4YDF
 
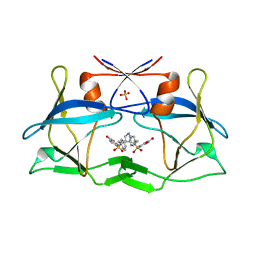 | | Crystal structure of compound 9 in complex with HTLV-1 Protease | | Descriptor: | HTLV-1 Protease, N-benzyl-N-[(3S,4S)-4-{benzyl[(4-nitrophenyl)sulfonyl]amino}pyrrolidin-3-yl]-3-nitrobenzenesulfonamide, SULFATE ION | | Authors: | Kuhnert, M, Blum, A, Steuber, H, Diederich, W.E. | | Deposit date: | 2015-02-22 | | Release date: | 2016-02-03 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.804 Å) | | Cite: | Privileged Structures Meet Human T-Cell Leukemia Virus-1 (HTLV-1): C2-Symmetric 3,4-Disubstituted Pyrrolidines as Nonpeptidic HTLV-1 Protease Inhibitors.
J.Med.Chem., 58, 2015
|
|
2N9C
 
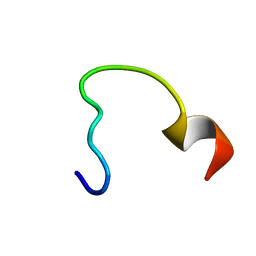 | | NRAS Isoform 5 | | Descriptor: | GTPase NRas | | Authors: | Markowitz, J, Mal, T.K, Yuan, C, Courtney, N.B, Patel, M, Stiff, A.R, Blachly, J, Walker, C, Eisfeld, A, de la Chapelle, A, Carson III, W.E. | | Deposit date: | 2015-11-13 | | Release date: | 2016-03-23 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural characterization of NRAS isoform 5.
Protein Sci., 25, 2016
|
|
4QB4
 
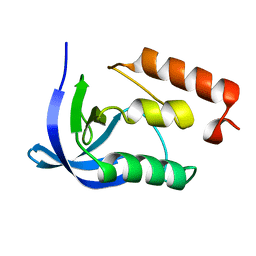 | |
9BVY
 
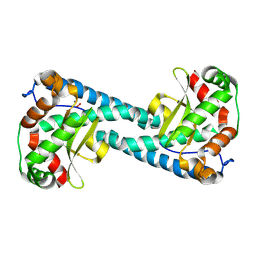 | | Neutron Structure of Peroxide-Soaked Tyr34Phe MnSOD | | Descriptor: | HYDROGEN PEROXIDE, MANGANESE (II) ION, Superoxide dismutase [Mn], ... | | Authors: | Azadmanesh, J, Slobodnik, K, Struble, L.R, Cone, E.A, Dasgupta, M, Lutz, W.E, Kumar, S, Natarajan, A, Coates, L, Weiss, K.L, Myles, D.A.A, Kroll, T, Borgstahl, G.E.O. | | Deposit date: | 2024-05-20 | | Release date: | 2025-03-12 | | Method: | NEUTRON DIFFRACTION (2.3 Å) | | Cite: | The role of Tyr34 in proton coupled electron transfer and product inhibition of manganese superoxide dismutase.
Nat Commun, 16, 2025
|
|
9BWQ
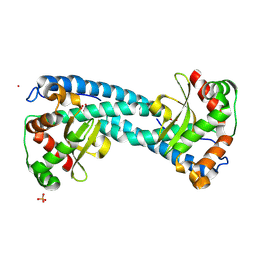 
 | | X-ray Counterpart to the Neutron Structure of Peroxide-Soaked Tyr34Phe MnSOD | | Descriptor: | HYDROGEN PEROXIDE, MANGANESE (II) ION, PHOSPHATE ION, ... | | Authors: | Azadmanesh, J, Slobodnik, K, Struble, L.R, Cone, E.A, Dasgupta, M, Lutz, W.E, Kumar, S, Natarajan, A, Coates, L, Weiss, K.L, Myles, D.A.A, Kroll, T, Borgstahl, G.E.O. | | Deposit date: | 2024-05-21 | | Release date: | 2025-03-12 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | The role of Tyr34 in proton coupled electron transfer and product inhibition of manganese superoxide dismutase.
Nat Commun, 16, 2025
|
|
9BWR
 
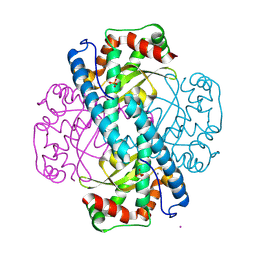 | | X-ray Counterpart to the Neutron Structure of Reduced Tyr34Phe MnSOD | | Descriptor: | MANGANESE (II) ION, PHOSPHATE ION, POTASSIUM ION, ... | | Authors: | Azadmanesh, J, Slobodnik, K, Struble, L.R, Cone, E.A, Dasgupta, M, Lutz, W.E, Kumar, S, Natarajan, A, Coates, L, Weiss, K.L, Myles, D.A.A, Kroll, T, Borgstahl, G.E.O. | | Deposit date: | 2024-05-21 | | Release date: | 2025-03-12 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The role of Tyr34 in proton coupled electron transfer and product inhibition of manganese superoxide dismutase.
Nat Commun, 16, 2025
|
|
9BWM
 
