7VG5
 
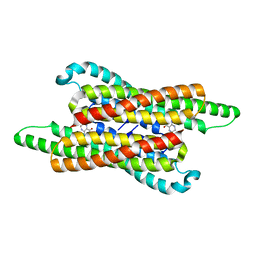 | | 10,5-methenyltetrahydrofolate cyclohydrolase from Methylobacterium extorquens AM1 with tetrahydrofolate | | Descriptor: | (6S)-5,6,7,8-TETRAHYDROFOLATE, Methenyltetrahydrofolate cyclohydrolase | | Authors: | Kim, S, Lee, S, Kim, I.-K, Seo, H, Kim, K.-J. | | Deposit date: | 2021-09-14 | | Release date: | 2022-07-06 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structural insight into a molecular mechanism of methenyltetrahydrofolate cyclohydrolase from Methylobacterium extorquens AM1.
Int.J.Biol.Macromol., 202, 2022
|
|
2QMQ
 
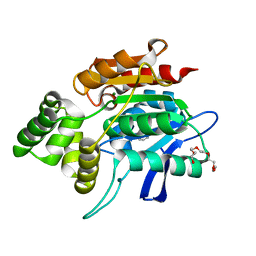 | |
4REP
 
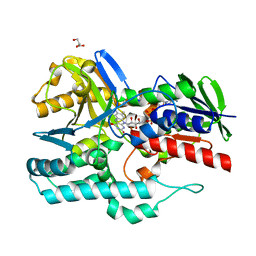 | | Crystal Structure of gamma-carotenoid desaturase | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, GLYCEROL, Gamma-carotene desaturase | | Authors: | Ahn, J.-W, Kim, E.-J, Kim, S, Kim, K.-J. | | Deposit date: | 2014-09-23 | | Release date: | 2015-07-15 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Crystal structure of 1'-OH-carotenoid 3,4-desaturase from Nonlabens dokdonensis DSW-6.
Enzyme.Microb.Technol., 77, 2015
|
|
2XVJ
 
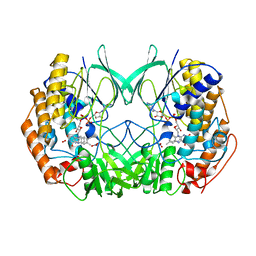 | |
2XVE
 
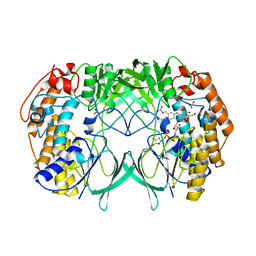 | | Crystal structure of bacterial flavin-containing monooxygenase | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, FLAVIN-CONTAINING MONOOXYGENASE, GLYCEROL, ... | | Authors: | Cho, H.J, Kang, B.S. | | Deposit date: | 2010-10-26 | | Release date: | 2011-05-04 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Structural and Functional Analysis of Bacterial Flavin-Containing Monooxygenase Reveals its Ping-Pong-Type Reaction Mechanism.
J.Struct.Biol., 175, 2011
|
|
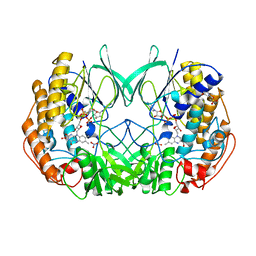
2XVI
 
 | |
6KD7
 
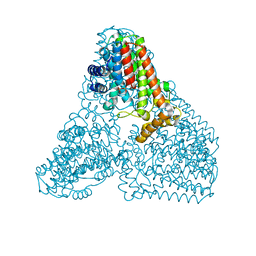 | | Crystal structure of geranylgeranyl pyrophosphate synthase | | Descriptor: | GLYCEROL, MAGNESIUM ION, PYROPHOSPHATE, ... | | Authors: | Kim, S, Kim, K.-J. | | Deposit date: | 2019-07-01 | | Release date: | 2019-09-11 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of geranylgeranyl pyrophosphate synthase (crtE) from Nonlabens dokdonensis DSW-6.
Biochem.Biophys.Res.Commun., 518, 2019
|
|
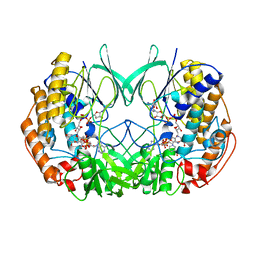
2XVH
 
 | |
2XVF
 
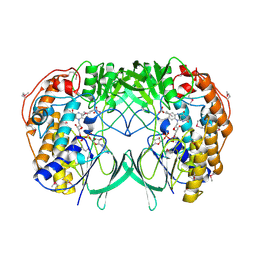 | | Crystal structure of bacterial flavin-containing monooxygenase | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, FLAVIN-CONTAINING MONOOXYGENASE, GLYCEROL, ... | | Authors: | Cho, H.J, Kang, B.S. | | Deposit date: | 2010-10-26 | | Release date: | 2011-05-04 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural and Functional Analysis of Bacterial Flavin-Containing Monooxygenase Reveals its Ping-Pong-Type Reaction Mechanism.
J.Struct.Biol., 175, 2011
|
|
8I70
 
