8YVA
 
 | |
8YV9
 
 | |
6IFE
 
 | | A Glycoside Hydrolase Family 43 beta-Xylosidase | | 分子名称: | Beta-xylosidase, GLYCEROL | | 著者 | Li, N, Liu, Y, Zhang, R, Zhou, J.P, Huang, Z.X. | | 登録日 | 2018-09-20 | | 公開日 | 2019-03-13 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (1.804 Å) | | 主引用文献 | Biochemical and structural properties of a low-temperature-active glycoside hydrolase family 43 beta-xylosidase: Activity and instability at high neutral salt concentrations.
Food Chem, 301, 2019
|
|
5KE6
 
 | |
5KEB
 
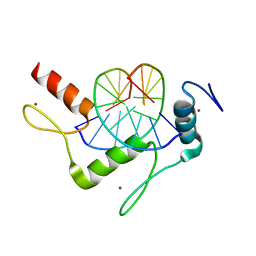 | |
4UTB
 
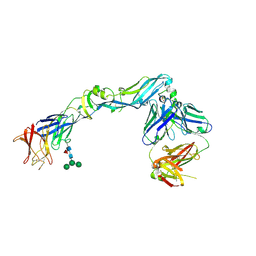 | | Crystal structure of dengue 2 virus envelope glycoprotein in complex with the Fab fragment of the broadly neutralizing human antibody EDE2 A11 | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, BROADLY NEUTRALIZING HUMAN ANTIBODY EDE2 A11, ... | | 著者 | Rouvinski, A, Guardado-Calvo, P, Barba-Spaeth, G, Duquerroy, S, Vaney, M.C, Rey, F.A. | | 登録日 | 2014-07-18 | | 公開日 | 2015-01-28 | | 最終更新日 | 2024-10-16 | | 実験手法 | X-RAY DIFFRACTION (3.85 Å) | | 主引用文献 | Recognition Determinants of Broadly Neutralizing Human Antibodies Against Dengue Viruses.
Nature, 520, 2015
|
|
6IIJ
 
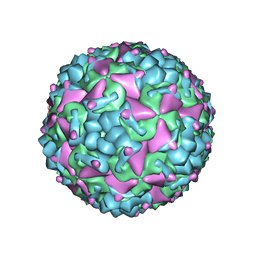 | | Cryo-EM structure of CV-A10 mature virion | | 分子名称: | SPHINGOSINE, VP1, VP2, ... | | 著者 | Chen, J.H, Ye, X.H, Cong, Y, Huang, Z. | | 登録日 | 2018-10-06 | | 公開日 | 2018-11-07 | | 最終更新日 | 2025-07-02 | | 実験手法 | ELECTRON MICROSCOPY (2.84 Å) | | 主引用文献 | Coxsackievirus A10 atomic structure facilitating the discovery of a broad-spectrum inhibitor against human enteroviruses.
Cell Discov, 5, 2019
|
|
5K5L
 
 | |
5K5H
 
 | |
5KE8
 
 | |
5YAD
 
 | |
6IIO
 
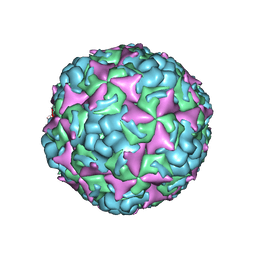 | | Cryo-EM structure of CV-A10 native empty particle | | 分子名称: | VP0, VP1, VP3 | | 著者 | Chen, J.H, Ye, X.H, Cong, Y, Huang, Z. | | 登録日 | 2018-10-07 | | 公開日 | 2018-11-07 | | 最終更新日 | 2025-06-25 | | 実験手法 | ELECTRON MICROSCOPY (3.12 Å) | | 主引用文献 | Coxsackievirus A10 atomic structure facilitating the discovery of a broad-spectrum inhibitor against human enteroviruses.
Cell Discov, 5, 2019
|
|
5K5J
 
