5E0Z
 
 | |
6SWG
 
 | |
4M6I
 
 | | Structure of the reduced, Zn-bound form of Mycobacterium tuberculosis peptidoglycan amidase Rv3717 | | 分子名称: | Peptidoglycan Amidase Rv3717, ZINC ION | | 著者 | Prigozhin, D.M, Mavrici, D, Huizar, J.P, Vansell, H.J, Alber, T, TB Structural Genomics Consortium (TBSGC) | | 登録日 | 2013-08-09 | | 公開日 | 2013-09-18 | | 最終更新日 | 2023-09-20 | | 実験手法 | X-RAY DIFFRACTION (2.666 Å) | | 主引用文献 | Structural and Biochemical Analyses of Mycobacterium tuberculosis N-Acetylmuramyl-L-alanine Amidase Rv3717 Point to a Role in Peptidoglycan Fragment Recycling.
J.Biol.Chem., 288, 2013
|
|
4M6H
 
 | | Structure of the reduced, metal-free form of Mycobacterium tuberculosis peptidoglycan amidase Rv3717 | | 分子名称: | Peptidoglycan Amidase Rv3717 | | 著者 | Prigozhin, D.M, Mavrici, D, Huizar, J.P, Vansell, H.J, Alber, T, TB Structural Genomics Consortium (TBSGC) | | 登録日 | 2013-08-09 | | 公開日 | 2013-09-18 | | 最終更新日 | 2023-09-20 | | 実験手法 | X-RAY DIFFRACTION (2.194 Å) | | 主引用文献 | Structural and Biochemical Analyses of Mycobacterium tuberculosis N-Acetylmuramyl-L-alanine Amidase Rv3717 Point to a Role in Peptidoglycan Fragment Recycling.
J.Biol.Chem., 288, 2013
|
|
3OUV
 
 | |
4M6G
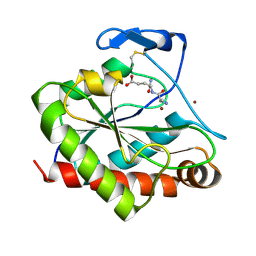 
 | | Structure of the Mycobacterium tuberculosis peptidoglycan amidase Rv3717 in complex with L-Alanine-iso-D-Glutamine reaction product | | 分子名称: | ALANINE, D-alpha-glutamine, Peptidoglycan Amidase Rv3717, ... | | 著者 | Prigozhin, D.M, Mavrici, D, Huizar, J.P, Vansell, H.J, Alber, T, TB Structural Genomics Consortium (TBSGC) | | 登録日 | 2013-08-09 | | 公開日 | 2013-09-18 | | 最終更新日 | 2023-11-15 | | 実験手法 | X-RAY DIFFRACTION (2.104 Å) | | 主引用文献 | Structural and Biochemical Analyses of Mycobacterium tuberculosis N-Acetylmuramyl-L-alanine Amidase Rv3717 Point to a Role in Peptidoglycan Fragment Recycling.
J.Biol.Chem., 288, 2013
|
|
5E12
 | |
5E10
 
 | |
5E0Y
 
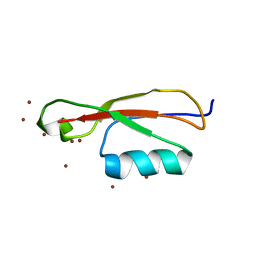 | |
4PPR
 
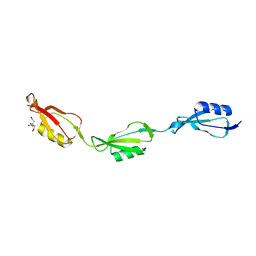
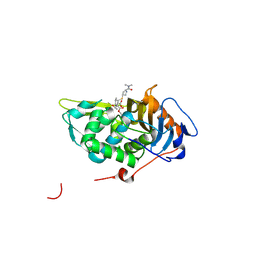 | | Crystal structure of Mycobacterium tuberculosis D,D-peptidase Rv3330 in complex with meropenem | | 分子名称: | (4R,5S)-3-{[(3S,5S)-5-(dimethylcarbamoyl)pyrrolidin-3-yl]sulfanyl}-5-[(2S,3R)-3-hydroxy-1-oxobutan-2-yl]-4-methyl-4,5-d ihydro-1H-pyrrole-2-carboxylic acid, Penicillin-binding protein DacB1 | | 著者 | Prigozhin, D.M, Huizar, J.P, Mavrici, D, Alber, T, TB Structural Genomics Consortium (TBSGC) | | 登録日 | 2014-02-27 | | 公開日 | 2014-11-05 | | 最終更新日 | 2023-09-20 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Subfamily-specific adaptations in the structures of two penicillin-binding proteins from Mycobacterium tuberculosis.
Plos One, 9, 2014
|
|
8QFB
 
