6BY4
 
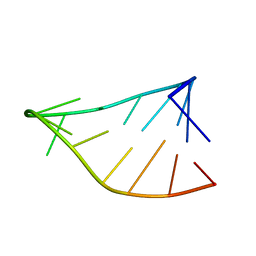 | | Single-State 14-mer UUCG Tetraloop calculated from Exact NOEs | | 分子名称: | RNA (5'-R(P*GP*GP*CP*AP*CP*UP*UP*CP*GP*GP*UP*GP*CP*C)-3') | | 著者 | Nichols, P.J, Henen, M.A, Born, A, Strotz, D, Guntert, P, Vogeli, B. | | 登録日 | 2017-12-19 | | 公開日 | 2018-06-13 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | High-resolution small RNA structures from exact nuclear Overhauser enhancement measurements without additional restraints.
Commun Biol, 1, 2018
|
|
6BY5
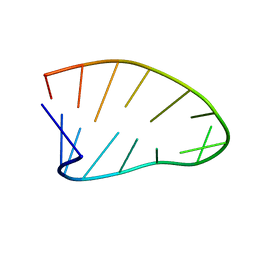 
 | | Two-State 14-mer UUCG Tetraloop calculated from Exact NOEs (State one: Conformers 1-5, State Two: Conformers 6-10) | | 分子名称: | RNA (5'-R(P*GP*GP*CP*AP*CP*UP*UP*CP*GP*GP*UP*GP*CP*C)-3') | | 著者 | Nichols, P.J, Henen, M.A, Born, A, Strotz, D, Guntert, P, Vogeli, B. | | 登録日 | 2017-12-19 | | 公開日 | 2018-06-13 | | 最終更新日 | 2024-05-22 | | 実験手法 | SOLUTION NMR | | 主引用文献 | High-resolution small RNA structures from exact nuclear Overhauser enhancement measurements without additional restraints.
Commun Biol, 1, 2018
|
|
8GBD
 
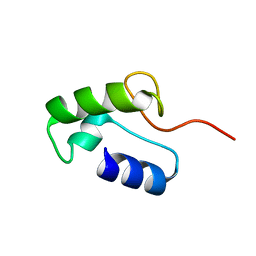 | | Homo sapiens Zalpha mutant - P193A | | 分子名称: | Double-stranded RNA-specific adenosine deaminase | | 著者 | Langeberg, C.J, Vogeli, B, Nichols, P.J, Henen, M, Vicens, Q. | | 登録日 | 2023-02-25 | | 公開日 | 2023-03-22 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Differential Structural Features of Two Mutant ADAR1p150 Z alpha Domains Associated with Aicardi-Goutieres Syndrome.
J.Mol.Biol., 435, 2023
|
|
8GBC
 
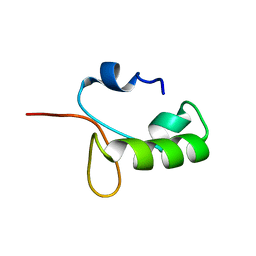 | | Homo sapiens Zalpha mutant - N173S | | 分子名称: | Double-stranded RNA-specific adenosine deaminase | | 著者 | Langeberg, C.J, Nichols, P.J, Henen, M, Vicens, Q, Vogeli, B. | | 登録日 | 2023-02-25 | | 公開日 | 2023-03-22 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Differential Structural Features of Two Mutant ADAR1p150 Z alpha Domains Associated with Aicardi-Goutieres Syndrome.
J.Mol.Biol., 435, 2023
|
|
4Y3B
 
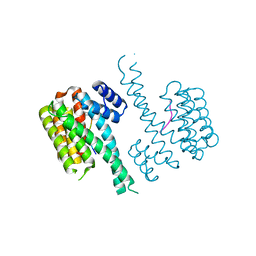 | | Crystal structure of C-terminal modified Tau peptide-hybrid 201D with 14-3-3sigma | | 分子名称: | (2S)-2-(2-methoxyethyl)pyrrolidine, 14-3-3 protein sigma, ARG-THR-PRO-SEP-LEU-PRO-THR-[H][C@@]1(C(C2=CC=CC=C2)C3=CC=CC=C3)CCCN1C | | 著者 | Bartel, M, Milroy, L.G, Brunsveld, L, Ottmann, C. | | 登録日 | 2015-02-10 | | 公開日 | 2015-12-23 | | 最終更新日 | 2024-10-16 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | Stabilizer-Guided Inhibition of Protein-Protein Interactions.
Angew.Chem.Int.Ed.Engl., 54, 2015
|
|
5HF3
 
