7RN1
 
 | |
4UA7
 
 | |
6Y9C
 
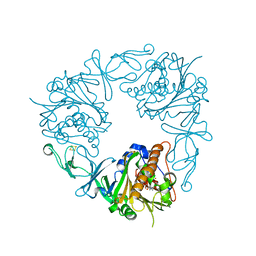 | | The structure of a quaternary ammonium Rieske monooxygenase reveals insights into carnitine oxidation by gut microbiota and inter-subunit electron transfer | | Descriptor: | CARNITINE, Carnitine monooxygenase oxygenase subunit, FE (III) ION, ... | | Authors: | Quareshy, M, Shanmugam, M, Bugg, T.D, Cameron, A, Chen, Y. | | Deposit date: | 2020-03-06 | | Release date: | 2021-03-31 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Characterisation of an unusual cysteine pair in the Rieske carnitine monooxygenase CntA catalytic site.
Febs J., 2023
|
|
6ZGP
 
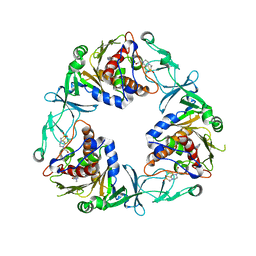 | | Crystal structure of the quaternary ammonium Rieske monooxygenase CntA in complex with inhibitor MMV12 (MMV020670) | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Carnitine monooxygenase oxygenase subunit, FE (III) ION, ... | | Authors: | Quareshy, M, Shanmugam, M, Bugg, T.D.H, Cameron, A, Chen, Y. | | Deposit date: | 2020-06-19 | | Release date: | 2020-11-18 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Structural basis of carnitine monooxygenase CntA substrate specificity, inhibition, and intersubunit electron transfer.
J.Biol.Chem., 296, 2020
|
|
4V8U
 
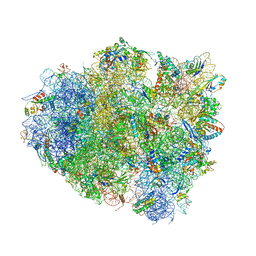 | | Crystal Structure of 70S Ribosome with Both Cognate tRNAs in the E and P Sites Representing an Authentic Elongation Complex. | | Descriptor: | 16S RIBOSOMAL RNA, 23S RIBOSOMAL RNA, 30S RIBOSOMAL PROTEIN S10, ... | | Authors: | Gao, Y.G, Feng, S, Chen, Y. | | Deposit date: | 2012-08-28 | | Release date: | 2014-07-09 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (3.7 Å) | | Cite: | Crystal structure of 70S ribosome with both cognate tRNAs in the E and P sites representing an authentic elongation complex.
PLoS ONE, 8, 2013
|
|
3K9R
 
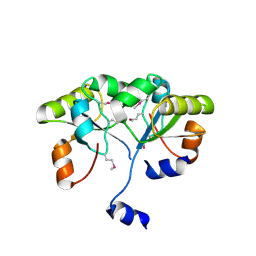 | | X-ray structure of the Rhodanese-like domain of the Alr3790 protein from Anabaena sp. Northeast Structural Genomics Consortium Target NsR437c. | | Descriptor: | Alr3790 protein | | Authors: | Vorobiev, S, Chen, Y, Seetharaman, J, Maglaqui, M, Ciccosanti, C, Mao, L, Xiao, R, Acton, T.B, Montelione, G.T, Tong, L, Hunt, J.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2009-10-16 | | Release date: | 2009-10-27 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | X-ray structure of the Rhodanese-like domain of the Alr3790 protein from Anabaena sp.
To be Published
|
|
1U9C
 
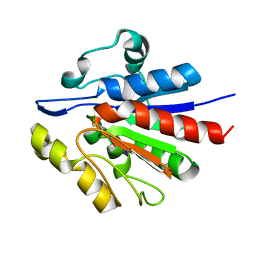 | | Crystallographic structure of APC35852 | | Descriptor: | APC35852 | | Authors: | Borek, D, Chen, Y, Shao, D, Collart, F, Joachimiak, A, Otwinowski, Z, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2004-08-09 | | Release date: | 2004-10-05 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Structural analysis of DJI superfamily
To be Published
|
|
1TZ9
 
