7CM4
 
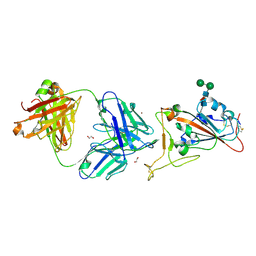 | | Crystal Structure of COVID-19 virus spike receptor-binding domain complexed with a neutralizing antibody CT-P59 | | 分子名称: | 1,2-ETHANEDIOL, IgG heavy chain, IgG light chain, ... | | 著者 | Kim, Y.G, Jeong, J.H, Bae, J.S, Lee, J. | | 登録日 | 2020-07-24 | | 公開日 | 2021-01-20 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (2.71 Å) | | 主引用文献 | A therapeutic neutralizing antibody targeting receptor binding domain of SARS-CoV-2 spike protein.
Nat Commun, 12, 2021
|
|
7F7P
 
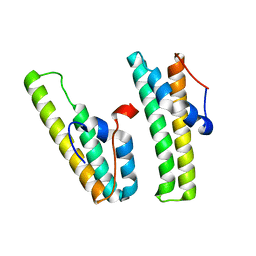 | | AcrIIC4 | | 分子名称: | anti-CRISPR protein AcrIIC4 | | 著者 | Kim, G.E, Park, H.H. | | 登録日 | 2021-06-30 | | 公開日 | 2022-05-25 | | 最終更新日 | 2024-05-29 | | 実験手法 | X-RAY DIFFRACTION (2.03 Å) | | 主引用文献 | Crystal structure of the anti-CRISPR, AcrIIC4.
Protein Sci., 30, 2021
|
|
7COE
 
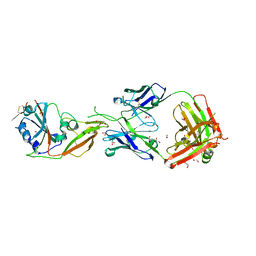 | | Crystal structure of Receptor binding domain of MERS-CoV and KNIH90-F1 Fab complex | | 分子名称: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, Heavy chain, ... | | 著者 | Lee, J.Y, Song, J.Y, Lee, H.S, Hong, E, Jang, T.H. | | 登録日 | 2020-08-04 | | 公開日 | 2021-08-04 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (2.05 Å) | | 主引用文献 | The structure of a novel antibody against the spike protein inhibits Middle East respiratory syndrome coronavirus infections.
Sci Rep, 12, 2022
|
|
7XI1
 
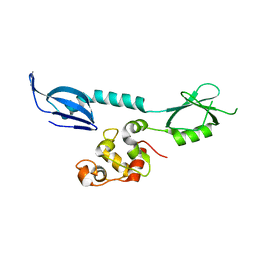 | | AcrIF 24 | | 分子名称: | anti-CRISPR protein AcrIF24 | | 著者 | Kim, G.E, Park, H.H. | | 登録日 | 2022-04-11 | | 公開日 | 2023-03-22 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (2.53 Å) | | 主引用文献 | Molecular basis of dual anti-CRISPR and auto-regulatory functions of AcrIF24.
Nucleic Acids Res., 50, 2022
|
|
4MZG
 
 | | Crystal structure of human Spindlin1 bound to histone H3K4me3 peptide | | 分子名称: | (4R)-2-METHYLPENTANE-2,4-DIOL, (4S)-2-METHYL-2,4-PENTANEDIOL, CHLORIDE ION, ... | | 著者 | Su, X, Ding, X, Li, H. | | 登録日 | 2013-09-30 | | 公開日 | 2014-03-26 | | 実験手法 | X-RAY DIFFRACTION (1.698 Å) | | 主引用文献 | Molecular basis underlying histone H3 lysine-arginine methylation pattern readout by Spin/Ssty repeats of Spindlin1
Genes Dev., 28, 2014
|
|
4MZF
 
 | | Crystal structure of human Spindlin1 bound to histone H3(K4me3-R8me2a) peptide | | 分子名称: | CHLORIDE ION, MAGNESIUM ION, Peptide from Histone H3.2, ... | | 著者 | Su, X, Ding, X, Li, H. | | 登録日 | 2013-09-30 | | 公開日 | 2014-03-26 | | 実験手法 | X-RAY DIFFRACTION (2.098 Å) | | 主引用文献 | Molecular basis underlying histone H3 lysine-arginine methylation pattern readout by Spin/Ssty repeats of Spindlin1
Genes Dev., 28, 2014
|
|
4MZH
 
 | |
5YNS
 
 | |
7BRA
 | | Bacillus subtilis IRG1 | | 分子名称: | 3-CYCLOHEXYL-1-PROPYLSULFONIC ACID, Bacillus subtilis IRG1, SULFATE ION | | 著者 | Park, H.H, Chun, H.L. | | 登録日 | 2020-03-27 | | 公開日 | 2021-02-03 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (1.785 Å) | | 主引用文献 | Enzymatic reaction mechanism of cis-aconitate decarboxylase based on the crystal structure of IRG1 from Bacillus subtilis.
Sci Rep, 10, 2020
|
|
6BPQ
 
