1V2N
 
 | | Potent factor XA inhibitor in complex with bovine trypsin variant X(99/175/190)bT | | Descriptor: | 2,7-BIS-(4-AMIDINOBENZYLIDENE)-CYCLOHEPTAN-1-ONE, CALCIUM ION, Trypsin | | Authors: | Rauh, D, Klebe, G, Stubbs, M.T. | | Deposit date: | 2003-10-17 | | Release date: | 2004-06-01 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Understanding protein-ligand interactions: the price of protein flexibility
J.Mol.Biol., 335, 2004
|
|
1J17
 
 | |
1J14
 
 | | BENZAMIDINE IN COMPLEX WITH RAT TRYPSIN MUTANT X99RT | | Descriptor: | BENZAMIDINE, CALCIUM ION, SULFATE ION, ... | | Authors: | Stubbs, M.T. | | Deposit date: | 2002-11-30 | | Release date: | 2002-12-23 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Reconstructing the Binding Site of Factor Xa in Trypsin Reveals Ligand-induced Structural Plasticity
J.MOL.BIOL., 325, 2003
|
|
1J16
 
 | | BENZAMIDINE IN COMPLEX WITH RAT TRYPSIN MUTANT X99/175/190RT | | Descriptor: | BENZAMIDINE, CALCIUM ION, SULFATE ION, ... | | Authors: | Stubbs, M.T. | | Deposit date: | 2002-11-30 | | Release date: | 2002-12-23 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Reconstructing the Binding Site of Factor Xa in Trypsin Reveals Ligand-induced Structural Plasticity
J.MOL.BIOL., 325, 2003
|
|
1J15
 
 | | BENZAMIDINE IN COMPLEX WITH RAT TRYPSIN MUTANT X99/175/190RT | | Descriptor: | BENZAMIDINE, CALCIUM ION, SULFATE ION, ... | | Authors: | Stubbs, M.T. | | Deposit date: | 2002-11-30 | | Release date: | 2002-12-23 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Reconstructing the Binding Site of Factor Xa in Trypsin Reveals Ligand-induced Structural Plasticity
J.MOL.BIOL., 325, 2003
|
|
5TEG
 
 | |
1V2L
 
 | | Benzamidine in complex with bovine trypsin variant X(triple.Glu)bT.D1 | | Descriptor: | BENZAMIDINE, CALCIUM ION, SULFATE ION, ... | | Authors: | Rauh, D, Klebe, G, Stubbs, M.T. | | Deposit date: | 2003-10-17 | | Release date: | 2004-06-01 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Understanding protein-ligand interactions: the price of protein flexibility
J.Mol.Biol., 335, 2004
|
|
1V2P
 
 | | Trypsin inhibitor in complex with bovine trypsin variant X(SSYI)bT.A4 | | Descriptor: | CALCIUM ION, METHYL N-[(4-METHYLPHENYL)SULFONYL]GLYCYL-3-[AMINO(IMINO)METHYL]-D-PHENYLALANINATE, SULFATE ION, ... | | Authors: | Rauh, D, Klebe, G, Stubbs, M.T. | | Deposit date: | 2003-10-17 | | Release date: | 2004-06-01 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | Understanding protein-ligand interactions: the price of protein flexibility
J.Mol.Biol., 335, 2004
|
|
1V2W
 | | Trypsin inhibitor in complex with bovine trypsin variant X(SSAI)bT.B4 | | Descriptor: | CALCIUM ION, METHYL N-[(4-METHYLPHENYL)SULFONYL]GLYCYL-3-[AMINO(IMINO)METHYL]-D-PHENYLALANINATE, SULFATE ION, ... | | Authors: | Rauh, D, Klebe, G, Stubbs, M.T. | | Deposit date: | 2003-10-17 | | Release date: | 2004-06-01 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Understanding protein-ligand interactions: the price of protein flexibility
J.Mol.Biol., 335, 2004
|
|
1V2R
 | | Trypsin inhibitor in complex with bovine trypsin variant X(SSRI)bT.B4 | | Descriptor: | CALCIUM ION, METHYL N-[(4-METHYLPHENYL)SULFONYL]GLYCYL-3-[AMINO(IMINO)METHYL]-D-PHENYLALANINATE, SULFATE ION, ... | | Authors: | Rauh, D, Klebe, G, Stubbs, M.T. | | Deposit date: | 2003-10-17 | | Release date: | 2004-06-01 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Understanding protein-ligand interactions: the price of protein flexibility
J.Mol.Biol., 335, 2004
|
|
3ZOJ
 
