3WP0
 
 | | Crystal structure of Dlg GK in complex with a phosphor-Lgl2 peptide | | 分子名称: | Disks large homolog 4, GLYCEROL, Lethal(2) giant larvae protein homolog 2 | | 著者 | Zhu, J, Shang, Y, Wan, Q, Xia, Y, Chen, J, Du, Q, Zhang, M. | | 登録日 | 2014-01-08 | | 公開日 | 2014-03-19 | | 最終更新日 | 2024-11-13 | | 実験手法 | X-RAY DIFFRACTION (2.039 Å) | | 主引用文献 | Phosphorylation-dependent interaction between tumor suppressors Dlg and Lgl
Cell Res., 24, 2014
|
|
6K9R
 
 | |
6K9O
 
 | | Crystal Structure Analysis of Protein | | 分子名称: | Endo-1,4-beta-xylanase 2, GLYCEROL, IODIDE ION | | 著者 | Li, C, Wan, Q. | | 登録日 | 2019-06-17 | | 公開日 | 2020-06-17 | | 最終更新日 | 2025-03-12 | | 実験手法 | X-RAY DIFFRACTION (1.06 Å) | | 主引用文献 | Studying the Role of a Single Mutation of a Family 11 Glycoside Hydrolase Using High-Resolution X-ray Crystallography.
Protein J., 39, 2020
|
|
6JUG
 
 | |
3TBA
 
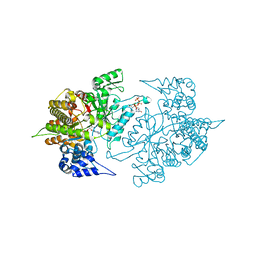 | | Structure of Yeast Ribonucleotide Reductase 1 Q288A with dGTP and ADP | | 分子名称: | 2'-DEOXYGUANOSINE-5'-TRIPHOSPHATE, ADENOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, ... | | 著者 | Ahmad, M.F, Kaushal, P.S, Wan, Q, Wijeratna, S.R, Huang, M, Dealwis, C. | | 登録日 | 2011-08-05 | | 公開日 | 2012-04-04 | | 最終更新日 | 2024-02-28 | | 実験手法 | X-RAY DIFFRACTION (2.8 Å) | | 主引用文献 | Role of Arginine 293 and Glutamine 288 in Communication between Catalytic and Allosteric Sites in Yeast Ribonucleotide Reductase.
J.Mol.Biol., 419, 2012
|
|
3TB9
 
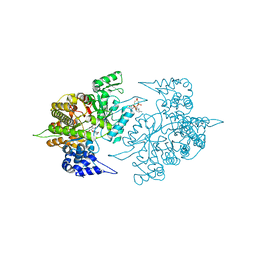 | | Structure of Yeast Ribonucleotide Reductase 1 Q288A with AMPPNP and CDP | | 分子名称: | CYTIDINE-5'-DIPHOSPHATE, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, ... | | 著者 | Ahmad, M.F, Kaushal, P.S, Wan, Q, Wijeratna, S.R, Huang, M, Dealwis, C.D. | | 登録日 | 2011-08-05 | | 公開日 | 2012-04-04 | | 最終更新日 | 2023-09-13 | | 実験手法 | X-RAY DIFFRACTION (2.53 Å) | | 主引用文献 | Role of Arginine 293 and Glutamine 288 in Communication between Catalytic and Allosteric Sites in Yeast Ribonucleotide Reductase.
J.Mol.Biol., 419, 2012
|
|
3S8B
 
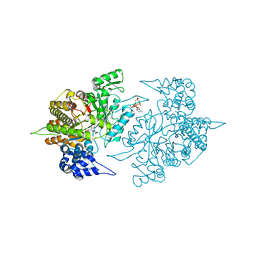 | | Structure of Yeast Ribonucleotide Reductase 1 with AMPPNP and CDP | | 分子名称: | CYTIDINE-5'-DIPHOSPHATE, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, ... | | 著者 | Ahmad, M.F, Kaushal, P.S, Wan, Q, Wijeratna, S.R, Huang, M, Dealwis, C.D. | | 登録日 | 2011-05-27 | | 公開日 | 2012-04-11 | | 最終更新日 | 2024-02-28 | | 実験手法 | X-RAY DIFFRACTION (2.8 Å) | | 主引用文献 | Structural and biochemical basis of lethal mutant R293A of yeast ribonucleotide reductase
To be Published
|
|
8JST
 
