8ADG
 
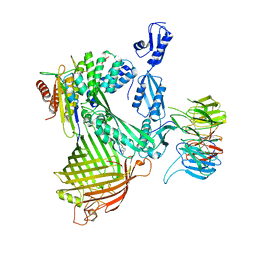 | | Cryo-EM structure of Darobactin 22 bound BAM complex | | Descriptor: | Darobactin 22, Outer membrane protein assembly factor BamA, Outer membrane protein assembly factor BamB, ... | | Authors: | Yuan, B, Marlovits, T.C. | | Deposit date: | 2022-07-08 | | Release date: | 2023-01-11 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Darobactins Exhibiting Superior Antibiotic Activity by Cryo-EM Structure Guided Biosynthetic Engineering.
Angew.Chem.Int.Ed.Engl., 62, 2023
|
|
8ADI
 
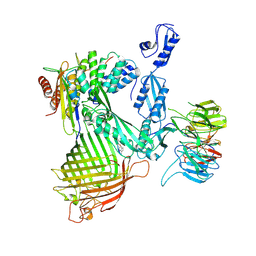 | | Cryo-EM structure of Darobactin 9 bound BAM complex | | Descriptor: | Darobactin 9, Outer membrane protein assembly factor BamA, Outer membrane protein assembly factor BamB, ... | | Authors: | Yuan, B, Marlovits, T.C. | | Deposit date: | 2022-07-08 | | Release date: | 2023-01-11 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Darobactins Exhibiting Superior Antibiotic Activity by Cryo-EM Structure Guided Biosynthetic Engineering.
Angew.Chem.Int.Ed.Engl., 62, 2023
|
|
6HAI
 
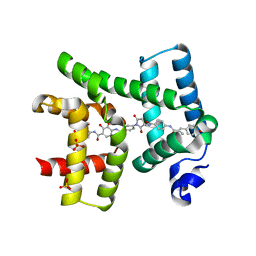 | | AlbAM131A mutant in complex with albicidin , albicidin resistance protein | | Descriptor: | 4-[[4-[[4-[(3~{S})-3-[[4-[[(~{E})-3-(4-hydroxyphenyl)-2-methyl-prop-2-enoyl]amino]phenyl]carbonylamino]-2,5-bis(oxidanylidene)pyrrolidin-1-yl]phenyl]carbonylamino]-3-methoxy-2-oxidanyl-phenyl]carbonylamino]-3-methoxy-2-oxidanyl-benzoic acid, Albicidin resistance protein, SULFATE ION | | Authors: | Koehnke, J, Sikandar, A. | | Deposit date: | 2018-08-07 | | Release date: | 2018-11-21 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Adaptation of a Bacterial Multidrug Resistance System Revealed by the Structure and Function of AlbA.
J.Am.Chem.Soc., 140, 2018
|
|
3FO0
 
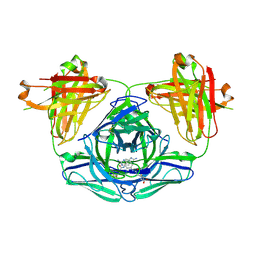
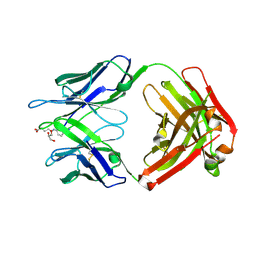 | | Crystal structure of hapten complex of catalytic elimination antibody 13G5 (wild-type) | | Descriptor: | 5-[(2-AMINO-1H-BENZIMIDAZOL-6-YL)AMINO]-5-OXOPENTANOIC ACID, Catalytic antibody Fab 13G5 IgG2b heavy chain chimera, Catalytic antibody Fab 13G5 kappa light chain chimera, ... | | Authors: | Debler, E.W, Wilson, I.A. | | Deposit date: | 2008-12-27 | | Release date: | 2009-11-10 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | An aspartate and a water molecule mediate efficient acid-base catalysis in a tailored antibody pocket.
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
2GSA
 
 | |
5HU7
 
 | |
5HQW
 
 | | Crystal structure of a trans-AT PKS dehydratase domain of C0ZGQ6 from Brevibacillus brevis | | Descriptor: | MAGNESIUM ION, Putative polyketide synthase | | Authors: | Jakob, R.P, Hauswirth, P, Dilmi, J, Herbst, D.A, Maier, T. | | Deposit date: | 2016-01-22 | | Release date: | 2017-01-25 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.39 Å) | | Cite: | Crystal Structures of Dehydratase Domains from trans AT Polyketide Biosynthetic Pathway.
To Be Published
|
|
5HWO
 
