1LYV
 
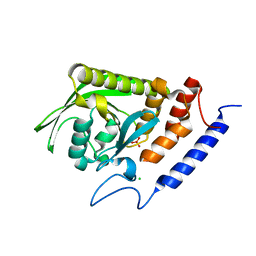 | | High-resolution structure of the catalytically inactive yersinia tyrosine phosphatase C403A mutant in complex with phosphate. | | 分子名称: | CHLORIDE ION, PHOSPHATE ION, PROTEIN-TYROSINE PHOSPHATASE YOPH | | 著者 | Evdokimov, A.G, Waugh, D.S, Routzahn, K, Tropea, J, Cherry, S. | | 登録日 | 2002-06-08 | | 公開日 | 2002-07-03 | | 最終更新日 | 2024-02-14 | | 実験手法 | X-RAY DIFFRACTION (1.363 Å) | | 主引用文献 | High-resolution structure of the catalytically inactive yersinia tyrosine phosphatase C403A mutant in complex with phosphate.
To be Published
|
|
5VFI
 
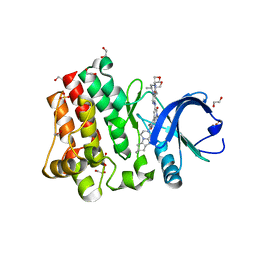 | | Bruton's tyrosine kinase (BTK) with GDC-0853 | | 分子名称: | 1,2-ETHANEDIOL, 2-[3'-(hydroxymethyl)-1-methyl-5-({5-[(2S)-2-methyl-4-(oxetan-3-yl)piperazin-1-yl]pyridin-2-yl}amino)-6-oxo[1,6-dihydro[3,4'-bipyridine]]-2'-yl]-7,7-dimethyl-3,4,7,8-tetrahydro-2H-cyclopenta[4,5]pyrrolo[1,2-a]pyrazin-1(6H)-one, SULFATE ION, ... | | 著者 | Steinbacher, S, Eigenbrot, C. | | 登録日 | 2017-04-07 | | 公開日 | 2018-02-28 | | 最終更新日 | 2024-03-13 | | 実験手法 | X-RAY DIFFRACTION (1.59 Å) | | 主引用文献 | Discovery of GDC-0853: A Potent, Selective, and Noncovalent Bruton's Tyrosine Kinase Inhibitor in Early Clinical Development.
J. Med. Chem., 61, 2018
|
|
3UN0
 
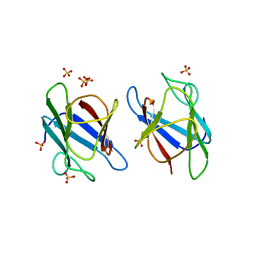 | | Crystal Structure of MDC1 FHA Domain | | 分子名称: | Mediator of DNA damage checkpoint protein 1, SULFATE ION | | 著者 | Clapperton, J.A, Lloyd, J, Haire, L.F, Li, J, Smerdon, S.J. | | 登録日 | 2011-11-15 | | 公開日 | 2011-12-28 | | 最終更新日 | 2024-02-28 | | 実験手法 | X-RAY DIFFRACTION (2.3 Å) | | 主引用文献 | The molecular basis of ATM-dependent dimerization of the Mdc1 DNA damage checkpoint mediator.
Nucleic Acids Res., 40, 2012
|
|
3UOT
 
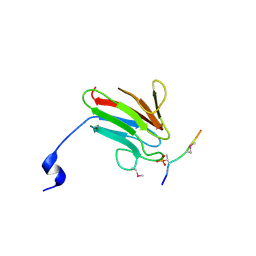 | | Crystal Structure of MDC1 FHA Domain in Complex with a Phosphorylated Peptide from the MDC1 N-terminus | | 分子名称: | Mediator of DNA damage checkpoint protein 1 | | 著者 | Clapperton, J.A, Lloyd, J, Haire, L.F, Li, J, Smerdon, S.J. | | 登録日 | 2011-11-17 | | 公開日 | 2011-12-28 | | 最終更新日 | 2012-07-18 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | The molecular basis of ATM-dependent dimerization of the Mdc1 DNA damage checkpoint mediator.
Nucleic Acids Res., 40, 2012
|
|
4UMN
 
