2XU6
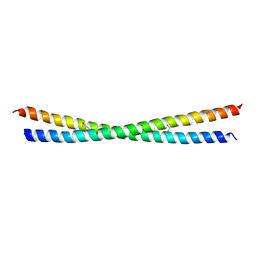 
 | | MDV1 coiled coil domain | | Descriptor: | MDV1 COILED COIL | | Authors: | Koirala, S, Bui, H.T, Schubert, H.L, Eckert, D.M, Hill, C.P, Kay, M.S, Shaw, J.M. | | Deposit date: | 2010-10-14 | | Release date: | 2010-10-27 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Molecular Architecture of a Dynamin Adaptor: Implications for Assembly of Mitochondrial Fission Complexes
J.Cell Biol., 191, 2010
|
|
1NV8
 
 | | N5-glutamine methyltransferase, HemK | | Descriptor: | N5-METHYLGLUTAMINE, S-ADENOSYLMETHIONINE, hemK protein | | Authors: | Schubert, H.L, Phillips, J.D, Hill, C.P. | | Deposit date: | 2003-02-02 | | Release date: | 2003-05-27 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structures along the Catalytic Pathway of PrmC/HemK, an N(5)-Glutamine
AdoMet-Dependent Methyltransferase
Biochemistry, 42, 2003
|
|
1NV9
 
 | | HemK, apo structure | | Descriptor: | S-ADENOSYL-L-HOMOCYSTEINE, hemK protein | | Authors: | Schubert, H.L, Phillips, J.D, Hill, C.P. | | Deposit date: | 2003-02-02 | | Release date: | 2003-05-27 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.356 Å) | | Cite: | Structures along the Catalytic Pathway of PrmC/HemK, an N(5)-Glutamine
AdoMet-Dependent Methyltransferase
Biochemistry, 42, 2003
|
|
1QB7
 
 | | CRYSTAL STRUCTURES OF ADENINE PHOSPHORIBOSYLTRANSFERASE FROM LEISHMANIA DONOVANI. | | Descriptor: | ADENINE, ADENINE PHOSPHORIBOSYLTRANSFERASE, CITRIC ACID, ... | | Authors: | Phillips, C.L, Ullman, B, Brennan, R.G, Hill, C.P. | | Deposit date: | 1999-04-30 | | Release date: | 1999-07-21 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystal structures of adenine phosphoribosyltransferase from Leishmania donovani.
EMBO J., 18, 1999
|
|
1QCC
 
 | | CRYSTAL STRUCTURES OF ADENINE PHOSPHORIBOSYLTRANSFERASE FROM LEISHMANIA DONOVANI | | Descriptor: | ADENINE PHOSPHORIBOSYLTRANSFERASE, CITRIC ACID | | Authors: | Phillips, C.L, Ullman, B, Brennan, R.G, Hill, C.P. | | Deposit date: | 1999-05-01 | | Release date: | 1999-07-21 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Crystal structures of adenine phosphoribosyltransferase from Leishmania donovani.
EMBO J., 18, 1999
|
|
1QCD
 
 | | CRYSTAL STRUCTURES OF ADENINE PHOSPHORIBOSYLTRANSFERASE FROM LEISHMANIA DONOVANI | | Descriptor: | ADENINE PHOSPHORIBOSYLTRANSFERASE, SULFATE ION | | Authors: | Phillips, C.L, Ullman, B, Brennan, R.G, Hill, C.P. | | Deposit date: | 1999-05-01 | | Release date: | 1999-07-21 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.48 Å) | | Cite: | Crystal structures of adenine phosphoribosyltransferase from Leishmania donovani.
EMBO J., 18, 1999
|
|
1R3Y
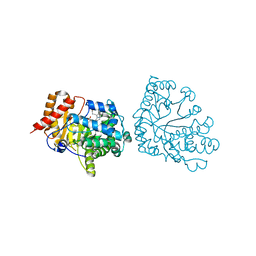 
 | | Uroporphyrinogen Decarboxylase in complex with coproporphyrinogen-III | | Descriptor: | COPROPORPHYRINOGEN III, Uroporphyrinogen Decarboxylase | | Authors: | Phillips, J.D, Whitby, F.G, Kushner, J.P, Hill, C.P. | | Deposit date: | 2003-10-03 | | Release date: | 2003-12-09 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.755 Å) | | Cite: | Structural basis for tetrapyrrole coordination by uroporphyrinogen decarboxylase
Embo J., 22, 2003
|
|
3EIE
 
 | | Crystal Structure of S.cerevisiae Vps4 in the SO4-bound state | | Descriptor: | SULFATE ION, Vacuolar protein sorting-associated protein 4 | | Authors: | Gonciarz, M.D, Whitby, F.G, Eckert, D.M, Kieffer, C, Heroux, A, Sundquist, W.I, Hill, C.P. | | Deposit date: | 2008-09-15 | | Release date: | 2008-09-30 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Biochemical and structural studies of yeast vps4 oligomerization.
J.Mol.Biol., 384, 2008
|
|
3FRT
 
 | |
3FRR
 
 | |
3FRS
 
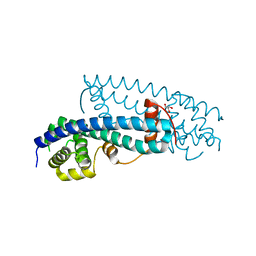 | | Structure of human IST1(NTD) (residues 1-189)(p43212) | | Descriptor: | GLYCEROL, Uncharacterized protein KIAA0174 | | Authors: | Schubert, H.L, Hill, C.P, Bajorek, M, Sundquist, W.I. | | Deposit date: | 2009-01-08 | | Release date: | 2009-06-30 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.61 Å) | | Cite: | Structural basis for ESCRT-III protein autoinhibition.
Nat.Struct.Mol.Biol., 16, 2009
|
|
3EIH
 