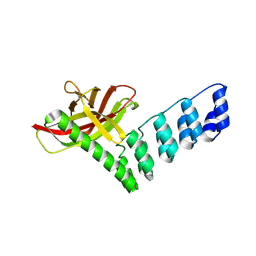 | |
4CGE
 
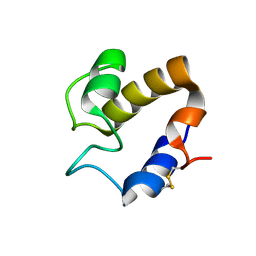 | |
4ESQ
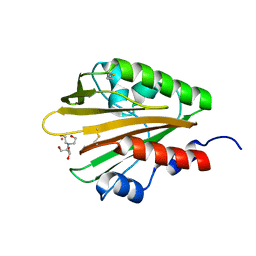 
 | | Crystal structure of the extracellular domain of PknH from Mycobacterium tuberculosis | | 分子名称: | 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Serine/threonine protein kinase, TERBIUM(III) ION | | 著者 | Cavazos, A, Prigozhin, D.M, Alber, T. | | 登録日 | 2012-04-23 | | 公開日 | 2012-07-18 | | 最終更新日 | 2017-11-15 | | 実験手法 | X-RAY DIFFRACTION (1.7 Å) | | 主引用文献 | Structure of the sensor domain of Mycobacterium tuberculosis PknH receptor kinase reveals a conserved binding cleft.
J.Mol.Biol., 422, 2012
|
|
4P0M
 
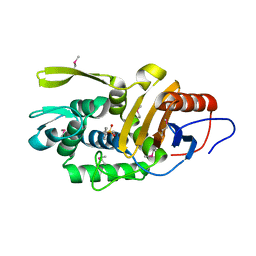 | | Crystal structure of an evolved putative penicillin-binding protein homolog, Rv2911, from Mycobacterium tuberculosis | | 分子名称: | D-alanyl-D-alanine carboxypeptidase | | 著者 | Krieger, I, Yu, M, Bursey, E, Hung, L.-W, Terwilliger, T.C, TB Structural Genomics Consortium (TBSGC) | | 登録日 | 2014-02-21 | | 公開日 | 2014-03-12 | | 最終更新日 | 2023-12-27 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Subfamily-Specific Adaptations in the Structures of Two Penicillin-Binding Proteins from Mycobacterium tuberculosis.
Plos One, 9, 2014
|
|
6VPT
 
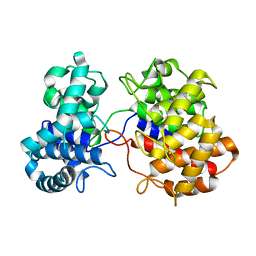 | |
6TL1
 
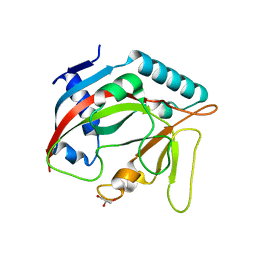 | | Crystal structure of the TASOR pseudo-PARP domain | | 分子名称: | GLYCEROL, Protein TASOR | | 著者 | Douse, C.H, Timms, R.T, Freund, S.M.V, Modis, Y. | | 登録日 | 2019-11-29 | | 公開日 | 2020-09-16 | | 最終更新日 | 2024-05-15 | | 実験手法 | X-RAY DIFFRACTION (2.03 Å) | | 主引用文献 | TASOR is a pseudo-PARP that directs HUSH complex assembly and epigenetic transposon control.
Nat Commun, 11, 2020
|
|
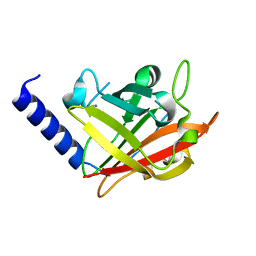
4G1H
 | |
4G1J
 