 | |
4UT7
 
 | | CRYSTAL STRUCTURE OF THE SCFV FRAGMENT OF THE BROADLY NEUTRALIZING HUMAN ANTIBODY EDE2 A11 | | 分子名称: | BROADLY NEUTRALIZING HUMAN ANTIBODY EDE2 A11 | | 著者 | Rouvinski, A, Guardado-Calvo, P, Barba-Spaeth, G, Duquerroy, S, Vaney, M.C, Rey, F.A. | | 登録日 | 2014-07-18 | | 公開日 | 2015-01-28 | | 最終更新日 | 2024-11-13 | | 実験手法 | X-RAY DIFFRACTION (1.7 Å) | | 主引用文献 | Recognition Determinants of Broadly Neutralizing Human Antibodies Against Dengue Viruses.
Nature, 520, 2015
|
|
5KE9
 
 | |
7Y9J
 
 | | Crystal structure of P450 BM3-TMK from Bacillus megaterium in complex with 5-nitro-1,2-benzisoxazole | | 分子名称: | 5-nitro-1,2-benzoxazole, Bifunctional cytochrome P450/NADPH--P450 reductase, PROTOPORPHYRIN IX CONTAINING FE | | 著者 | Wang, Q, Zhang, L.L, Liu, W.D, Huang, J.-W, Yang, Y, Chen, C.-C, Guo, R.-T. | | 登録日 | 2022-06-24 | | 公開日 | 2023-06-28 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (1.83 Å) | | 主引用文献 | Engineering of a P450-based Kemp eliminase with a new mechanism
Chinese J Catal, 47, 2023
|
|
7Y9K
 
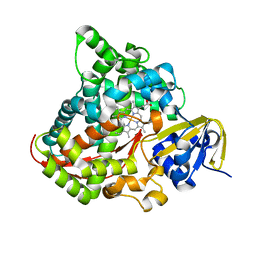 | | Crystal structure of P450 BM3-TMK from Bacillus megaterium | | 分子名称: | Bifunctional cytochrome P450/NADPH--P450 reductase, PROTOPORPHYRIN IX CONTAINING FE | | 著者 | Wang, Q, Zhang, L.L, Liu, W.D, Huang, J.-W, Yang, Y, Chen, C.-C, Guo, R.-T. | | 登録日 | 2022-06-25 | | 公開日 | 2023-06-28 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (2.23 Å) | | 主引用文献 | Engineering of a P450-based Kemp eliminase with a new mechanism
Chinese J Catal, 47, 2023
|
|
5YAA
 
 | |
5KEA
 
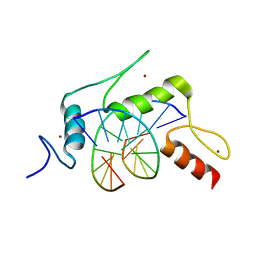 | |
7Z8O
 
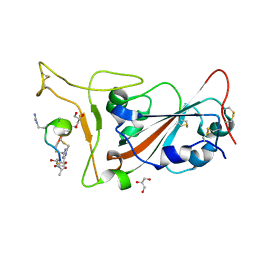 | | Crystal structure of SARS-CoV-2 S RBD in complex with a stapled peptide | | 分子名称: | 2,4,6-tris(chloromethyl)-1,3,5-triazine, GLYCEROL, Spike protein S1, ... | | 著者 | Brear, P, Chen, L, Gaynor, K, Harman, M, Dods, R, Hyvonen, M. | | 登録日 | 2022-03-18 | | 公開日 | 2023-06-28 | | 最終更新日 | 2024-02-07 | | 実験手法 | X-RAY DIFFRACTION (0.96 Å) | | 主引用文献 | Multivalent bicyclic peptides are an effective antiviral modality that can potently inhibit SARS-CoV-2.
Nat Commun, 14, 2023
|
|
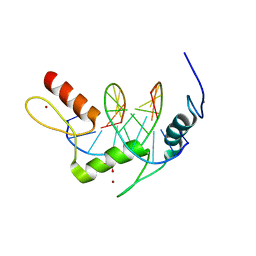
5KL6
 