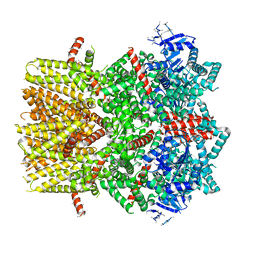 | | Structure of the cold- and menthol-sensing ion channel TRPM8 | | 分子名称: | Transient receptor potential cation channel subfamily M member 8 | | 著者 | Yin, Y, Wu, M, Zubcevic, L, Borschel, W.F, Lander, G.C, Lee, S.-Y. | | 登録日 | 2017-11-25 | | 公開日 | 2017-12-13 | | 最終更新日 | 2024-03-13 | | 実験手法 | ELECTRON MICROSCOPY (4.1 Å) | | 主引用文献 | Structure of the cold- and menthol-sensing ion channel TRPM8.
Science, 359, 2018
|
|
6BW5
 
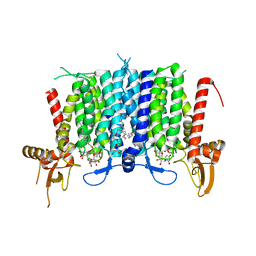 | | Human GPT (DPAGT1) in complex with tunicamycin | | 分子名称: | (1R)-2-{[(S)-{[(2S)-2,3-dihydroxypropyl]oxy}(hydroxy)phosphoryl]oxy}-1-[(hexadecanoyloxy)methyl]ethyl (9Z)-octadec-9-enoate, Tunicamycin, UDP-N-acetylglucosamine--dolichyl-phosphate N-acetylglucosaminephosphotransferase | | 著者 | Yoo, J, Kuk, A.C.Y, Mashalidis, E.H, Lee, S.-Y. | | 登録日 | 2017-12-14 | | 公開日 | 2018-02-21 | | 最終更新日 | 2023-10-04 | | 実験手法 | X-RAY DIFFRACTION (3.1 Å) | | 主引用文献 | GlcNAc-1-P-transferase-tunicamycin complex structure reveals basis for inhibition of N-glycosylation.
Nat. Struct. Mol. Biol., 25, 2018
|
|
6BW6
 
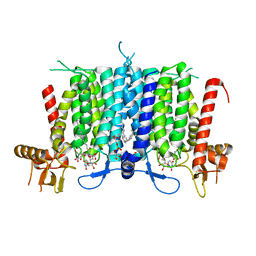 | | Human GPT (DPAGT1) H129 variant in complex with tunicamycin | | 分子名称: | (1R)-2-{[(S)-{[(2S)-2,3-dihydroxypropyl]oxy}(hydroxy)phosphoryl]oxy}-1-[(hexadecanoyloxy)methyl]ethyl (9Z)-octadec-9-enoate, Tunicamycin, UDP-N-acetylglucosamine--dolichyl-phosphate N-acetylglucosaminephosphotransferase | | 著者 | Yoo, J, Kuk, A.C.Y, Mashalidis, E.H, Lee, S.-Y. | | 登録日 | 2017-12-14 | | 公開日 | 2018-02-21 | | 最終更新日 | 2023-10-04 | | 実験手法 | X-RAY DIFFRACTION (2.95 Å) | | 主引用文献 | GlcNAc-1-P-transferase-tunicamycin complex structure reveals basis for inhibition of N-glycosylation.
Nat. Struct. Mol. Biol., 25, 2018
|
|
7BR9
 
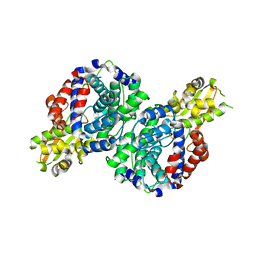 | | Crystal structure of mus musculus IRG1 | | 分子名称: | Cis-aconitate decarboxylase | | 著者 | Park, H.H, Chun, H.L. | | 登録日 | 2020-03-27 | | 公開日 | 2021-02-03 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (3.3 Å) | | 主引用文献 | The crystal structure of mouse IRG1 suggests that cis-aconitate decarboxylase has an open and closed conformation.
Plos One, 15, 2020
|
|
6K1D
 
