7FIL
 
 | |
7FIK
 
 | | The cryo-EM structure of the CR subunit from X. laevis NPC | | 分子名称: | MGC154553 protein, MGC83295 protein, MGC83926 protein, ... | | 著者 | Shi, Y, Huang, G, Zhan, X. | | 登録日 | 2021-07-31 | | 公開日 | 2022-11-09 | | 最終更新日 | 2024-06-12 | | 実験手法 | ELECTRON MICROSCOPY (3.7 Å) | | 主引用文献 | Structure of the cytoplasmic ring of the Xenopus laevis nuclear pore complex.
Science, 376, 2022
|
|
2IAM
 
 | | Structural basis for recognition of mutant self by a tumor-specific, MHC class II-restricted TCR | | 分子名称: | 15-mer peptide from Triosephosphate isomerase, CD4+ T cell receptor E8 alpha chain, CD4+ T cell receptor E8 beta chain, ... | | 著者 | Deng, L, Langley, R.J, Mariuzza, R.A. | | 登録日 | 2006-09-08 | | 公開日 | 2007-04-03 | | 最終更新日 | 2024-11-13 | | 実験手法 | X-RAY DIFFRACTION (2.8 Å) | | 主引用文献 | Structural basis for the recognition of mutant self by a tumor-specific, MHC class II-restricted T cell receptor
Nat.Immunol., 8, 2007
|
|
2IAN
 
 | | Structural basis for recognition of mutant self by a tumor-specific, MHC class II-restricted TCR | | 分子名称: | 15-mer peptide from Triosephosphate isomerase, CD4+ T cell receptor E8 alpha chain, CD4+ T cell receptor E8 beta chain, ... | | 著者 | Deng, L, Langley, R.J, Mariuzza, R.A. | | 登録日 | 2006-09-08 | | 公開日 | 2007-04-03 | | 最終更新日 | 2024-10-30 | | 実験手法 | X-RAY DIFFRACTION (2.8 Å) | | 主引用文献 | Structural basis for the recognition of mutant self by a tumor-specific, MHC class II-restricted T cell receptor
Nat.Immunol., 8, 2007
|
|
7EPX
 
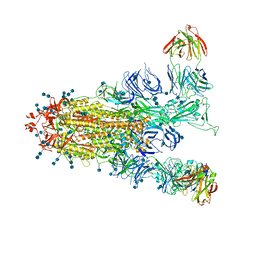 | | S protein of SARS-CoV-2 in complex with GW01 | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein, ... | | 著者 | Shen, Y.P, Zhang, Y.Y, Yan, R.H, Li, Y.N, Zhou, Q. | | 登録日 | 2021-04-28 | | 公開日 | 2022-05-18 | | 最終更新日 | 2024-11-06 | | 実験手法 | ELECTRON MICROSCOPY (3 Å) | | 主引用文献 | Novel sarbecovirus bispecific neutralizing antibodies with exceptional breadth and potency against currently circulating SARS-CoV-2 variants and sarbecoviruses.
Cell Discov, 8, 2022
|
|
2YYF
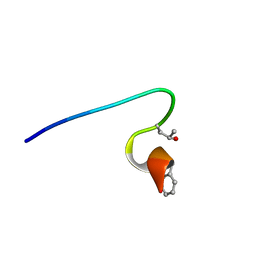 
 | | Purification and structural characterization of a D-amino acid containing conopeptide, marmophine, from Conus marmoreus | | 分子名称: | M-conotoxin mr12 | | 著者 | Huang, F, Du, W, Han, Y, Wang, C. | | 登録日 | 2007-04-29 | | 公開日 | 2008-04-08 | | 最終更新日 | 2022-03-16 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Purification and structural characterization of a D-amino acid-containing conopeptide, conomarphin, from Conus marmoreus.
Febs J., 275, 2008
|
|
7FAH
 
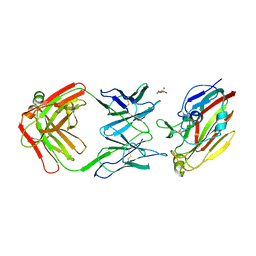 | | Immune complex of head region of CA09 HA and neutralizing antibody 12H5 | | 分子名称: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 2-acetamido-2-deoxy-beta-D-glucopyranose, Hemagglutinin, ... | | 著者 | Li, T.T, Xue, W.H, Gu, Y, Li, S.W. | | 登録日 | 2021-07-06 | | 公開日 | 2022-07-06 | | 最終更新日 | 2024-10-30 | | 実験手法 | X-RAY DIFFRACTION (3.151 Å) | | 主引用文献 | Identification of a cross-neutralizing antibody that targets the receptor binding site of H1N1 and H5N1 influenza viruses.
Nat Commun, 13, 2022
|
|
8HWT
 