 | | High-resolution structure of Pichia Pastoris aquaporin Aqy1 at 0.88 A | | Descriptor: | AQUAPORIN, CHLORIDE ION, octyl beta-D-glucopyranoside | | Authors: | Kosinska-Eriksson, U, Fischer, G, Friemann, R, Enkavi, G, Tajkhorshid, E, Neutze, R. | | Deposit date: | 2013-02-21 | | Release date: | 2013-06-26 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (0.88 Å) | | Cite: | Subangstrom Resolution X-Ray Structure Details Aquaporin-Water Interactions
Science, 340, 2013
|
|
1QRY
 
 | | Homeobox protein VND (ventral nervous system defective protein) | | Descriptor: | PROTEIN (HOMEOBOX VENTRAL NERVOUS SYSTEM DEFECTIVE PROTEIN) | | Authors: | Xiang, B. | | Deposit date: | 1999-06-16 | | Release date: | 1999-07-06 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Distortion of the three-dimensional structure of the vnd/NK-2 homeodomain bound to DNA induced by an embryonically lethal A35T point mutation.
Biochemistry, 42, 2003
|
|
1WC2
 
 | | Beta-1,4-D-endoglucanase Cel45A from blue mussel Mytilus edulis at 1.2A | | Descriptor: | ACETATE ION, DI(HYDROXYETHYL)ETHER, ENDOGLUCANASE | | Authors: | Jakobsson, E, Mahdi, S, Kleywegt, G.J, Stahlberg, J. | | Deposit date: | 2004-11-08 | | Release date: | 2006-05-24 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Glucomannan and beta-glucan degradation by Mytilus edulis Cel45A: Crystal structure and activity comparison with GH45 subfamily A, B and C.
Carbohydr Polym, 277, 2022
|
|
1GQF
 
 | |
1PMC
 
 | |
2FL2
 
 | | crystal structure of KSP in complex with inhibitor 19 | | Descriptor: | (1S)-1-CYCLOPROPYL-2-[(2S)-4-(2,5-DIFLUOROPHENYL)-2-PHENYL-2,5-DIHYDRO-1H-PYRROL-1-YL]-2-OXOETHANAMINE, ADENOSINE-5'-DIPHOSPHATE, Kinesin-like protein KIF11, ... | | Authors: | Yan, Y. | | Deposit date: | 2006-01-05 | | Release date: | 2006-02-07 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Kinesin spindle protein (KSP) inhibitors. Part 2: the design, synthesis, and characterization of 2,4-diaryl-2,5-dihydropyrrole inhibitors of the mitotic kinesin KSP.
Bioorg.Med.Chem.Lett., 16, 2006
|
|
2FL6
 
 | | crystal structure of KSP in complex with inhibitor 6 | | Descriptor: | (2S)-4-(2,5-DIFLUOROPHENYL)-N,N-DIMETHYL-2-PHENYL-2,5-DIHYDRO-1H-PYRROLE-1-CARBOXAMIDE, ADENOSINE-5'-DIPHOSPHATE, Kinesin-like protein KIF11, ... | | Authors: | Yan, Y. | | Deposit date: | 2006-01-05 | | Release date: | 2006-02-07 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Kinesin spindle protein (KSP) inhibitors. Part 2: the design, synthesis, and characterization of 2,4-diaryl-2,5-dihydropyrrole inhibitors of the mitotic kinesin KSP.
Bioorg.Med.Chem.Lett., 16, 2006
|
|
2FKY
 
 | | crystal structure of KSP in complex with inhibitor 13 | | Descriptor: | (2S)-4-(2,5-DIFLUOROPHENYL)-N-METHYL-2-PHENYL-N-PIPERIDIN-4-YL-2,5-DIHYDRO-1H-PYRROLE-1-CARBOXAMIDE, ADENOSINE-5'-DIPHOSPHATE, Kinesin-like protein KIF11, ... | | Authors: | Yan, Y. | | Deposit date: | 2006-01-05 | | Release date: | 2006-02-07 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Kinesin spindle protein (KSP) inhibitors. Part 2: the design, synthesis, and characterization of 2,4-diaryl-2,5-dihydropyrrole inhibitors of the mitotic kinesin KSP.
Bioorg.Med.Chem.Lett., 16, 2006
|
|
2G1Q
 
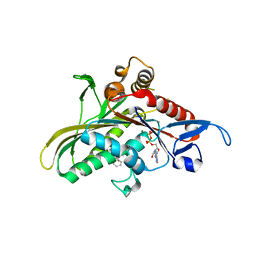 | | crystal structure of KSP in complex with inhibitor 9h | | Descriptor: | (5S)-5-(3-AMINOPROPYL)-3-(2,5-DIFLUOROPHENYL)-N-ETHYL-5-PHENYL-4,5-DIHYDRO-1H-PYRAZOLE-1-CARBOXAMIDE, ADENOSINE-5'-DIPHOSPHATE, Kinesin-like protein KIF11, ... | | Authors: | Yan, Y. | | Deposit date: | 2006-02-14 | | Release date: | 2006-10-03 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.51 Å) | | Cite: | Kinesin spindle protein (KSP) inhibitors. Part 4: Structure-based design of 5-alkylamino-3,5-diaryl-4,5-dihydropyrazoles as potent, water-soluble inhibitors of the mitotic kinesin KSP.
Bioorg.Med.Chem.Lett., 16, 2006
|
|
2H5E
 