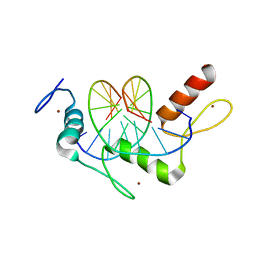 | | Wilms Tumor Protein (WT1) Q369R ZnF2-4 in complex with DNA | | 分子名称: | DNA (5'-D(*AP*GP*CP*GP*TP*GP*GP*GP*GP*GP*T)-3'), DNA (5'-D(*TP*AP*CP*CP*CP*CP*CP*AP*CP*GP*C)-3'), Wilms tumor protein, ... | | 著者 | Hashimoto, H, Cheng, X. | | 登録日 | 2016-06-23 | | 公開日 | 2016-09-14 | | 最終更新日 | 2023-09-27 | | 実験手法 | X-RAY DIFFRACTION (1.641 Å) | | 主引用文献 | Denys-Drash syndrome associated WT1 glutamine 369 mutants have altered sequence-preferences and altered responses to epigenetic modifications.
Nucleic Acids Res., 44, 2016
|
|
5KL4
 
 | |
8YY8
 
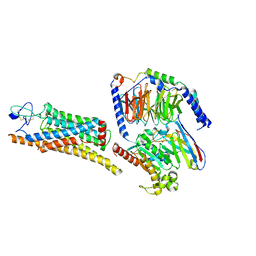 | | Fzd7 -Gs complex | | 分子名称: | Frizzled-7, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | 著者 | Chen, B, Xu, L, Han, G.W, Xu, F. | | 登録日 | 2024-04-03 | | 公開日 | 2024-04-24 | | 最終更新日 | 2025-06-25 | | 実験手法 | ELECTRON MICROSCOPY (3.22 Å) | | 主引用文献 | Cryo-EM structure of constitutively active human Frizzled 7 in complex with heterotrimeric G s .
Cell Res., 31, 2021
|
|
8AAA
 
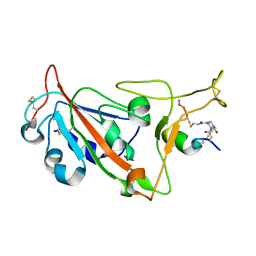 | | Crystal structure of SARS-CoV-2 S RBD in complex with a stapled peptide | | 分子名称: | 1,1',1''-(1,3,5-triazinane-1,3,5-triyl)tripropan-1-one, Spike protein S1, Stapled peptide | | 著者 | Brear, P, Chen, L, Gaynor, K, Harman, M, Dods, R, Hyvonen, M. | | 登録日 | 2022-06-30 | | 公開日 | 2023-06-28 | | 最終更新日 | 2024-10-23 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | Multivalent bicyclic peptides are an effective antiviral modality that can potently inhibit SARS-CoV-2.
Nat Commun, 14, 2023
|
|
4V40
 
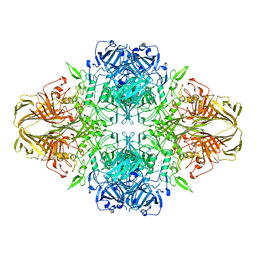 | | BETA-GALACTOSIDASE | | 分子名称: | BETA-GALACTOSIDASE, MAGNESIUM ION | | 著者 | Jacobson, R.H, Zhang, X, Dubose, R.F, Matthews, B.W. | | 登録日 | 1994-07-18 | | 公開日 | 2014-07-09 | | 最終更新日 | 2024-02-28 | | 実験手法 | X-RAY DIFFRACTION (2.5 Å) | | 主引用文献 | Three-dimensional structure of beta-galactosidase from E. coli.
Nature, 369, 1994
|
|