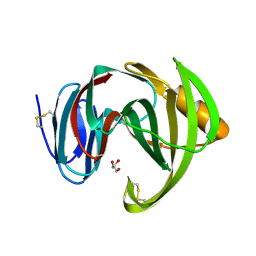 | |
5ZF3
 
 | | Crystal Structures of Endo-beta-1,4-xylanase II Complexed with Xylotriose | | 分子名称: | Endo-1,4-beta-xylanase 2, GLYCEROL, IODIDE ION, ... | | 著者 | Zhang, X, Wan, Q, Li, Z. | | 登録日 | 2018-03-02 | | 公開日 | 2019-03-06 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (1.2 Å) | | 主引用文献 | Crystal Structures of Endo-beta-1,4-xylanase II Complexed with Xylotriose
To be published
|
|
5ZH0
 
 | | Crystal Structures of Endo-beta-1,4-xylanase II | | 分子名称: | Endo-1,4-beta-xylanase 2, GLYCEROL, IODIDE ION | | 著者 | Zhang, X, Wan, Q, Li, Z. | | 登録日 | 2018-03-10 | | 公開日 | 2019-03-13 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (1.08 Å) | | 主引用文献 | Crystal Structures of Endo-beta-1,4-xylanase II
To be published
|
|
5ZH9
 
 | |
6K9W
 
 | |
6KWD
 
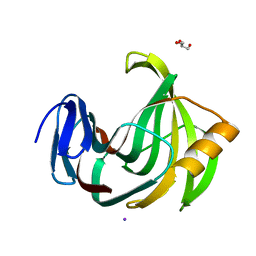 | | Crystal Structure Analysis of Endo-beta-1,4-Xylanase II Complexed with Xylotriose | | 分子名称: | Endo-1,4-beta-xylanase 2, GLYCEROL, IODIDE ION, ... | | 著者 | Li, C, Wan, Q. | | 登録日 | 2019-09-06 | | 公開日 | 2020-12-30 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (1.298 Å) | | 主引用文献 | Studying the Role of a Single Mutation of a Family 11 Glycoside Hydrolase Using High-Resolution X-ray Crystallography.
Protein J., 39, 2020
|
|
6KW9
 
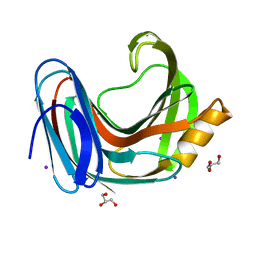 | |
4LLA
 
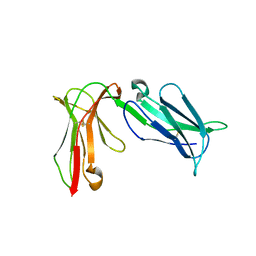 | | Crystal structure of D3D4 domain of the LILRB2 molecule | | 分子名称: | Leukocyte immunoglobulin-like receptor subfamily B member 2 | | 著者 | Nam, G, Shi, Y, Ryu, M, Wang, Q, Song, H, Liu, J, Yan, J, Qi, J, Gao, G.F. | | 登録日 | 2013-07-09 | | 公開日 | 2013-09-11 | | 最終更新日 | 2024-11-13 | | 実験手法 | X-RAY DIFFRACTION (2.502 Å) | | 主引用文献 | Crystal structures of the two membrane-proximal Ig-like domains (D3D4) of LILRB1/B2: alternative models for their involvement in peptide-HLA binding
Protein Cell, 4, 2013
|
|
4LL9
 
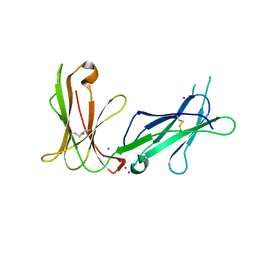 | | Crystal structure of D3D4 domain of the LILRB1 molecule | | 分子名称: | IODIDE ION, Leukocyte immunoglobulin-like receptor subfamily B member 1 | | 著者 | Nam, G, Shi, Y, Ryu, M, Wang, Q, Song, H, Liu, J, Yan, J, Qi, J, Gao, G.F. | | 登録日 | 2013-07-09 | | 公開日 | 2013-09-11 | | 最終更新日 | 2024-10-30 | | 実験手法 | X-RAY DIFFRACTION (2.686 Å) | | 主引用文献 | Crystal structures of the two membrane-proximal Ig-like domains (D3D4) of LILRB1/B2: alternative models for their involvement in peptide-HLA binding
Protein Cell, 4, 2013
|
|
1KAK
 