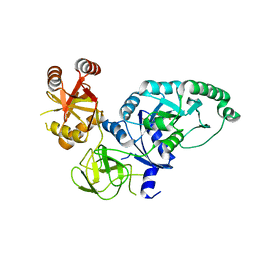 | | Crystal structure of E.coli polypeptide release factor RF3 | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, Peptide chain release factor RF-3 | | Authors: | Song, H.W, Zhou, Z.H. | | Deposit date: | 2006-05-26 | | Release date: | 2007-05-15 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | RF3 induces ribosomal conformational changes responsible for dissociation of class I release factors
Cell(Cambridge,Mass.), 129, 2007
|
|
5U8F
 
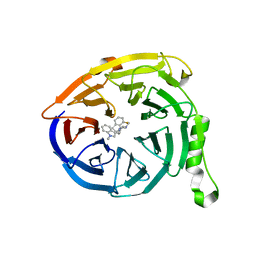 | | Polycomb protein EED in complex with inhibitor: (3R,4S)-1-[(1S)-7-fluoro-2,3-dihydro-1H-inden-1-yl]-N,N-dimethyl-4-(1-methyl-1H-indol-3-yl)pyrrolidin-3-amine | | Descriptor: | (3R,4S)-1-[(1S)-7-fluoro-2,3-dihydro-1H-inden-1-yl]-N,N-dimethyl-4-(1-methyl-1H-indol-3-yl)pyrrolidin-3-amine, Polycomb protein EED | | Authors: | Jakob, C.G, Zhu, H. | | Deposit date: | 2016-12-14 | | Release date: | 2017-03-15 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.343 Å) | | Cite: | SAR of amino pyrrolidines as potent and novel protein-protein interaction inhibitors of the PRC2 complex through EED binding.
Bioorg. Med. Chem. Lett., 27, 2017
|
|
5TJT
 
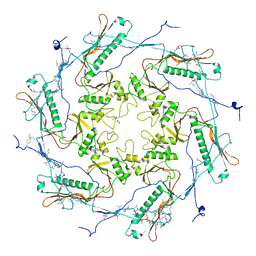 | |
5U69
 
 | | Polycomb protein EED in complex with inhibitor: (3R,4S)-1-[(2-methoxyphenyl)methyl]-N,N-dimethyl-4-(1-methylindol-3-yl)pyrrolidin-3-amine | | Descriptor: | (3R,4S)-1-[(2-methoxyphenyl)methyl]-N,N-dimethyl-4-(1-methylindol-3-yl)pyrrolidin-3-amine, Polycomb protein EED | | Authors: | Jakob, C.G, Zhu, H. | | Deposit date: | 2016-12-07 | | Release date: | 2017-03-15 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.28 Å) | | Cite: | SAR of amino pyrrolidines as potent and novel protein-protein interaction inhibitors of the PRC2 complex through EED binding.
Bioorg. Med. Chem. Lett., 27, 2017
|
|
5U8A
 
 | | Polycomb protein EED in complex with inhibitor: (3R,4S)-1-[(2-bromo-6-fluorophenyl)methyl]-N,N-dimethyl-4-(1-methyl-1H-indol-3-yl)pyrrolidin-3-amine | | Descriptor: | (3R,4S)-1-[(2-bromo-6-fluorophenyl)methyl]-N,N-dimethyl-4-(1-methyl-1H-indol-3-yl)pyrrolidin-3-amine, Polycomb protein EED | | Authors: | Jakob, C.G, Zhu, H. | | Deposit date: | 2016-12-14 | | Release date: | 2017-03-15 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | SAR of amino pyrrolidines as potent and novel protein-protein interaction inhibitors of the PRC2 complex through EED binding.
Bioorg. Med. Chem. Lett., 27, 2017
|
|
1QLS
 
 | | S100C (S100A11),OR CALGIZZARIN, IN COMPLEX WITH ANNEXIN I N-TERMINUS | | Descriptor: | ANNEXIN I, CALCIUM ION, S100C PROTEIN | | Authors: | Rety, S, Sopkova, J, Renouard, M, Osterloh, D, Gerke, V, Russo-Marie, F, Lewit-Bentley, A. | | Deposit date: | 1999-09-15 | | Release date: | 2000-02-25 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural Basis of the Ca2+ Dependent Association between S100C (S100A11) and its Target, the N-Terminal Part of Annexin I
Structure, 8, 2000
|
|
