5CA6
 
 | |
4JDI
 
 | |
4JDJ
 
 | |
4JDH
 
 | |
3D3M
 
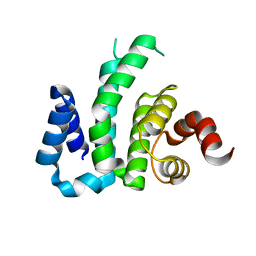 | |
4D8S
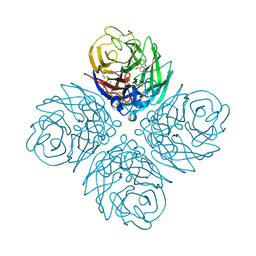 
 | | Influenza NA in complex with antiviral compound | | Descriptor: | CALCIUM ION, Neuraminidase, pentan-3-yl 2-acetamido-2,4-dideoxy-alpha-L-threo-hex-4-enopyranosiduronic acid | | Authors: | Kerry, P.S, Russell, R.J.M.R. | | Deposit date: | 2012-01-11 | | Release date: | 2013-02-13 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.398 Å) | | Cite: | Exploring the interactions of unsaturated glucuronides with influenza virus sialidase.
J.Med.Chem., 55, 2012
|
|
5OBK
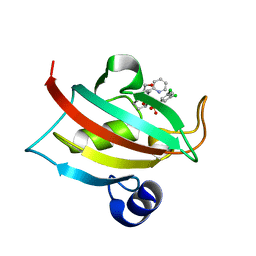 
 | | The Fk1 domain of FKBP51 in complex with (1S,5S,6R)-10-((3,5-dichlorophenyl)sulfonyl)-5-(hydroxymethyl)-3-(pyridin-2-ylmethyl)-3,10-diazabicyclo[4.3.1]decan-2-one | | Descriptor: | (1~{S},5~{S},6~{R})-10-[3,5-bis(chloranyl)phenyl]sulfonyl-5-(hydroxymethyl)-3-(pyridin-2-ylmethyl)-3,10-diazabicyclo[4.3.1]decan-2-one, Peptidyl-prolyl cis-trans isomerase FKBP5 | | Authors: | Pomplun, S, Sippel, C, Haehle, A, Bracher, A, Hausch, F. | | Deposit date: | 2017-06-28 | | Release date: | 2018-04-04 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1 Å) | | Cite: | Chemogenomic Profiling of Human and Microbial FK506-Binding Proteins.
J. Med. Chem., 61, 2018
|
|
3SIS
 
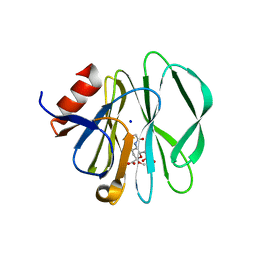 | |
3SIT
 
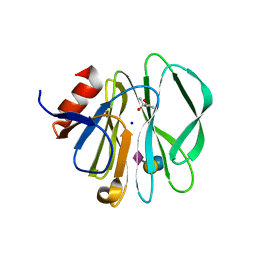 | |
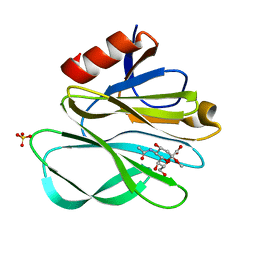
2P3J
 | |
1GH9
 
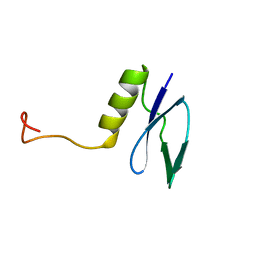 | |
3G45
 
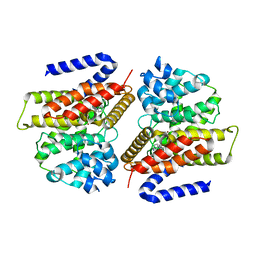 | |
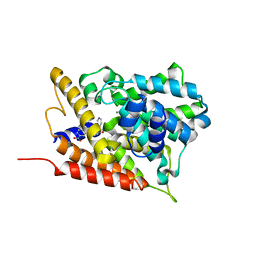
3G4G
 
 | |
3TB0
 
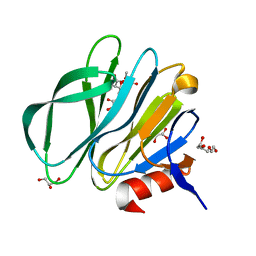 | |
3TAY
 
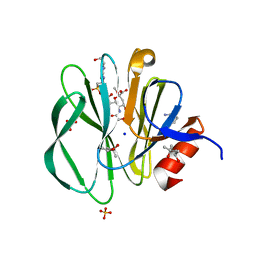 | |
3G4L
 
 | | Crystal structure of human phosphodiesterase 4d with roflumilast | | Descriptor: | 1,2-ETHANEDIOL, 3-(CYCLOPROPYLMETHOXY)-N-(3,5-DICHLOROPYRIDIN-4-YL)-4-(DIFLUOROMETHOXY)BENZAMIDE, MAGNESIUM ION, ... | | Authors: | Staker, B.L. | | Deposit date: | 2009-02-03 | | Release date: | 2010-01-19 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Design of phosphodiesterase 4D (PDE4D) allosteric modulators for enhancing cognition with improved safety.
Nat.Biotechnol., 28, 2010
|
|
3G58
 