 | |
6K1C
 
 | |
6K18
 | | Crystal structure of EXD2 exonuclease domain soaked in Mn | | 分子名称: | Exonuclease 3'-5' domain-containing protein 2, MANGANESE (II) ION | | 著者 | Park, J, Lee, C. | | 登録日 | 2019-05-10 | | 公開日 | 2019-05-22 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.303 Å) | | 主引用文献 | The structure of human EXD2 reveals a chimeric 3' to 5' exonuclease domain that discriminates substrates via metal coordination.
Nucleic Acids Res., 47, 2019
|
|
6K1A
 
 | | Crystal structure of EXD2 exonuclease domain soaked in Mn and Mg | | 分子名称: | Exonuclease 3'-5' domain-containing protein 2, MAGNESIUM ION, MANGANESE (II) ION | | 著者 | Park, J, Lee, C. | | 登録日 | 2019-05-10 | | 公開日 | 2019-05-22 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.602 Å) | | 主引用文献 | The structure of human EXD2 reveals a chimeric 3' to 5' exonuclease domain that discriminates substrates via metal coordination.
Nucleic Acids Res., 47, 2019
|
|
6K1B
 
 | |
6K19
 
 | | Crystal structure of EXD2 exonuclease domain soaked in Mg | | 分子名称: | Exonuclease 3'-5' domain-containing protein 2, MAGNESIUM ION | | 著者 | Park, J, Lee, C. | | 登録日 | 2019-05-10 | | 公開日 | 2019-05-22 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (2.202 Å) | | 主引用文献 | The structure of human EXD2 reveals a chimeric 3' to 5' exonuclease domain that discriminates substrates via metal coordination.
Nucleic Acids Res., 47, 2019
|
|
6K17
 
 | | Crystal structure of EXD2 exonuclease domain | | 分子名称: | Exonuclease 3'-5' domain-containing protein 2, SODIUM ION | | 著者 | Park, J, Lee, C. | | 登録日 | 2019-05-10 | | 公開日 | 2019-05-22 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (1.602 Å) | | 主引用文献 | The structure of human EXD2 reveals a chimeric 3' to 5' exonuclease domain that discriminates substrates via metal coordination.
Nucleic Acids Res., 47, 2019
|
|
6K1E
 
 | |
5K4B
 
 | | Structure of eukaryotic translation initiation factor 3 subunit D (eIF3d) cap binding domain from Nasonia vitripennis, Crystal form 1 | | 分子名称: | CHLORIDE ION, Eukaryotic translation initiation factor 3 subunit D | | 著者 | Kranzusch, P.J, Lee, A.S.Y, Doudna, J.A, Cate, J.H.D. | | 登録日 | 2016-05-20 | | 公開日 | 2016-07-27 | | 最終更新日 | 2024-03-06 | | 実験手法 | X-RAY DIFFRACTION (1.399 Å) | | 主引用文献 | eIF3d is an mRNA cap-binding protein that is required for specialized translation initiation.
Nature, 536, 2016
|
|
5K4D
 
 | | Structure of eukaryotic translation initiation factor 3 subunit D (eIF3d) cap binding domain from Nasonia vitripennis, Crystal form 3 | | 分子名称: | Eukaryotic translation initiation factor 3 subunit D | | 著者 | Kranzusch, P.J, Lee, A.S.Y, Doudna, J.A, Cate, J.H.D. | | 登録日 | 2016-05-20 | | 公開日 | 2016-07-27 | | 最終更新日 | 2024-03-06 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | eIF3d is an mRNA cap-binding protein that is required for specialized translation initiation.
Nature, 536, 2016
|
|
7CHR
 
 | | AcrIF9 | | 分子名称: | anti-CRISPR AcrIF9 | | 著者 | Kim, G.E, Park, H.H. | | 登録日 | 2020-07-06 | | 公開日 | 2020-12-23 | | 最終更新日 | 2024-03-27 | | 実験手法 | X-RAY DIFFRACTION (1.21 Å) | | 主引用文献 | A high-resolution (1.2 angstrom ) crystal structure of the anti-CRISPR protein AcrIF9.
Febs Open Bio, 10, 2020
|
|
7C3K
 
 | | Crystal Structure of mIRGB10 | | 分子名称: | GUANOSINE-5'-DIPHOSPHATE, Immunity-related GTPase family member b10 | | 著者 | Ha, H.J, Jeong, J.H, Kim, Y.G, Park, H.H. | | 登録日 | 2020-05-12 | | 公開日 | 2021-04-21 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (2.6 Å) | | 主引用文献 | Molecular basis of IRGB10 oligomerization and membrane association for pathogen membrane disruption.
Commun Biol, 4, 2021
|
|