 | |
8HWS
 
 | | The complex structure of Omicron BA.4 RBD with BD604, S309, and S304 | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, BD-604 Fab Heavy chain, BD-604 Fab Light chain, ... | | 著者 | He, Q.W, Xu, Z.P, Xie, Y.F. | | 登録日 | 2023-01-02 | | 公開日 | 2023-12-06 | | 最終更新日 | 2024-10-23 | | 実験手法 | ELECTRON MICROSCOPY (2.36 Å) | | 主引用文献 | An updated atlas of antibody evasion by SARS-CoV-2 Omicron sub-variants including BQ.1.1 and XBB.
Cell Rep Med, 4, 2023
|
|
8HU4
 
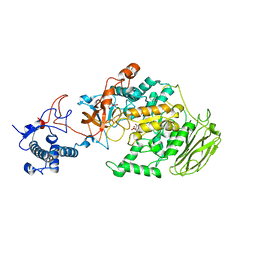 | | Limosilactobacillus reuteri N1 GtfB | | 分子名称: | CITRIC ACID, DI(HYDROXYETHYL)ETHER, SODIUM ION, ... | | 著者 | Dong, J.J, Bai, Y.X. | | 登録日 | 2022-12-22 | | 公開日 | 2023-12-27 | | 最終更新日 | 2024-03-27 | | 実験手法 | X-RAY DIFFRACTION (2.76 Å) | | 主引用文献 | Insights into the Structure-Function Relationship of GH70 GtfB alpha-Glucanotransferases from the Crystal Structure and Molecular Dynamic Simulation of a Newly Characterized Limosilactobacillus reuteri N1 GtfB Enzyme.
J.Agric.Food Chem., 72, 2024
|
|
8HWK
 
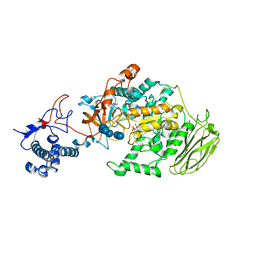 | | Limosilactobacillus reuteri N1 GtfB-maltohexaose | | 分子名称: | CITRIC ACID, DI(HYDROXYETHYL)ETHER, SODIUM ION, ... | | 著者 | Dong, J.J, Bai, Y.X. | | 登録日 | 2022-12-30 | | 公開日 | 2024-01-03 | | 最終更新日 | 2024-03-27 | | 実験手法 | X-RAY DIFFRACTION (2.9 Å) | | 主引用文献 | Insights into the Structure-Function Relationship of GH70 GtfB alpha-Glucanotransferases from the Crystal Structure and Molecular Dynamic Simulation of a Newly Characterized Limosilactobacillus reuteri N1 GtfB Enzyme.
J.Agric.Food Chem., 72, 2024
|
|
8HW3
 
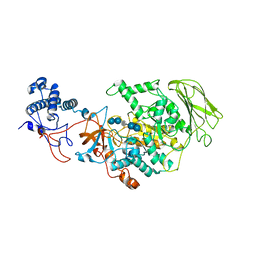 | | Limosilactobacillus reuteri N1 GtfB-acarbose | | 分子名称: | 4,6-dideoxy-4-{[(1S,4R,5S,6S)-4,5,6-trihydroxy-3-(hydroxymethyl)cyclohex-2-en-1-yl]amino}-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-beta-D-glucopyranose, GLYCEROL, SODIUM ION, ... | | 著者 | Dong, J.J, Bai, Y.X. | | 登録日 | 2022-12-28 | | 公開日 | 2024-01-03 | | 最終更新日 | 2024-03-27 | | 実験手法 | X-RAY DIFFRACTION (2.66 Å) | | 主引用文献 | Insights into the Structure-Function Relationship of GH70 GtfB alpha-Glucanotransferases from the Crystal Structure and Molecular Dynamic Simulation of a Newly Characterized Limosilactobacillus reuteri N1 GtfB Enzyme.
J.Agric.Food Chem., 72, 2024
|
|
6XEY
 