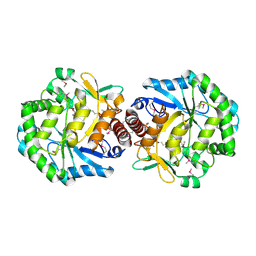 | | Crystal Structure of the Putative Mannonate Dehydratase from Enterococcus faecalis, Northeast Structural Genomics Target EfR41 | | Descriptor: | Mannonate dehydratase | | Authors: | Forouhar, F, Chen, Y, Xiao, R, Cooper, B, Shastry, R, Acton, T.A, Montelione, G.T, Hunt, J.F, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2004-07-09 | | Release date: | 2004-07-20 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal Structure of the Putative Mannonate Dehydratase from Enterococcus faecalis, Northeast Structural Genomics Target EfR41
To be Published
|
|
5Y2V
 | | Strcutrue of the full-length CcmR complexed with 2-OG from Synechocystis PCC6803 | | Descriptor: | 2-OXOGLUTARIC ACID, PHOSPHATE ION, Rubisco operon transcriptional regulator | | Authors: | Jiang, Y.L, Wang, X.P, Sun, H, Cheng, W, Han, S.J, Li, W.F, Chen, Y, Zhou, C.Z. | | Deposit date: | 2017-07-27 | | Release date: | 2017-12-27 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Coordinating carbon and nitrogen metabolic signaling through the cyanobacterial global repressor NdhR.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
3D5N
 
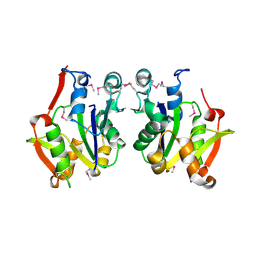 | | Crystal structure of the Q97W15_SULSO protein from Sulfolobus solfataricus. NESG target SsR125. | | Descriptor: | Q97W15_SULSO | | Authors: | Vorobiev, S.M, Chen, Y, Seetharaman, J, Lee, D, Foote, R.E, Maglaqui, M, Janjua, H, Xiao, R, Acton, T.B, Montelione, G.T, Tong, L, Hunt, J.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-05-16 | | Release date: | 2008-07-15 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal structure of the Q97W15_SULSO protein from Sulfolobus solfataricus. NESG target SsR125.
To be Published
|
|
3CEU
 
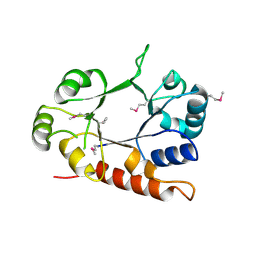 | | Crystal structure of thiamine phosphate pyrophosphorylase (BT_0647) from Bacteroides thetaiotaomicron. Northeast Structural Genomics Consortium target BtR268 | | Descriptor: | Thiamine phosphate pyrophosphorylase | | Authors: | Forouhar, F, Chen, Y, Seetharaman, J, Janjua, H, Mao, L, Xiao, R, Foote, E.L, Maglaqui, M, Ciccosanti, C, Zhao, L, Baran, M.C, Acton, T.B, Montelione, G.T, Hunt, J.F, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-02-29 | | Release date: | 2008-03-25 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of thiamine phosphate pyrophosphorylase (BT_0647) from Bacteroides thetaiotaomicron. Northeast Structural Genomics Consortium target BtR268.
To be Published
|
|
5XU0
 | | Structure of the membrane fusion protein Spr0693 from Streptococcus pneumoniae R6 | | Descriptor: | Membrane-fusion protein | | Authors: | Yang, H.B, Jiang, Y.L, Hou, W.T, Chen, M.T, Chen, Y, Zhou, C.Z. | | Deposit date: | 2017-06-22 | | Release date: | 2018-01-24 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Structure of a MacAB-like efflux pump from Streptococcus pneumoniae.
Nat Commun, 9, 2018
|
|
5XU1
 
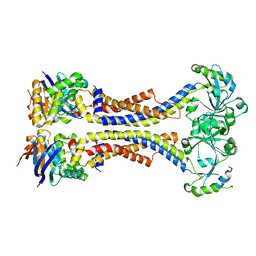 | | Structure of a non-canonical ABC transporter from Streptococcus pneumoniae R6 | | Descriptor: | ABC transporter ATP-binding protein, ABC transporter permeae, MAGNESIUM ION | | Authors: | Yang, H.B, Jiang, Y.L, Hou, W.T, Chen, M.T, Chen, Y, Zhou, C.Z. | | Deposit date: | 2017-06-22 | | Release date: | 2018-01-24 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Structure of a MacAB-like efflux pump from Streptococcus pneumoniae.
Nat Commun, 9, 2018
|
|
5K5S
 