 | | MvaS in complex with 3-hydroxy-3-methylglutaryl coenzyme A | | Descriptor: | 3-HYDROXY-3-METHYLGLUTARYL-COENZYME A, GLYCEROL, Hydroxymethylglutaryl-CoA synthase, ... | | Authors: | Bock, T, Kasten, J, Blankenfeldt, W. | | Deposit date: | 2016-01-29 | | Release date: | 2016-05-11 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Crystal Structure of the HMG-CoA Synthase MvaS from the Gram-Negative Bacterium Myxococcus xanthus.
Chembiochem, 17, 2016
|
|
5HWP
 
 | | MvaS with acetylated Cys115 in complex with coenzyme A | | Descriptor: | COENZYME A, GLYCEROL, Hydroxymethylglutaryl-CoA synthase, ... | | Authors: | Bock, T, Kasten, J, Blankenfeldt, W. | | Deposit date: | 2016-01-29 | | Release date: | 2016-05-11 | | Last modified: | 2016-07-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal Structure of the HMG-CoA Synthase MvaS from the Gram-Negative Bacterium Myxococcus xanthus.
Chembiochem, 17, 2016
|
|
5HST
 
 | |
3FO2
 | |
3FO1
 
 | |
5HWQ
 
 | | MvaS in complex with acetoacetyl coenzyme A | | Descriptor: | ACETOACETYL-COENZYME A, GLYCEROL, Hydroxymethylglutaryl-CoA synthase, ... | | Authors: | Bock, T, Kasten, J, Blankenfeldt, W. | | Deposit date: | 2016-01-29 | | Release date: | 2016-05-11 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystal Structure of the HMG-CoA Synthase MvaS from the Gram-Negative Bacterium Myxococcus xanthus.
Chembiochem, 17, 2016
|
|
5HWR
 
 | | MvaS in complex with coenzyme A | | Descriptor: | COENZYME A, GLYCEROL, Hydroxymethylglutaryl-CoA synthase, ... | | Authors: | Bock, T, Kasten, J, Blankenfeldt, W. | | Deposit date: | 2016-01-29 | | Release date: | 2016-05-11 | | Last modified: | 2016-07-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystal Structure of the HMG-CoA Synthase MvaS from the Gram-Negative Bacterium Myxococcus xanthus.
Chembiochem, 17, 2016
|
|
5O28
 
 | | E. coli FolD apo | | Descriptor: | Bifunctional protein FolD | | Authors: | Koehnke, J, Sikandar, A. | | Deposit date: | 2017-05-19 | | Release date: | 2018-02-28 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | The natural product carolacton inhibits folate-dependent C1 metabolism by targeting FolD/MTHFD.
Nat Commun, 8, 2017
|
|
5N5D
 
 | | Crystal Structure of the O-Methyltransferase TomG from Streptomyces achromogenes involved in Tomaymycin synthesis in complex with SAM | | Descriptor: | (R,R)-2,3-BUTANEDIOL, GLYCEROL, Methyltransferase, ... | | Authors: | Pippel, J, Blankenfeldt, W. | | Deposit date: | 2017-02-13 | | Release date: | 2017-09-20 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Total Biosynthesis of the Pyrrolo[4,2]benzodiazepine Scaffold Tomaymycin on an In Vitro Reconstituted NRPS System.
Cell Chem Biol, 24, 2017
|
|
5O2A
 
 | | FolD Q98H | | Descriptor: | Bifunctional protein FolD | | Authors: | Koehnke, J, Sikandar, A. | | Deposit date: | 2017-05-19 | | Release date: | 2018-02-28 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The natural product carolacton inhibits folate-dependent C1 metabolism by targeting FolD/MTHFD.
Nat Commun, 8, 2017
|
|
5O22
 
 | | E. coli FolD in complex with carolacton | | Descriptor: | Bifunctional protein FolD, Carolacton | | Authors: | Koehnke, J, Sikandar, A. | | Deposit date: | 2017-05-19 | | Release date: | 2018-02-28 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The natural product carolacton inhibits folate-dependent C1 metabolism by targeting FolD/MTHFD.
Nat Commun, 8, 2017
|
|
6QRI
 