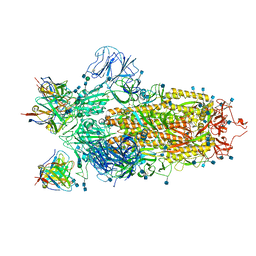 | | Cryo-EM structure of the SARS-CoV-2 spike glycoprotein bound to Fab 2-4 | | 分子名称: | 2-4 Heavy Chain, 2-4 Light Chain, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | 著者 | Rapp, M, Shapiro, L, Ho, D.D. | | 登録日 | 2020-06-14 | | 公開日 | 2020-07-22 | | 最終更新日 | 2024-11-20 | | 実験手法 | ELECTRON MICROSCOPY (3.25 Å) | | 主引用文献 | Potent neutralizing antibodies against multiple epitopes on SARS-CoV-2 spike.
Nature, 584, 2020
|
|
9JDU
 
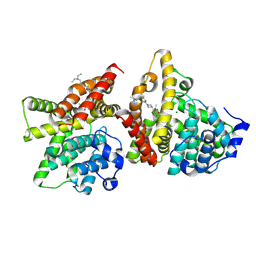 | | The crystal structure of PDE10A complexed with inhibitor 2061 | | 分子名称: | MAGNESIUM ION, ZINC ION, cAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A, ... | | 著者 | Huang, Y.-Y, Guo, L, Luo, H.-B. | | 登録日 | 2024-09-01 | | 公開日 | 2025-06-25 | | 実験手法 | X-RAY DIFFRACTION (2.3000176 Å) | | 主引用文献 | Discovery of novel azetidine-based imidazopyridines as selective and orally bioavailable inhibitors of phosphodiesterase 10A for the treatment of pulmonary arterial hypertension.
Eur.J.Med.Chem., 290, 2025
|
|
9LUG
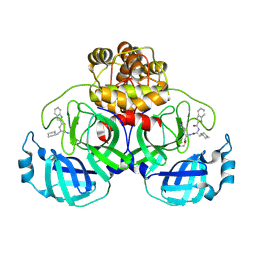 
 | |
9LUF
 
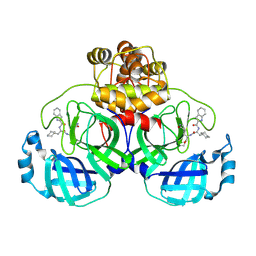 | |
5MEZ
 
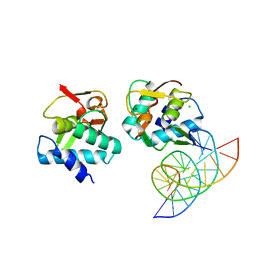 | | Crystal structure of Smad4-MH1 bound to the GGCT site. | | 分子名称: | CHLORIDE ION, DNA (5'-D(P*GP*CP*AP*GP*GP*CP*TP*AP*GP*CP*CP*TP*GP*CP*A)-3'), MH1 domain of human Smad4, ... | | 著者 | Kaczmarska, Z, Freier, R, Marquez, J.A, Macias, M.J. | | 登録日 | 2016-11-16 | | 公開日 | 2017-11-15 | | 最終更新日 | 2024-01-17 | | 実験手法 | X-RAY DIFFRACTION (2.98 Å) | | 主引用文献 | Structural basis for genome wide recognition of 5-bp GC motifs by SMAD transcription factors.
Nat Commun, 8, 2017
|
|
5MEY
 
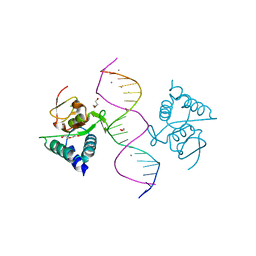 | | Crystal structure of Smad4-MH1 bound to the GGCGC site. | | 分子名称: | 1,2-ETHANEDIOL, CALCIUM ION, CHLORIDE ION, ... | | 著者 | Kaczmarska, Z, Freier, R, Marquez, J.A, Macias, M.J. | | 登録日 | 2016-11-16 | | 公開日 | 2017-11-15 | | 最終更新日 | 2024-01-17 | | 実験手法 | X-RAY DIFFRACTION (2.05 Å) | | 主引用文献 | Structural basis for genome wide recognition of 5-bp GC motifs by SMAD transcription factors.
Nat Commun, 8, 2017
|
|
5HU9
 