 | | Structure of a stapled peptide antagonist bound to Nutlin-resistant Mdm2. | | 分子名称: | E3 ubiquitin-protein ligase Mdm2, M06 | | 著者 | Chee, S, Wongsantichon, J, Quah, S, Robinson, R.C, Verma, C, Lane, D.P, Brown, C.J, Ghadessy, F.J. | | 登録日 | 2014-05-20 | | 公開日 | 2014-05-28 | | 最終更新日 | 2024-02-07 | | 実験手法 | X-RAY DIFFRACTION (1.99 Å) | | 主引用文献 | Structure of a stapled peptide antagonist bound to nutlin-resistant Mdm2.
PLoS ONE, 9, 2014
|
|
7LFV
 
 | |
7LFU
 
 | |
4FR2
 
 | | Alcohol dehydrogenase from Oenococcus oeni | | 分子名称: | 1,3-propanediol dehydrogenase, NICKEL (II) ION | | 著者 | Fodor, K, Skander, E, Wilmanns, M. | | 登録日 | 2012-06-26 | | 公開日 | 2013-06-05 | | 最終更新日 | 2023-09-13 | | 実験手法 | X-RAY DIFFRACTION (3.2 Å) | | 主引用文献 | Structural and biochemical characterisation of a NAD(+)-dependent alcohol dehydrogenase from Oenococcus oeni as a new model molecule for industrial biotechnology applications.
Appl.Microbiol.Biotechnol., 97, 2013
|
|
7MWI
 
 | | Crystal structure of human BAZ2A | | 分子名称: | Bromodomain adjacent to zinc finger domain protein 2A, UNKNOWN ATOM OR ION | | 著者 | Liu, K, Dong, A, Li, Y, Loppnau, P, Edwards, A.M, Arrowsmith, C.H, Min, J, Structural Genomics Consortium (SGC) | | 登録日 | 2021-05-17 | | 公開日 | 2021-12-29 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | Structural basis of the TAM domain of BAZ2A in binding to DNA or RNA independent of methylation status.
J.Biol.Chem., 297, 2021
|
|
5CZJ
 
 | |
8K8A
 
 | |
8K89
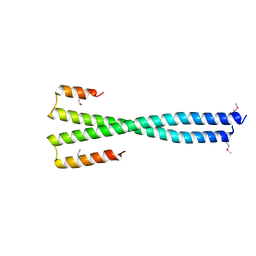 
 | | Crystal structure of NFIL3 | | 分子名称: | Nuclear factor interleukin-3-regulated protein | | 著者 | Min, J.R, Chen, S.Z, Liu, K. | | 登録日 | 2023-07-29 | | 公開日 | 2024-03-06 | | 最終更新日 | 2024-03-27 | | 実験手法 | X-RAY DIFFRACTION (2.1 Å) | | 主引用文献 | Structural basis for specific DNA sequence recognition by the transcription factor NFIL3.
J.Biol.Chem., 300, 2024
|
|
8K8D
 
 | |
8K86
 
 | | Crystal structure of NFIL3 in complex with TTATGTAA DNA | | 分子名称: | DNA (5'-D(*CP*AP*TP*TP*AP*TP*GP*TP*AP*AP*CP*G)-3'), DNA (5'-D(*CP*GP*TP*TP*AP*CP*AP*TP*AP*AP*TP*G)-3'), Nuclear factor interleukin-3-regulated protein | | 著者 | Min, J.R, Chen, S.Z, Liu, K. | | 登録日 | 2023-07-28 | | 公開日 | 2024-03-06 | | 最終更新日 | 2024-03-27 | | 実験手法 | X-RAY DIFFRACTION (2.06 Å) | | 主引用文献 | Structural basis for specific DNA sequence recognition by the transcription factor NFIL3.
J.Biol.Chem., 300, 2024
|
|
8K8C
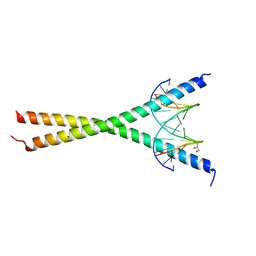 
 | | Crystal structure of C/EBPalpha BZIP domain bound to a high affinity DNA | | 分子名称: | CCAAT/enhancer-binding protein alpha, DNA (5'-D(*CP*AP*TP*TP*AP*CP*GP*TP*AP*AP*TP*GP*A)-3'), DNA (5'-D(*CP*AP*TP*TP*AP*CP*GP*TP*AP*AP*TP*GP*T)-3'), ... | | 著者 | Min, J.R, Chen, S.Z, Liu, K. | | 登録日 | 2023-07-29 | | 公開日 | 2024-03-06 | | 最終更新日 | 2024-03-27 | | 実験手法 | X-RAY DIFFRACTION (2.06 Å) | | 主引用文献 | Structural basis for specific DNA sequence recognition by the transcription factor NFIL3.
J.Biol.Chem., 300, 2024
|
|
6XDF
 