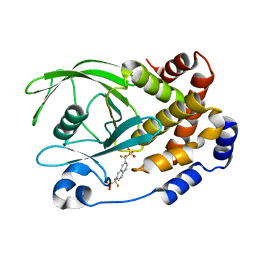 | | Human Tyrosine Phosphatase 1B Complexed with an Inhibitor | | 分子名称: | PROTEIN-TYROSINE PHOSPHATASE, NON-RECEPTOR TYPE 1, {[7-(DIFLUORO-PHOSPHONO-METHYL)-NAPHTHALEN-2-YL]-DIFLUORO-METHYL}-PHOSPHONIC ACID | | 著者 | Jia, Z, Ye, Q, Dinaut, A.N, Wang, Q, Waddleton, D, Payette, P, Ramachandran, C, Kennedy, B, Hum, G, Taylor, S.D. | | 登録日 | 2001-11-02 | | 公開日 | 2002-06-19 | | 最終更新日 | 2023-08-16 | | 実験手法 | X-RAY DIFFRACTION (2.5 Å) | | 主引用文献 | Structure of protein tyrosine phosphatase 1B in complex with inhibitors bearing two phosphotyrosine mimetics.
J.Med.Chem., 44, 2001
|
|
1KAV
 
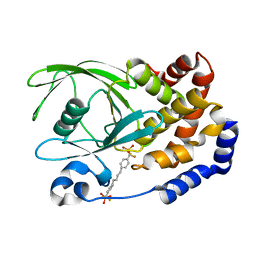 | | Human Tyrosine Phosphatase 1B Complexed with an Inhibitor | | 分子名称: | PROTEIN-TYROSINE PHOSPHATASE, NON-RECEPTOR TYPE 1, [(4-{4-[4-(DIFLUORO-PHOSPHONO-METHYL)-PHENYL]-BUTYL}-PHENYL)-DIFLUORO-METHYL]-PHOSPHONIC ACID | | 著者 | Jia, Z, Ye, Q, Dinaut, A.N, Wang, Q, Waddleton, D, Payette, P, Ramachandran, C, Kennedy, B, Hum, G, Taylor, S.D. | | 登録日 | 2001-11-03 | | 公開日 | 2002-06-19 | | 最終更新日 | 2023-08-16 | | 実験手法 | X-RAY DIFFRACTION (2.35 Å) | | 主引用文献 | Structure of protein tyrosine phosphatase 1B in complex with inhibitors bearing two phosphotyrosine mimetics.
J.Med.Chem., 44, 2001
|
|
8BZN
 
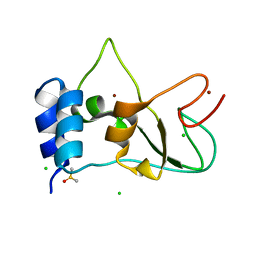 | | SARS-CoV-2 non-structural protein 10 (nsp10) variant T102I | | 分子名称: | CHLORIDE ION, DIMETHYL SULFOXIDE, Replicase polyprotein 1ab, ... | | 著者 | Wang, H, Rizvi, S.R.A, Dong, D, Lou, J, Wang, Q, Sopipong, W, Najar, F, Agarwal, P.K, Kozielski, F, Haider, S. | | 登録日 | 2022-12-15 | | 公開日 | 2023-12-27 | | 最終更新日 | 2024-01-10 | | 実験手法 | X-RAY DIFFRACTION (2.19 Å) | | 主引用文献 | Emerging variants of SARS-CoV-2 NSP10 highlight strong functional conservation of its binding to two non-structural proteins, NSP14 and NSP16.
Elife, 12, 2023
|
|
7H9O
 