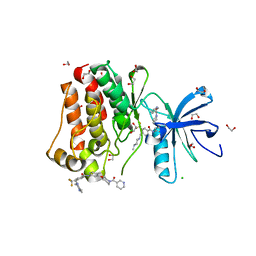 | | Crystal structure of ABL1 in complex with CHMFL-074 | | 分子名称: | 1,2-ETHANEDIOL, 4-[(4-methylpiperazin-1-yl)methyl]-N-(4-methyl-3-{[1-(pyridin-3-ylcarbonyl)piperidin-4-yl]oxy}phenyl)-3-(trifluoromethyl)benzamide, CHLORIDE ION, ... | | 著者 | Kong, L.L, Yun, C.H. | | 登録日 | 2016-01-27 | | 公開日 | 2016-07-13 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (1.529 Å) | | 主引用文献 | Discovery and characterization of a novel potent type II native and mutant BCR-ABL inhibitor (CHMFL-074) for Chronic Myeloid Leukemia (CML)
Oncotarget, 7, 2016
|
|
1GP7
 
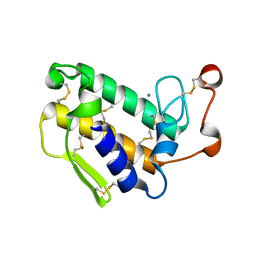 | |
8HLE
 
 | | Structure of DddY-DMSOP complex | | 分子名称: | 3-[dimethyl(oxidanyl)-$l^{4}-sulfanyl]propanoic acid, DMSP lyase DddY, ZINC ION | | 著者 | Peng, M, Li, C.Y, Zhang, Y.Z. | | 登録日 | 2022-11-30 | | 公開日 | 2023-10-04 | | 最終更新日 | 2024-11-20 | | 実験手法 | X-RAY DIFFRACTION (1.91 Å) | | 主引用文献 | DMSOP-cleaving enzymes are diverse and widely distributed in marine microorganisms.
Nat Microbiol, 8, 2023
|
|
8HLF
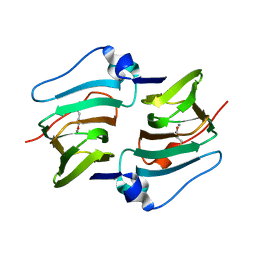 
 | | Crystal structure of DddK-DMSOP complex | | 分子名称: | 3-[dimethyl(oxidanyl)-$l^{4}-sulfanyl]propanoic acid, MANGANESE (II) ION, Novel protein with potential Cupin domain | | 著者 | Peng, M, Li, C.Y, Zhang, Y.Z. | | 登録日 | 2022-11-30 | | 公開日 | 2023-10-04 | | 最終更新日 | 2023-12-20 | | 実験手法 | X-RAY DIFFRACTION (1.62 Å) | | 主引用文献 | DMSOP-cleaving enzymes are diverse and widely distributed in marine microorganisms.
Nat Microbiol, 8, 2023
|
|
7FH7
 
 | | Friedel-Crafts alkylation enzyme CylK mutant Y37F | | 分子名称: | 5-[(2S,7R)-7-fluoranyl-2-methyl-undecyl]benzene-1,3-diol, CALCIUM ION, CHLORIDE ION, ... | | 著者 | Wang, H.Q, Wei, Z, Xiang, Z. | | 登録日 | 2021-07-29 | | 公開日 | 2022-02-16 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (1.42 Å) | | 主引用文献 | Structural Basis for the Friedel-Crafts Alkylation in Cylindrocyclophane Biosynthesis
ACS Catal., 12, 2022
|
|
7FH8
 
 | | Friedel-Crafts alkylation enzyme CylK mutant H391A | | 分子名称: | 2-[(5S,10S)-11-[3,5-bis(oxidanyl)phenyl]-10-methyl-undecan-5-yl]-5-[(2S,7R)-7-fluoranyl-2-methyl-undecyl]benzene-1,3-diol, CALCIUM ION, CHLORIDE ION, ... | | 著者 | Wang, H.Q, Wei, Z, Xiang, Z. | | 登録日 | 2021-07-29 | | 公開日 | 2022-02-16 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (1.32 Å) | | 主引用文献 | Structural Basis for the Friedel-Crafts Alkylation in Cylindrocyclophane Biosynthesis
ACS Catal., 12, 2022
|
|
7FH6
 
 | | Friedel-Crafts alkylation enzyme CylK | | 分子名称: | CALCIUM ION, CHLORIDE ION, CylK, ... | | 著者 | Liu, L, Wang, H.Q, Xiang, Z. | | 登録日 | 2021-07-29 | | 公開日 | 2022-02-16 | | 最終更新日 | 2024-05-29 | | 実験手法 | X-RAY DIFFRACTION (1.55 Å) | | 主引用文献 | Structural Basis for the Friedel-Crafts Alkylation in Cylindrocyclophane Biosynthesis
ACS Catal., 12, 2022
|
|