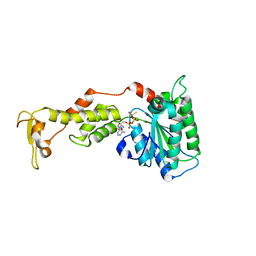 | | Crystal structure of S.cerevisiae Vps4 in the presence of ATPgammaS | | Descriptor: | 1,2-ETHANEDIOL, MAGNESIUM ION, PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER, ... | | Authors: | Gonciarz, M.D, Whitby, F.G, Eckert, D.M, Kieffer, C, Heroux, A, Sundquist, W.I, Hill, C.P. | | Deposit date: | 2008-09-15 | | Release date: | 2008-09-30 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (3.25 Å) | | Cite: | Biochemical and structural studies of yeast vps4 oligomerization.
J.Mol.Biol., 384, 2008
|
|
1R3V
 
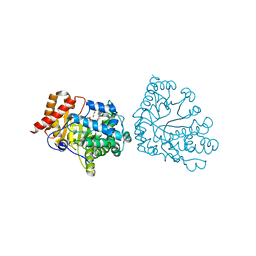 | | Uroporphyrinogen Decarboxylase single mutant D86E in complex with coproporphyrinogen-I | | Descriptor: | BETA-MERCAPTOETHANOL, COPROPORPHYRINOGEN I, Uroporphyrinogen Decarboxylase | | Authors: | Phillips, J.D, Whitby, F.G, Kushner, J.P, Hill, C.P. | | Deposit date: | 2003-10-03 | | Release date: | 2003-12-09 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural basis for tetrapyrrole coordination by uroporphyrinogen decarboxylase
Embo J., 22, 2003
|
|
1R3W
 
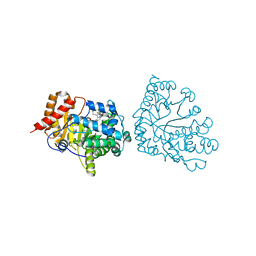 | | Uroporphyrinogen Decarboxylase Y164F mutant in complex with coproporphyrinogen-III | | Descriptor: | COPROPORPHYRINOGEN III, Uroporphyrinogen Decarboxylase | | Authors: | Phillips, J.D, Whitby, F.G, Kushner, J.P, Hill, C.P. | | Deposit date: | 2003-10-03 | | Release date: | 2003-12-09 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural basis for tetrapyrrole coordination by uroporphyrinogen decarboxylase
Embo J., 22, 2003
|
|
1R3S
 
 | | Uroporphyrinogen Decarboxylase single mutant D86G in complex with coproporphyrinogen-I | | Descriptor: | COPROPORPHYRINOGEN I, Uroporphyrinogen Decarboxylase | | Authors: | Phillips, J.D, Whitby, F.G, Kushner, J.P, Hill, C.P. | | Deposit date: | 2003-10-03 | | Release date: | 2003-12-09 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structural basis for tetrapyrrole coordination by uroporphyrinogen decarboxylase
Embo J., 22, 2003
|
|
1R3Q
 
 | | Uroporphyrinogen Decarboxylase in complex with coproporphyrinogen-I | | Descriptor: | CARBON DIOXIDE, COPROPORPHYRINOGEN I, Uroporphyrinogen Decarboxylase | | Authors: | Phillips, J.D, Whitby, F.G, Kushner, J.P, Hill, C.P. | | Deposit date: | 2003-10-03 | | Release date: | 2003-12-09 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural basis for tetrapyrrole coordination by uroporphyrinogen decarboxylase
Embo J., 22, 2003
|
|
1R3R
 
 | |
1R3T
 
 | | Uroporphyrinogen Decarboxylase single mutant D86G in complex with coproporphyrinogen-III | | Descriptor: | COPROPORPHYRINOGEN III, Uroporphyrinogen Decarboxylase | | Authors: | Phillips, J.D, Whitby, F.G, Kushner, J.P, Hill, C.P. | | Deposit date: | 2003-10-03 | | Release date: | 2003-12-09 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural basis for tetrapyrrole coordination by uroporphyrinogen decarboxylase
Embo J., 22, 2003
|
|
1TLB
 