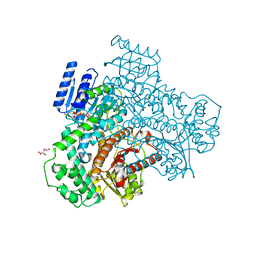 | |
8I6Z
 
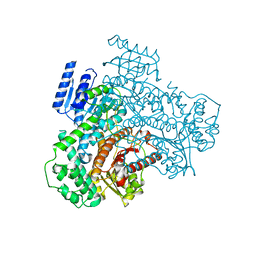 | |
8IDU
 
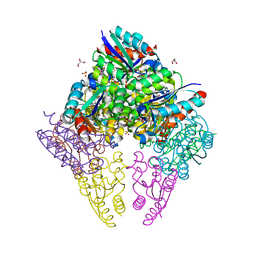 | | Crystal structure of substrate bound-form dehydroquinate dehydratase from Corynebacterium glutamicum | | Descriptor: | 1,3,4-TRIHYDROXY-5-OXO-CYCLOHEXANECARBOXYLIC ACID, 3-dehydroquinate dehydratase, GLYCEROL, ... | | Authors: | Lee, C.H, Kim, S, Kim, K.-J. | | Deposit date: | 2023-02-14 | | Release date: | 2023-12-27 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural and Biochemical Analysis of 3-Dehydroquinate Dehydratase from Corynebacterium glutamicum .
J Microbiol Biotechnol., 33, 2023
|
|
8IDR
 
 | |
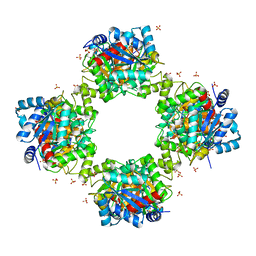
7VG9
 | |
8HKB
 
 | |
8HKA
 
 | |
8HK9
 
 | |
5E3Q
 
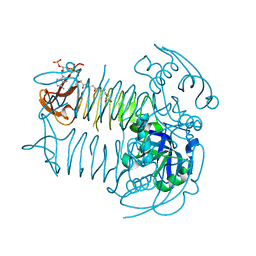 | | Crystal structure of DapD in complex with succinyl-CoA from Corynebacterium glutamicum | | Descriptor: | 2,3,4,5-tetrahydropyridine-2,6-dicarboxylate N-succinyltransferase, SUCCINYL-COENZYME A | | Authors: | Sagong, H.-Y, Kim, K.-J. | | Deposit date: | 2015-10-03 | | Release date: | 2015-11-04 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal Structure and Biochemical Characterization of Tetrahydrodipicolinate N-Succinyltransferase from Corynebacterium glutamicum.
J.Agric.Food Chem., 63, 2015
|
|
5E3R
 
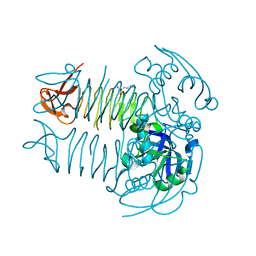 | | Crystal structure of DapD in complex with 2-aminopimelate from Corynebacterium glutamicum | | Descriptor: | (2S)-2-aminoheptanedioic acid, 2,3,4,5-tetrahydropyridine-2,6-dicarboxylate N-succinyltransferase | | Authors: | Sagong, H.-Y, Kim, K.-J. | | Deposit date: | 2015-10-03 | | Release date: | 2015-11-04 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Crystal Structure and Biochemical Characterization of Tetrahydrodipicolinate N-Succinyltransferase from Corynebacterium glutamicum.
J.Agric.Food Chem., 63, 2015
|
|
8I4Q
 
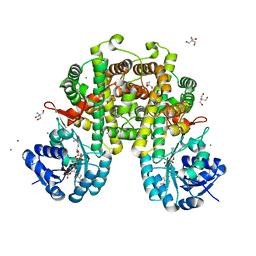 | |
8I4N
 
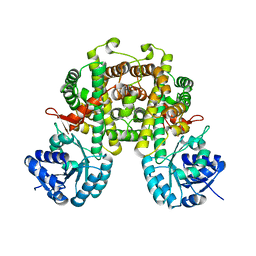 | |
7D02
 
 | | Lysozyme structure SASE2 from SASE mode | | Descriptor: | Lysozyme C | | Authors: | Kang, H.S, Lee, S.J. | | Deposit date: | 2020-09-09 | | Release date: | 2021-03-31 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | High-brightness self-seeded X-ray free-electron laser covering the 3.5 keV to 14.6 keV range
Nat Photonics, 2021
|
|
7D04
 
 | | Lysozyme structure SS3 from SS mode | | Descriptor: | Lysozyme C | | Authors: | Kang, H.S, Lee, S.J. | | Deposit date: | 2020-09-09 | | Release date: | 2021-03-31 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | High-brightness self-seeded X-ray free-electron laser covering the 3.5 keV to 14.6 keV range
Nat Photonics, 2021
|
|
7D05
 
 | | Lysozyme structure SASE3 from SASE mode | | Descriptor: | Lysozyme C | | Authors: | Kang, H.S, Lee, S.J. | | Deposit date: | 2020-09-09 | | Release date: | 2021-03-31 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | High-brightness self-seeded X-ray free-electron laser covering the 3.5 keV to 14.6 keV range
Nat Photonics, 2021
|
|
5E3P
 
 | | Crystal structure of DapD from Corynebacterium glutamicum | | Descriptor: | 2,3,4,5-tetrahydropyridine-2,6-dicarboxylate N-succinyltransferase, SULFATE ION, TETRAETHYLENE GLYCOL | | Authors: | Sagong, H.-Y, Kim, K.-J. | | Deposit date: | 2015-10-03 | | Release date: | 2015-11-04 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Crystal Structure and Biochemical Characterization of Tetrahydrodipicolinate N-Succinyltransferase from Corynebacterium glutamicum.
J.Agric.Food Chem., 63, 2015
|
|