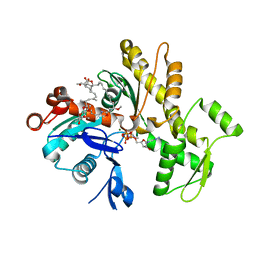 | | Structure of rabbit G-actin in complex with chivosazole A | | Descriptor: | (2~{R},3~{R},5~{S},6~{E},8~{E},10~{Z},12~{S},13~{R},16~{Z},18~{E},20~{Z},22~{E},24~{R},25~{S},26~{E},28~{Z})-13-[(2~{S},3~{S},5~{S})-3,5-bis(oxidanyl)hexan-2-yl]-25-[(2~{R},3~{R},4~{S},5~{R},6~{R})-3,4-dimethoxy-6-methyl-5-oxidanyl-oxan-2-yl]oxy-3-methoxy-2,12,22,24-tetramethyl-5-oxidanyl-14,32-dioxa-33-azabicyclo[28.2.1]tritriaconta-1(33),6,8,10,16,18,20,22,26,28,30-undecaen-15-one, ADENOSINE-5'-TRIPHOSPHATE, Actin, ... | | Authors: | Schneider, S, Wang, S, Zahler, S. | | Deposit date: | 2019-02-19 | | Release date: | 2019-07-03 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Chivosazole A Modulates Protein-Protein Interactions of Actin.
J.Nat.Prod., 82, 2019
|
|
6RYO
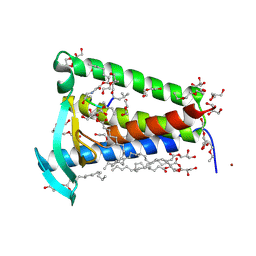 
 | | Bacterial membrane enzyme structure by the in meso method at 1.9 A resolution | | Descriptor: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, 2-(2-(2-(2-(2-(2-ETHOXYETHOXY)ETHOXY)ETHOXY)ETHOXY)ETHOXY)ETHANOL, GLYCEROL, ... | | Authors: | Huang, C.Y, Olatunji, S, Bailey, J, Yu, X, Olieric, V, Wang, M, Caffrey, M. | | Deposit date: | 2019-06-11 | | Release date: | 2020-01-15 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.924 Å) | | Cite: | Structures of lipoprotein signal peptidase II from Staphylococcus aureus complexed with antibiotics globomycin and myxovirescin.
Nat Commun, 11, 2020
|
|
6RYP
 
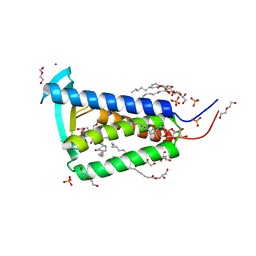 | | Bacterial membrane enzyme structure by the in meso method at 2.3 A resolution | | Descriptor: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, DI(HYDROXYETHYL)ETHER, Lipoprotein signal peptidase, ... | | Authors: | Huang, C.Y, Olatunji, S, Bailey, J, Yu, X, Olieric, V, Wang, M, Caffrey, M. | | Deposit date: | 2019-06-11 | | Release date: | 2020-01-15 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structures of lipoprotein signal peptidase II from Staphylococcus aureus complexed with antibiotics globomycin and myxovirescin.
Nat Commun, 11, 2020
|
|
6RX4
 
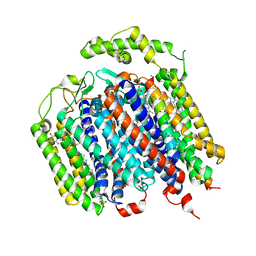 | | THE STRUCTURE OF BD OXIDASE FROM ESCHERICHIA COLI | | Descriptor: | 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine, CIS-HEME D HYDROXYCHLORIN GAMMA-SPIROLACTONE, Cytochrome bd-I ubiquinol oxidase subunit 1, ... | | Authors: | Rasmussen, T, Boettcher, B, Thesseling, A, Friedrich, T. | | Deposit date: | 2019-06-07 | | Release date: | 2019-11-20 | | Last modified: | 2024-05-22 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Homologous bd oxidases share the same architecture but differ in mechanism.
Nat Commun, 10, 2019
|
|
3GSB
 
 | |
5AA6
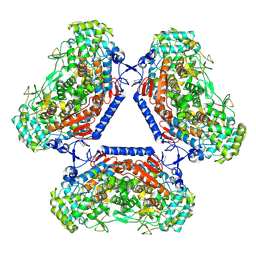 
 | | Homohexameric Structure of the second Vanadate-Dependent Bromoperoxidase (AnII) from Ascophyllum nodosum | | Descriptor: | VANADATE ION, VANADIUM-DEPENDENT BROMOPEROXIDASE 2 | | Authors: | Radlow, M, Jeudy, A, Dabin, J, Delage, L, Leblanc, C, Hartung, J, Czjzek, M. | | Deposit date: | 2015-07-23 | | Release date: | 2016-08-03 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Homohexameric Structure of the Second Vanadate Dependant Bromoperoxidase from Ascophyllum Nodosum
To be Published
|
|
1BT4
 
 | |