 | | Crystal structure of human phosphodiesterase 4d with d155988/pmnpq | | Descriptor: | 1,2-ETHANEDIOL, 8-(3-nitrophenyl)-6-(pyridin-4-ylmethyl)quinoline, MAGNESIUM ION, ... | | Authors: | Staker, B.L. | | Deposit date: | 2009-02-04 | | Release date: | 2010-01-19 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Design of phosphodiesterase 4D (PDE4D) allosteric modulators for enhancing cognition with improved safety.
Nat.Biotechnol., 28, 2010
|
|
3G4I
 
 | | Crystal structure of human phosphodiesterase 4d with d155871 | | Descriptor: | 1-(3-nitrophenyl)-3-(pyridin-4-ylmethyl)pyrido[2,3-d]pyrimidine-2,4(1H,3H)-dione, ETHANOL, MAGNESIUM ION, ... | | Authors: | Staker, B.L. | | Deposit date: | 2009-02-03 | | Release date: | 2010-01-19 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Design of phosphodiesterase 4D (PDE4D) allosteric modulators for enhancing cognition with improved safety.
Nat.Biotechnol., 28, 2010
|
|
3G4K
 
 | | Crystal structure of human phosphodiesterase 4d with rolipram | | Descriptor: | 1,2-ETHANEDIOL, MAGNESIUM ION, ROLIPRAM, ... | | Authors: | Staker, B.L. | | Deposit date: | 2009-02-03 | | Release date: | 2010-01-19 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Design of phosphodiesterase 4D (PDE4D) allosteric modulators for enhancing cognition with improved safety.
Nat.Biotechnol., 28, 2010
|
|
2P3I
 
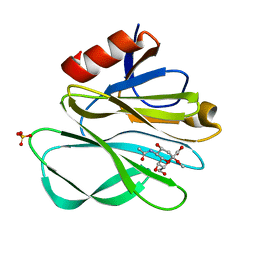 | | Crystal structure of Rhesus Rotavirus VP8* at 295K | | Descriptor: | 2-O-methyl-5-N-acetyl-alpha-D-neuraminic acid, SULFATE ION, VP4 | | Authors: | Blanchard, H. | | Deposit date: | 2007-03-09 | | Release date: | 2008-03-11 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Effects on sialic acid recognition of amino acid mutations in the carbohydrate-binding cleft of the rotavirus spike protein
Glycobiology, 19, 2009
|
|
2P3K
 
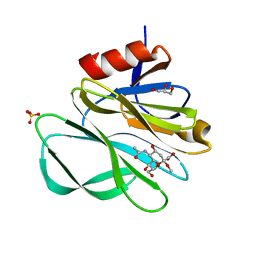 | | Crystal structure of Rhesus rotavirus VP8* at 100K | | Descriptor: | 2-O-methyl-5-N-acetyl-alpha-D-neuraminic acid, GLYCEROL, SULFATE ION, ... | | Authors: | Blanchard, H. | | Deposit date: | 2007-03-09 | | Release date: | 2008-03-11 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Effects on sialic acid recognition of amino acid mutations in the carbohydrate-binding cleft of the rotavirus spike protein
Glycobiology, 19, 2009
|
|
2GJZ
 
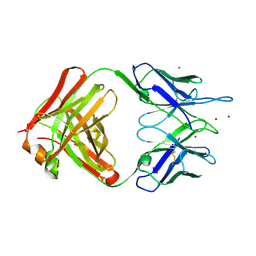 | |
2GK0
 
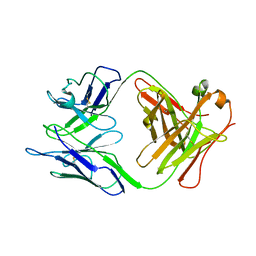 | |
1EJE
 
 | | CRYSTAL STRUCTURE OF AN FMN-BINDING PROTEIN | | Descriptor: | FLAVIN MONONUCLEOTIDE, FMN-BINDING PROTEIN, NICKEL (II) ION, ... | | Authors: | Christendat, D, Saridakis, V, Bochkarev, A, Arrowsmith, C, Edwards, A.M, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2000-03-02 | | Release date: | 2000-10-11 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural proteomics of an archaeon.
Nat.Struct.Biol., 7, 2000
|
|
3GS3
 
 | |
