3FTP
 
 | |
3EZO
 
 | |
3GNC
 
 | |
3GVF
 
 | |
3GK3
 
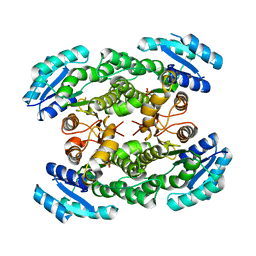 | |
3GTD
 
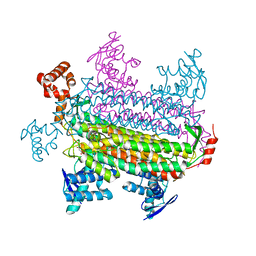 | |
3GVC
 
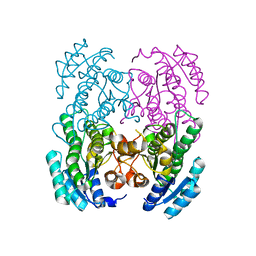 | |
3GK0
 
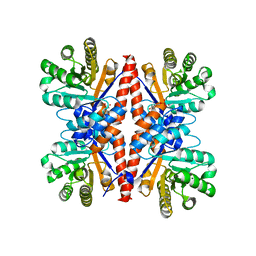 | |
3GW8
 
 | |
3GP5
 
 | |
3GVG
 
 | |
3GWC
 
 | |
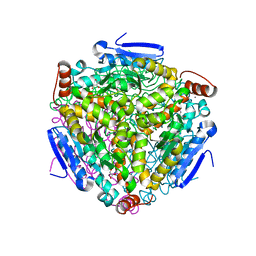
3H81
 
 | |
3GQT
 
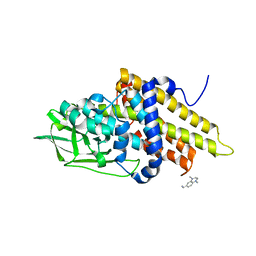 | | Crystal structure of glutaryl-CoA dehydrogenase from Burkholderia pseudomallei with fragment (1,4-dimethyl-1,2,3,4-tetrahydroquinoxalin-6-yl)methylamine | | Descriptor: | 1-(1,4-dimethyl-1,2,3,4-tetrahydroquinoxalin-6-yl)methanamine, Glutaryl-CoA dehydrogenase | | Authors: | Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2009-03-24 | | Release date: | 2009-04-28 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Probing conformational states of glutaryl-CoA dehydrogenase by fragment screening.
Acta Crystallogr.,Sect.F, 67, 2011
|
|
4FZI
 
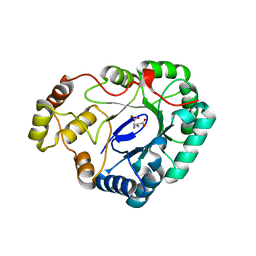 | |
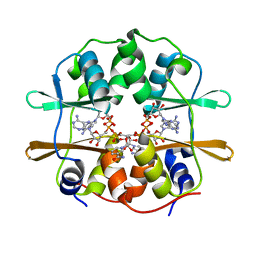
4FRY
 
 | |
4G1K
 
 | |
4G5D
 
 | |
4G67
 
 | |
4G6Z
 
 | |
4G50
 
 | |
4GHK
 
 | |
4GIE
 
 | |
4GK6
 
 | |
4HR3
 
 | |