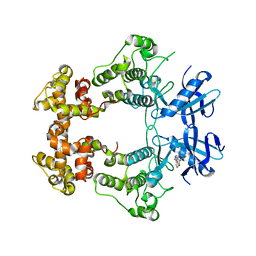 | | Crystal structure of IRE1a in complex with G-4100 | | 分子名称: | 4-amino-N-(6-chloro-2-fluoro-3-{[(pyrrolidin-1-yl)sulfonyl]amino}phenyl)quinazoline-8-carboxamide, SODIUM ION, Serine/threonine-protein kinase/endoribonuclease IRE1 | | 著者 | Steinbacher, S, Wang, W. | | 登録日 | 2020-06-10 | | 公開日 | 2021-04-21 | | 最終更新日 | 2024-03-06 | | 実験手法 | X-RAY DIFFRACTION (2.54 Å) | | 主引用文献 | Identification of BRaf-Sparing Amino-Thienopyrimidines with Potent IRE1 alpha Inhibitory Activity.
Acs Med.Chem.Lett., 11, 2020
|
|
6BOZ
 
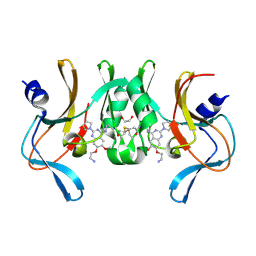 | | Structure of human SETD8 in complex with covalent inhibitor MS4138 | | 分子名称: | 1,2-ETHANEDIOL, N-(3-{[7-(2-aminoethoxy)-6-methoxy-2-(pyrrolidin-1-yl)quinazolin-4-yl]amino}propyl)prop-2-enamide, N-lysine methyltransferase KMT5A | | 著者 | Babault, N, Anqi, M, Jin, J. | | 登録日 | 2017-11-21 | | 公開日 | 2019-05-01 | | 最終更新日 | 2019-12-04 | | 実験手法 | X-RAY DIFFRACTION (2.4 Å) | | 主引用文献 | The dynamic conformational landscape of the protein methyltransferase SETD8.
Elife, 8, 2019
|
|
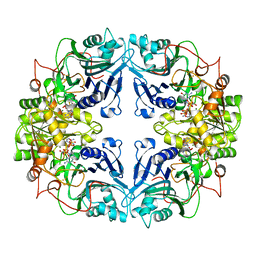
1AO0
 
