1FXT
 
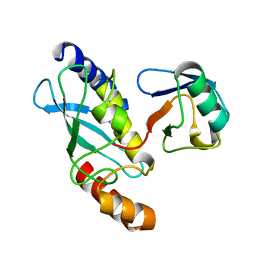 | | STRUCTURE OF A CONJUGATING ENZYME-UBIQUITIN THIOLESTER COMPLEX | | Descriptor: | UBIQUITIN, UBIQUITIN-CONJUGATING ENZYME E2-24 KDA | | Authors: | Hamilton, K.S, Shaw, G.S, Williams, R.S, Huzil, J.T, McKenna, S, Ptak, C, Glover, M, Ellison, M.J. | | Deposit date: | 2000-09-26 | | Release date: | 2001-10-10 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structure of a conjugating enzyme-ubiquitin thiolester intermediate reveals a novel role for the ubiquitin tail.
Structure, 9, 2001
|
|
1FZY
 
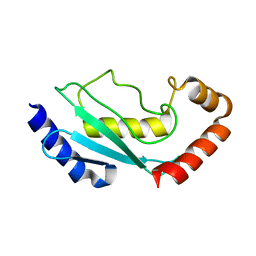 | |
7L34
 
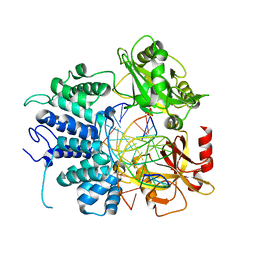 | | Human DNA Ligase 1 - R641L nicked DNA complex | | Descriptor: | ADENOSINE MONOPHOSPHATE, DNA (5'-D(*GP*CP*TP*GP*AP*TP*GP*CP*GP*TP*C)-3'), DNA (5'-D(*GP*TP*CP*CP*GP*AP*CP*GP*AP*CP*GP*CP*AP*TP*CP*AP*GP*C)-3'), ... | | Authors: | Tumbale, P.P, Williams, R.S, Schellenberg, M.S. | | Deposit date: | 2020-12-17 | | Release date: | 2021-01-13 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.901 Å) | | Cite: | LIG1 syndrome mutations remodel a cooperative network of ligand binding interactions to compromise ligation efficiency.
Nucleic Acids Res., 49, 2021
|
|
7L35
 
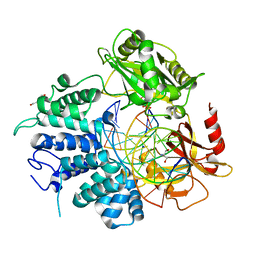 | | Human DNA Ligase 1 - R771W nicked DNA complex | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, ADENOSINE MONOPHOSPHATE, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Tumbale, P.P, Williams, R.S, Schellenberg, M.S. | | Deposit date: | 2020-12-17 | | Release date: | 2021-01-13 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | LIG1 syndrome mutations remodel a cooperative network of ligand binding interactions to compromise ligation efficiency.
Nucleic Acids Res., 49, 2021
|
|
7JL5
 
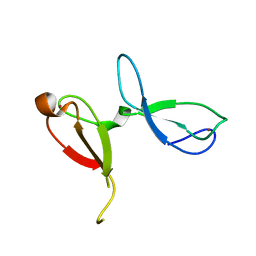 | | Crystal structure of human NEIL3 tandem zinc finger GRF domains | | Descriptor: | Endonuclease 8-like 3, ZINC ION | | Authors: | Rodriguez, A.A, Wojtaszek, J.L, Williams, R.S, Eichman, B.F. | | Deposit date: | 2020-07-29 | | Release date: | 2020-09-09 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | An autoinhibitory role for the GRF zinc finger domain of DNA glycosylase NEIL3.
J.Biol.Chem., 295, 2020
|
|
3PXE
 