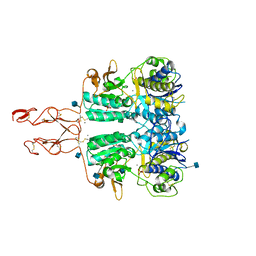 | | Crystal structure of the active form of human calcium-sensing receptor extracellular domain | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, Extracellular calcium-sensing receptor, ... | | Authors: | Geng, Y, Mosyak, L, Kurinov, I, Zuo, H, Sturchler, E, Cheng, T.C, Subramanyam, P, Brown, A.P, Brennan, S.C, Mun, H.-C, Bush, M, Chen, Y, Nguyen, T, Cao, B, Chang, D, Quick, M, Conigrave, A, Colecraft, H.M, McDonald, P, Fan, Q.R. | | Deposit date: | 2016-05-23 | | Release date: | 2016-08-03 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural mechanism of ligand activation in human calcium-sensing receptor.
Elife, 5, 2016
|
|
7S0V
 
 | |
5DEP
 
 | | Structure of Pseudomonas aeruginosa LpxA in complex with UDP-GlcNAc | | Descriptor: | Acyl-[acyl-carrier-protein]--UDP-N-acetylglucosamine O-acyltransferase, PHOSPHATE ION, URIDINE-DIPHOSPHATE-N-ACETYLGLUCOSAMINE | | Authors: | Smith, E.W, Chen, Y. | | Deposit date: | 2015-08-25 | | Release date: | 2015-09-16 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.16 Å) | | Cite: | Structures of Pseudomonas aeruginosa LpxA Reveal the Basis for Its Substrate Selectivity.
Biochemistry, 54, 2015
|
|
5DG3
 
 | |
4XSQ
 
 | | Structure of a variable lymphocyte receptor-like protein Bf66946 from Branchiostoma floridae | | Descriptor: | 1,2-ETHANEDIOL, GLYCEROL, variable lymphocyte receptor-like protein Bf66946 | | Authors: | Cao, D.D, Cheng, W, Jiang, Y.L, Wang, W.J, Li, Q, Chen, Y, Zhou, C.Z. | | Deposit date: | 2015-01-22 | | Release date: | 2016-03-23 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Structure of a variable lymphocyte receptor-like protein from the amphioxus Branchiostoma floridae.
Sci Rep, 6, 2016
|
|
9ORS
 
 | |
9OR7
 
 | | X-ray diffraction structure of CTX-M-14 beta-lactamase soaked with avibactam | | Descriptor: | (2S,5R)-1-formyl-5-[(sulfooxy)amino]piperidine-2-carboxamide, Beta-lactamase, PHOSPHATE ION | | Authors: | Vlahakis, N.W, Rodriguez, J.A, Jacobs, L.M.C, Chen, Y. | | Deposit date: | 2025-05-21 | | Release date: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Combining MicroED and native mass spectrometry for structural discovery of enzyme-small molecule complexes
To Be Published
|
|
9OR3
 
 | | X-ray diffraction structure of CTX-M-14 beta-lactamase co-crystallized with avibactam | | Descriptor: | (2S,5R)-1-formyl-5-[(sulfooxy)amino]piperidine-2-carboxamide, Beta-lactamase, GLYCEROL, ... | | Authors: | Vlahakis, N.W, Rodriguez, J.A, Jacobs, L.M.C, Chen, Y. | | Deposit date: | 2025-05-21 | | Release date: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Combining MicroED and native mass spectrometry for structural discovery of enzyme-small molecule complexes
To Be Published
|
|
9ORG
 
 | |
9ORB
 
 | | X-ray diffraction structure of the CTX-M-14 beta-lactamase-avibactam complex an inhibitor cocktail-soaked crystal | | Descriptor: | (2S,5R)-1-formyl-5-[(sulfooxy)amino]piperidine-2-carboxamide, Beta-lactamase, PHOSPHATE ION | | Authors: | Vlahakis, N.W, Rodriguez, J.A, Jacobs, L.M.C, Chen, Y. | | Deposit date: | 2025-05-21 | | Release date: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Combining MicroED and native mass spectrometry for structural discovery of enzyme-small molecule complexes
To Be Published
|
|
9ORH
 
 | |
9OQE
 
 | |