 | | PanDDA analysis group deposition -- Crystal structure of HSP90N in complex with Fr13146 | | 分子名称: | Heat shock protein HSP 90-alpha, {4-[(oxan-4-yl)oxy]phenyl}methanol | | 著者 | Huang, L, Wang, W, Zhu, Z, Li, Q, Li, M, Zhou, H, Xu, Q, Wen, W, Wang, Q, Yu, F. | | 登録日 | 2024-07-10 | | 公開日 | 2025-03-26 | | 実験手法 | X-RAY DIFFRACTION (1.55 Å) | | 主引用文献 | Novel starting points for fragment-based drug design against human heat-shock protein 90 identified using crystallographic fragment screening.
Iucrj, 12, 2025
|
|
7HB6
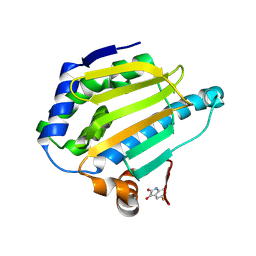 
 | | PanDDA analysis group deposition -- Crystal structure of HSP90N in complex with Fr13755 | | 分子名称: | 5-fluoranyl-1~{H}-indole-2,3-dione, Heat shock protein HSP 90-alpha | | 著者 | Huang, L, Wang, W, Zhu, Z, Li, Q, Li, M, Zhou, H, Xu, Q, Wen, W, Wang, Q, Yu, F. | | 登録日 | 2024-07-10 | | 公開日 | 2025-03-26 | | 実験手法 | X-RAY DIFFRACTION (2.09 Å) | | 主引用文献 | Novel starting points for fragment-based drug design against human heat-shock protein 90 identified using crystallographic fragment screening.
Iucrj, 12, 2025
|
|
7HBK
 
 | | PanDDA analysis group deposition -- Crystal structure of HSP90N in complex with Fr13232 | | 分子名称: | 6-(2,3-dimethylphenoxy)pyridin-3-amine, Heat shock protein HSP 90-alpha | | 著者 | Huang, L, Wang, W, Zhu, Z, Li, Q, Li, M, Zhou, H, Xu, Q, Wen, W, Wang, Q, Yu, F. | | 登録日 | 2024-07-10 | | 公開日 | 2025-03-26 | | 実験手法 | X-RAY DIFFRACTION (2.14 Å) | | 主引用文献 | Novel starting points for fragment-based drug design against human heat-shock protein 90 identified using crystallographic fragment screening.
Iucrj, 12, 2025
|
|
7HC1
 
 | | PanDDA analysis group deposition -- Crystal structure of HSP90N in complex with 10T-0263 | | 分子名称: | (3M)-3-(3-fluoro-4-methoxyphenyl)-4-methyl-1H-pyrazole, Heat shock protein HSP 90-alpha | | 著者 | Huang, L, Wang, W, Zhu, Z, Li, Q, Li, M, Zhou, H, Xu, Q, Wen, W, Wang, Q, Yu, F. | | 登録日 | 2024-07-10 | | 公開日 | 2025-03-26 | | 実験手法 | X-RAY DIFFRACTION (2.13 Å) | | 主引用文献 | Novel starting points for fragment-based drug design against human heat-shock protein 90 identified using crystallographic fragment screening.
Iucrj, 12, 2025
|
|
7H9V
 
 | | PanDDA analysis group deposition -- Crystal structure of HSP90N in complex with Fr12319 | | 分子名称: | 8-carbamoyl-1-benzopyran-1-ium, Heat shock protein HSP 90-alpha | | 著者 | Huang, L, Wang, W, Zhu, Z, Li, Q, Li, M, Zhou, H, Xu, Q, Wen, W, Wang, Q, Yu, F. | | 登録日 | 2024-07-10 | | 公開日 | 2025-03-26 | | 実験手法 | X-RAY DIFFRACTION (1.52 Å) | | 主引用文献 | Novel starting points for fragment-based drug design against human heat-shock protein 90 identified using crystallographic fragment screening.
Iucrj, 12, 2025
|
|
7H9W
 
 | | PanDDA analysis group deposition -- Crystal structure of HSP90N in complex with Fr12314 | | 分子名称: | 3,4-dihydro-2~{H}-chromene-6-carboxamide, Heat shock protein HSP 90-alpha | | 著者 | Huang, L, Wang, W, Zhu, Z, Li, Q, Li, M, Zhou, H, Xu, Q, Wen, W, Wang, Q, Yu, F. | | 登録日 | 2024-07-10 | | 公開日 | 2025-03-26 | | 実験手法 | X-RAY DIFFRACTION (1.64 Å) | | 主引用文献 | Novel starting points for fragment-based drug design against human heat-shock protein 90 identified using crystallographic fragment screening.
Iucrj, 12, 2025
|
|