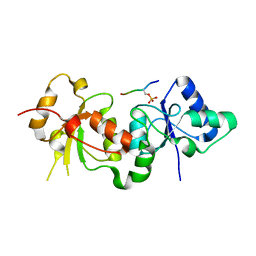 | |
3PXB
 
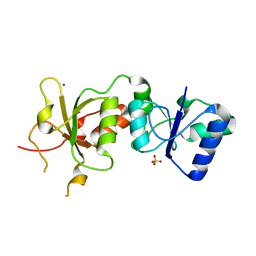 | |
3PXC
 
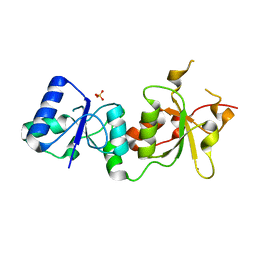 | |
3PXD
 
 | |
3PXA
 
 | |
3U7G
 
 | | Crystal structure of mPNKP catalytic fragment (D170A) bound to single-stranded DNA (TCCTAp) | | Descriptor: | Bifunctional polynucleotide phosphatase/kinase, DNA, GLYCEROL, ... | | Authors: | Coquelle, N, Havali, Z, Bernstein, N, Green, R, Glover, J.N.M. | | Deposit date: | 2011-10-13 | | Release date: | 2011-12-14 | | Last modified: | 2017-11-08 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural basis for the phosphatase activity of polynucleotide kinase/phosphatase on single- and double-stranded DNA substrates.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3U7F
 
 | | Crystal structure of mPNKP catalytic fragment (D170A) bound to single-stranded DNA (TCCTCp) | | Descriptor: | Bifunctional polynucleotide phosphatase/kinase, DNA, GLYCEROL, ... | | Authors: | Coquelle, N, Havali, Z, Bernstein, N, Green, R, Glover, J.N.M. | | Deposit date: | 2011-10-13 | | Release date: | 2011-12-14 | | Last modified: | 2017-11-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural basis for the phosphatase activity of polynucleotide kinase/phosphatase on single- and double-stranded DNA substrates.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3U7E
 
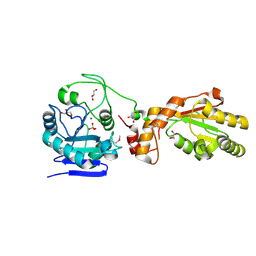 | | Crystal structure of mPNKP catalytic fragment (D170A) | | Descriptor: | Bifunctional polynucleotide phosphatase/kinase, GLYCEROL, MAGNESIUM ION, ... | | Authors: | Coquelle, N, Havali, Z, Bernstein, N, Green, R, Glover, J.N.M. | | Deposit date: | 2011-10-13 | | Release date: | 2011-12-14 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural basis for the phosphatase activity of polynucleotide kinase/phosphatase on single- and double-stranded DNA substrates.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3U7H
 
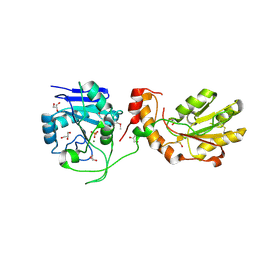 | | Crystal structure of mPNKP catalytic fragment (D170A) bound to single-stranded DNA (TCCTTp) | | Descriptor: | Bifunctional polynucleotide phosphatase/kinase, DNA, GLYCEROL, ... | | Authors: | Coquelle, N, Havali, Z, Bernstein, N, Green, R, Glover, J.N.M. | | Deposit date: | 2011-10-13 | | Release date: | 2011-12-14 | | Last modified: | 2017-11-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural basis for the phosphatase activity of polynucleotide kinase/phosphatase on single- and double-stranded DNA substrates.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
6XSR
 
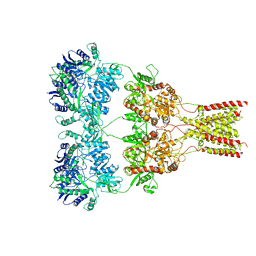 | |