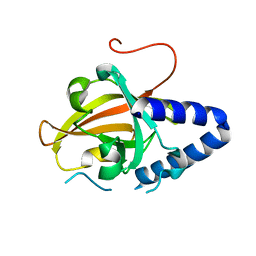 | |
4N8O
 
 | |
4N8N
 
 | |
5TQN
 
 | | Lipoxygenase-1 (soybean) L546A mutant at 293K | | 分子名称: | FE (II) ION, Seed linoleate 13S-lipoxygenase-1 | | 著者 | Poss, E.M, Fraser, J.S, Gee, C. | | 登録日 | 2016-10-24 | | 公開日 | 2017-11-01 | | 最終更新日 | 2023-10-04 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | Biophysical Characterization of a Disabled Double Mutant of Soybean Lipoxygenase: The "Undoing" of Precise Substrate Positioning Relative to Metal Cofactor and an Identified Dynamical Network.
J.Am.Chem.Soc., 141, 2019
|
|
5TQO
 
 | | Lipoxygenase-1 (soybean) L546A/L754A mutant at 300K | | 分子名称: | FE (III) ION, Seed linoleate 13S-lipoxygenase-1 | | 著者 | Poss, E.M, Fraser, J.S, Gee, C. | | 登録日 | 2016-10-24 | | 公開日 | 2017-11-01 | | 最終更新日 | 2023-10-04 | | 実験手法 | X-RAY DIFFRACTION (1.7 Å) | | 主引用文献 | Biophysical Characterization of a Disabled Double Mutant of Soybean Lipoxygenase: The "Undoing" of Precise Substrate Positioning Relative to Metal Cofactor and an Identified Dynamical Network.
J.Am.Chem.Soc., 141, 2019
|
|
5TQP
 
 | | LIPOXYGENASE-1 (SOYBEAN) I553G MUTANT AT 300K | | 分子名称: | FE (III) ION, Seed linoleate 13S-lipoxygenase-1 | | 著者 | Poss, E.M, Fraser, J.S. | | 登録日 | 2016-10-24 | | 公開日 | 2017-11-01 | | 最終更新日 | 2023-10-04 | | 実験手法 | X-RAY DIFFRACTION (1.7 Å) | | 主引用文献 | Hydrogen-Deuterium Exchange of Lipoxygenase Uncovers a Relationship between Distal, Solvent Exposed Protein Motions and the Thermal Activation Barrier for Catalytic Proton-Coupled Electron Tunneling.
ACS Cent Sci, 3, 2017
|
|
5TR0
 
 | | Lipoxygenase-1 (soybean) L754A mutant at 293K | | 分子名称: | FE (II) ION, Seed linoleate 13S-lipoxygenase-1 | | 著者 | Poss, E.M, Fraser, J.S. | | 登録日 | 2016-10-24 | | 公開日 | 2017-11-01 | | 最終更新日 | 2023-10-04 | | 実験手法 | X-RAY DIFFRACTION (1.85 Å) | | 主引用文献 | Biophysical Characterization of a Disabled Double Mutant of Soybean Lipoxygenase: The "Undoing" of Precise Substrate Positioning Relative to Metal Cofactor and an Identified Dynamical Network.
J.Am.Chem.Soc., 141, 2019
|
|
5T5V
 
 | | LIPOXYGENASE-1 (SOYBEAN) AT 293K | | 分子名称: | FE (III) ION, Seed linoleate 13S-lipoxygenase-1 | | 著者 | Poss, E.M, Fraser, J.S. | | 登録日 | 2016-08-31 | | 公開日 | 2017-09-06 | | 最終更新日 | 2023-10-04 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | Hydrogen-Deuterium Exchange of Lipoxygenase Uncovers a Relationship between Distal, Solvent Exposed Protein Motions and the Thermal Activation Barrier for Catalytic Proton-Coupled Electron Tunneling.
ACS Cent Sci, 3, 2017
|
|
