadruplexes.
Plos Pathog., 5, 2009
|
|
6QW3
 
 | | Calcium-bound gelsolin domain 2 | | Descriptor: | CALCIUM ION, Gelsolin | | Authors: | Scalone, E, Boni, F, Milani, M, Mastrangelo, E, de Rosa, M. | | Deposit date: | 2019-03-05 | | Release date: | 2019-08-28 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | High-resolution crystal structure of gelsolin domain 2 in complex with the physiological calcium ion.
Biochem.Biophys.Res.Commun., 518, 2019
|
|
2QEQ
 
 | |
6Q9Z
 
 | | Crystal structure of the pathological G167R variant of calcium-free human gelsolin, | | Descriptor: | GLYCEROL, Gelsolin, SULFATE ION | | Authors: | Boni, F, Scalone, E, Milani, M, Eloise, M, de Rosa, M. | | Deposit date: | 2018-12-18 | | Release date: | 2019-11-27 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (3.8 Å) | | Cite: | The structure of N184K amyloidogenic variant of gelsolin highlights the role of the H-bond network for protein stability and aggregation properties.
Eur.Biophys.J., 49, 2020
|
|
6QBF
 
 | | Crystal structure of the pathological D187N variant of calcium-free human gelsolin. | | Descriptor: | GLYCEROL, Gelsolin, SODIUM ION, ... | | Authors: | Scalone, E, Boni, F, Milani, M, Eloise, M, de Rosa, M. | | Deposit date: | 2018-12-21 | | Release date: | 2019-11-27 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (3.499 Å) | | Cite: | The structure of N184K amyloidogenic variant of gelsolin highlights the role of the H-bond network for protein stability and aggregation properties.
Eur.Biophys.J., 49, 2020
|
|
6Q9R
 
 | | Crystal structure of the pathological N184K variant of calcium-free human gelsolin | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CHLORIDE ION, GLYCEROL, ... | | Authors: | Scalone, E, Boni, F, Milani, M, Eloise, M, de Rosa, M. | | Deposit date: | 2018-12-18 | | Release date: | 2019-11-27 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.73 Å) | | Cite: | The structure of N184K amyloidogenic variant of gelsolin highlights the role of the H-bond network for protein stability and aggregation properties.
Eur.Biophys.J., 49, 2020
|
|