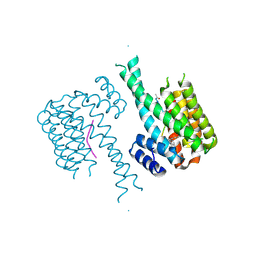 | | Crystal structure of C-terminal modified Tau peptide-hybrid 201D with 14-3-3sigma | | 分子名称: | 14-3-3 protein sigma, modified Tau peptide | | 著者 | Bartel, M, Milroy, L.G, Brunsveld, L, Ottmann, C. | | 登録日 | 2016-01-06 | | 公開日 | 2016-01-27 | | 最終更新日 | 2024-01-10 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | Stabilizer-Guided Inhibition of Protein-Protein Interactions.
Angew.Chem.Int.Ed.Engl., 54, 2015
|
|
4Y32
 
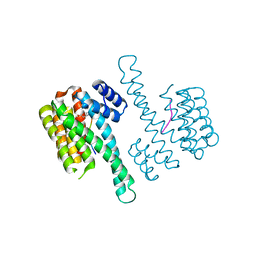 | | Crystal structure of C-terminal modified Tau peptide-hybrid 109B with 14-3-3sigma | | 分子名称: | (2S)-2-(2-methoxyethyl)pyrrolidine, 14-3-3 protein sigma, ARG-THR-PRO-SEP-LEU-PRO-CNC(C(C)O)C(=O)N1CCCC1CCOC | | 著者 | Bartel, M, Milroy, L, Bier, D, Brunsveld, L, Ottmann, C. | | 登録日 | 2015-02-10 | | 公開日 | 2015-12-23 | | 最終更新日 | 2024-10-23 | | 実験手法 | X-RAY DIFFRACTION (1.7 Å) | | 主引用文献 | Stabilizer-Guided Inhibition of Protein-Protein Interactions.
Angew.Chem.Int.Ed.Engl., 54, 2015
|
|
4Y5I
 
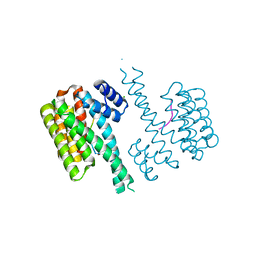 | | Crystal structure of C-terminal modified Tau peptide-hybrid 126B with 14-3-3sigma | | 分子名称: | 14-3-3 protein sigma, CHLORIDE ION, Microtubule-associated protein tau | | 著者 | Leysen, S, Bartel, M, Milroy, L, Brunsveld, L, Ottmann, C. | | 登録日 | 2015-02-11 | | 公開日 | 2016-01-13 | | 最終更新日 | 2024-01-10 | | 実験手法 | X-RAY DIFFRACTION (1.4 Å) | | 主引用文献 | Stabilizer-Guided Inhibition of Protein-Protein Interactions.
Angew.Chem.Int.Ed.Engl., 54, 2015
|
|
7SA5
 
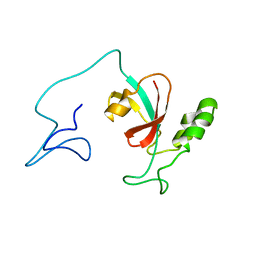 | | Two-state solution NMR structure of Apo Pin1 | | 分子名称: | Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1 | | 著者 | Born, A, Vogeli, B. | | 登録日 | 2021-09-22 | | 公開日 | 2021-10-20 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Reconstruction of Coupled Intra- and Interdomain Protein Motion from Nuclear and Electron Magnetic Resonance.
J.Am.Chem.Soc., 143, 2021
|
|
7SUR
 
 | |
7SUQ
 
 | |
8QPP
 
 | | Bacillus subtilis MutS2-collided disome complex (stalled 70S) | | 分子名称: | 16S rRNA (1533-MER), 23S ribosomal RNA, 30S ribosomal protein S10, ... | | 著者 | Park, E, Mackens-Kiani, T, Berhane, R, Esser, H, Erdenebat, C, Burroughs, A.M, Berninghausen, O, Aravind, L, Beckmann, R, Green, R, Buskirk, A.R. | | 登録日 | 2023-10-02 | | 公開日 | 2023-12-27 | | 最終更新日 | 2024-10-23 | | 実験手法 | ELECTRON MICROSCOPY (3.4 Å) | | 主引用文献 | B. subtilis MutS2 splits stalled ribosomes into subunits without mRNA cleavage.
Embo J., 43, 2024
|
|