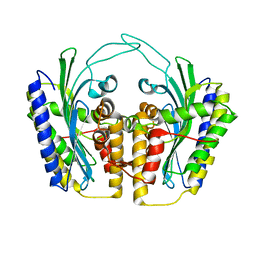 | | Yeast coproporphyrinogen oxidase | | Descriptor: | Coproporphyrinogen III oxidase, SULFATE ION | | Authors: | Phillip, J.D, Whitby, F.G, Warby, C.A, Labbe, P, Yang, C, Pflugrath, J.W, Ferrara, J.D, Robinson, H, Kushner, J.P, Hill, C.P. | | Deposit date: | 2004-06-09 | | Release date: | 2004-07-20 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of the oxygen-dependent coproporphyrinogen oxidase (Hem13p) of Saccharomyces cerevisiae
J.Biol.Chem., 279, 2004
|
|
1TK1
 
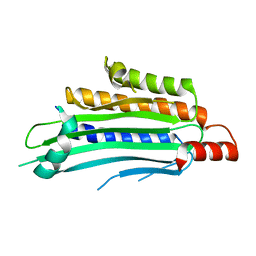 | | YEAST OXYGEN-DEPENDENT COPROPORPHYRINOGEN OXIDASE | | Descriptor: | Coproporphyrinogen III oxidase | | Authors: | Phillips, J.D, Whitby, F.G, Warby, C.A, Labbe, P, Yang, C, Pflugrath, J.W, Ferrara, J.D, Robinson, H, Kushner, J.P, Hill, C.P. | | Deposit date: | 2004-06-08 | | Release date: | 2004-07-20 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal Structure of the Oxygen-dependant Coproporphyrinogen Oxidase (Hem13p) of Saccharomyces cerevisiae
J.Biol.Chem., 279, 2004
|
|
1TKL
 
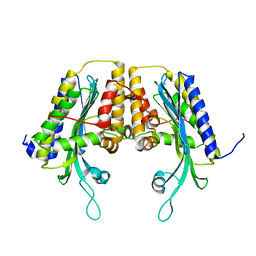 | | Yeast Oxygen-Dependent Coproporphyrinogen Oxidase | | Descriptor: | Coproporphyrinogen III oxidase | | Authors: | Phillips, J.D, Whitby, F.G, Warby, C.A, Labbe, P, Yang, C, Pflugrath, J.W, Ferrara, J.D, Robinson, H, Kushner, J.P, Hill, C.P. | | Deposit date: | 2004-06-08 | | Release date: | 2004-07-20 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal Structure of the Oxygen-dependant Coproporphyrinogen Oxidase (Hem13p) of Saccharomyces cerevisiae
J.Biol.Chem., 279, 2004
|
|
1URO
 
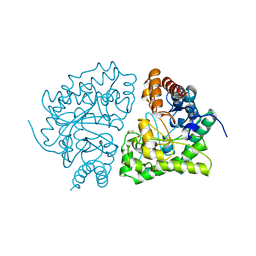 | | UROPORPHYRINOGEN DECARBOXYLASE | | Descriptor: | BETA-MERCAPTOETHANOL, PROTEIN (UROPORPHYRINOGEN DECARBOXYLASE) | | Authors: | Whitby, F.G, Phillips, J.D, Kushner, J.P, Hill, C.P. | | Deposit date: | 1998-08-21 | | Release date: | 1998-08-26 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of human uroporphyrinogen decarboxylase.
EMBO J., 17, 1998
|
|
5UIE
 
 | | Vps4-Vta1 complex | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, BERYLLIUM TRIFLUORIDE ION, DOA4-independent degradation protein 4, ... | | Authors: | Monroe, N, Shen, P, Han, H, Sundquist, W.I, Hill, C.P. | | Deposit date: | 2017-01-13 | | Release date: | 2017-04-12 | | Last modified: | 2020-01-01 | | Method: | ELECTRON MICROSCOPY (5.7 Å) | | Cite: | Structural basis of protein translocation by the Vps4-Vta1 AAA ATPase.
Elife, 6, 2017
|
|
5V1Z
 
 | | Crystal structure of the RPN13 PRU-RPN2 (932-953)-ubiquitin complex | | Descriptor: | 26S proteasome non-ATPase regulatory subunit 1, Proteasomal ubiquitin receptor ADRM1, Ubiquitin | | Authors: | Hemmis, C.W, VanderLinden, R.T, Yao, T, Robinson, H, Hill, C.P. | | Deposit date: | 2017-03-02 | | Release date: | 2017-05-03 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure and energetics of pairwise interactions between proteasome subunits RPN2, RPN13, and ubiquitin clarify a substrate recruitment mechanism.
J. Biol. Chem., 292, 2017
|
|
5VKO
 
 | | SPT6 tSH2-RPB1 1468-1500 pT1471, pS1493 | | Descriptor: | DNA-directed RNA polymerase II subunit RPB1, ISOPROPYL ALCOHOL, Transcription elongation factor SPT6 | | Authors: | Sdano, M.A, Whitby, F.G, Hill, C.P. | | Deposit date: | 2017-04-21 | | Release date: | 2017-09-20 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | A novel SH2 recognition mechanism recruits Spt6 to the doubly phosphorylated RNA polymerase II linker at sites of transcription.
Elife, 6, 2017
|
|