 | |
6ZUE
 
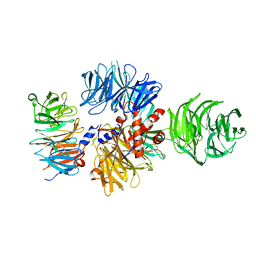 | |
6PI7
 
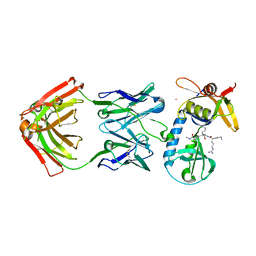 | |
5HZB
 
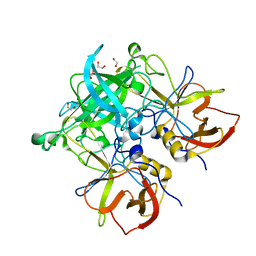 | | Crystal structure of GII.10 P domain in complex with 2-fucosyllactose (2'FL) | | 分子名称: | 1,2-ETHANEDIOL, Capsid protein, alpha-L-fucopyranose-(1-2)-beta-D-galactopyranose-(1-4)-alpha-D-glucopyranose | | 著者 | Hansman, G.S, Koromyslova, A.D, Singh, B.K.S. | | 登録日 | 2016-02-02 | | 公開日 | 2016-03-02 | | 最終更新日 | 2024-01-10 | | 実験手法 | X-RAY DIFFRACTION (1.553 Å) | | 主引用文献 | Structural Basis for Norovirus Inhibition by Human Milk Oligosaccharides.
J.Virol., 90, 2016
|
|
5HZA
 
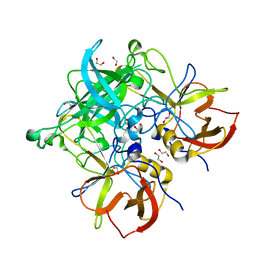 | | Crystal structure of GII.10 P domain in complex with 3-fucosyllactose (3 FL) | | 分子名称: | 1,2-ETHANEDIOL, Capsid protein, alpha-L-fucopyranose-(1-3)-[beta-D-galactopyranose-(1-4)]beta-D-glucopyranose | | 著者 | Hansman, G.S, Koromyslova, A.D, Singh, B.K. | | 登録日 | 2016-02-02 | | 公開日 | 2016-03-02 | | 最終更新日 | 2024-01-10 | | 実験手法 | X-RAY DIFFRACTION (1.35 Å) | | 主引用文献 | Structural Basis for Norovirus Inhibition by Human Milk Oligosaccharides.
J.Virol., 90, 2016
|
|
4REO
 
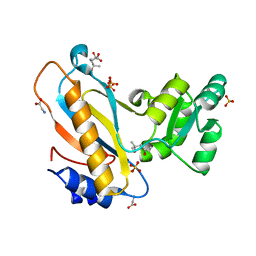 | | Mutant ribosomal protein l1 from thermus thermophilus with threonine 217 replaced by valine | | 分子名称: | (4S)-2-METHYL-2,4-PENTANEDIOL, 50S ribosomal protein L1, GLYCINE, ... | | 著者 | Gabdulkhakov, A.G, Nevskaya, N.A, Tishchenko, S.V, Nikonov, S.V. | | 登録日 | 2014-09-23 | | 公開日 | 2015-03-18 | | 最終更新日 | 2023-09-20 | | 実験手法 | X-RAY DIFFRACTION (1.35 Å) | | 主引用文献 | Protein-RNA affinity of ribosomal protein L1 mutants does not correlate with the number of intermolecular interactions.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
4IS6
 
 | | Crystal structure of HLA-DR4 bound to GP100 peptide | | 分子名称: | HLA class II histocompatibility antigen, DR alpha chain, DRB1-4 beta chain, ... | | 著者 | Li, Y. | | 登録日 | 2013-01-16 | | 公開日 | 2013-10-23 | | 最終更新日 | 2013-11-20 | | 実験手法 | X-RAY DIFFRACTION (2.5 Å) | | 主引用文献 | Structure-Based Design of Altered MHC Class II-Restricted Peptide Ligands with Heterogeneous Immunogenicity.
J.Immunol., 191, 2013
|
|
4QGB
 
 | | Crystal structure of mutant ribosomal protein G219V TthL1 | | 分子名称: | 50S ribosomal protein L1, ACETATE ION, CHLORIDE ION | | 著者 | Gabdulkhakov, A.G, Nevskaya, N.A, Nikonov, S.V. | | 登録日 | 2014-05-22 | | 公開日 | 2015-02-11 | | 最終更新日 | 2023-09-20 | | 実験手法 | X-RAY DIFFRACTION (2.6 Å) | | 主引用文献 | Protein-RNA affinity of ribosomal protein L1 mutants does not correlate with the number of intermolecular interactions.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