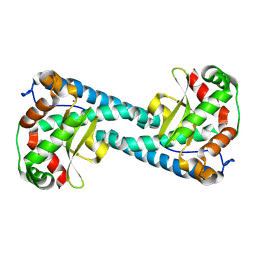 | | Neutron Structure of Oxidized Tyr34Phe MnSOD | | Descriptor: | MANGANESE (II) ION, Superoxide dismutase [Mn], mitochondrial | | Authors: | Azadmanesh, J, Slobodnik, K, Struble, L.R, Cone, E.A, Dasgupta, M, Lutz, W.E, Kumar, S, Natarajan, A, Coates, L, Weiss, K.L, Myles, D.A.A, Kroll, T, Borgstahl, G.E.O. | | Deposit date: | 2024-05-21 | | Release date: | 2025-03-12 | | Method: | NEUTRON DIFFRACTION (2.28 Å) | | Cite: | The role of Tyr34 in proton coupled electron transfer and product inhibition of manganese superoxide dismutase.
Nat Commun, 16, 2025
|
|
9BW2
 
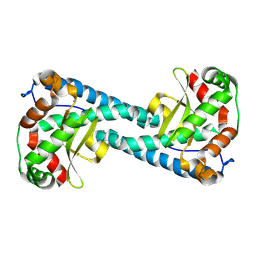 | | Neutron Structure of Reduced Tyr34Phe MnSOD | | Descriptor: | MANGANESE (II) ION, Superoxide dismutase [Mn], mitochondrial | | Authors: | Azadmanesh, J, Slobodnik, K, Struble, L.R, Cone, E.A, Dasgupta, M, Lutz, W.E, Kumar, S, Natarajan, A, Coates, L, Weiss, K.L, Myles, D.A.A, Kroll, T, Borgstahl, G.E.O. | | Deposit date: | 2024-05-20 | | Release date: | 2025-03-12 | | Method: | NEUTRON DIFFRACTION (2.5 Å) | | Cite: | The role of Tyr34 in proton coupled electron transfer and product inhibition of manganese superoxide dismutase.
Nat Commun, 16, 2025
|
|
4FBY
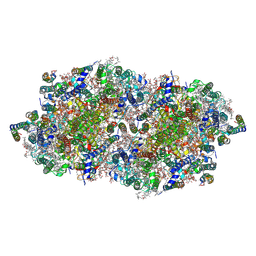 
 | | fs X-ray diffraction of Photosystem II | | Descriptor: | 1,2-DI-O-ACYL-3-O-[6-DEOXY-6-SULFO-ALPHA-D-GLUCOPYRANOSYL]-SN-GLYCEROL, 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE, 1,2-DISTEAROYL-MONOGALACTOSYL-DIGLYCERIDE, ... | | Authors: | Kern, J, Alonso-Mori, R, Hellmich, J, Tran, R, Hattne, J, Laksmono, H, Gloeckner, C, Echols, N, Sierra, R.G, Sellberg, J, Lassalle-Kaiser, B, Gildea, R.J, Glatzel, P, Grosse-Kunstleve, R.W, Latimer, M.J, Mcqueen, T.A, Difiore, D, Fry, A.R, Messerschmidt, M.M, Miahnahri, A, Schafer, D.W, Seibert, M.M, Sokaras, D, Weng, T.-C, Zwart, P.H, White, W.E, Adams, P.D, Bogan, M.J, Boutet, S, Williams, G.J, Messinger, J, Sauter, N.K, Zouni, A, Bergmann, U, Yano, J, Yachandra, V.K. | | Deposit date: | 2012-05-23 | | Release date: | 2012-06-20 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (6.56 Å) | | Cite: | Room temperature femtosecond X-ray diffraction of photosystem II microcrystals.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
6NS5
 
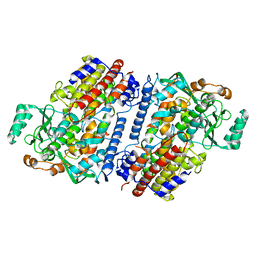 | | Crystal structure of fungal lipoxygenase from Fusarium graminearum. Second C2 crystal form. | | Descriptor: | FE (II) ION, lipoxygenase | | Authors: | Pakhomova, S, Boeglin, W.E, Neau, D.B, Bartlett, S.G, Brash, A.R, Newcomer, M.E. | | Deposit date: | 2019-01-24 | | Release date: | 2019-03-27 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.79 Å) | | Cite: | An ensemble of lipoxygenase structures reveals novel conformations of the Fe coordination sphere.
Protein Sci., 28, 2019
|
|
9B2G
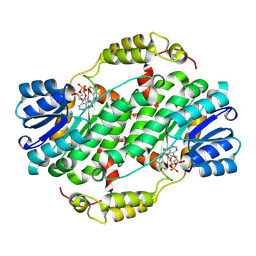 
 | | Crystal structure of short chain dehydrogenase reductase 9 (short chain dehydrogenase reductase 9C4) in complex with NADH | | Descriptor: | 1,4-DIHYDRONICOTINAMIDE ADENINE DINUCLEOTIDE, Dehydrogenase/reductase SDR family member 9 | | Authors: | Pakhomova, S, Belyaeva, O.V, Boeglin, W.E, Kedishvili, N.Y, Brash, A.R, Newcomer, M.E, Popov, K.M. | | Deposit date: | 2024-03-15 | | Release date: | 2025-03-19 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The large substrate binding pocket of dehydrogenase reductase 9 underlies its ability to oxidize diverse pro-inflammatory and pro-resolving oxylipins.
To Be Published
|
|
9B2F
 
 | | Crystal structure of short chain dehydrogenase reductase 9 (short chain dehydrogenase reductase 9C4) in complex with NAD+ | | Descriptor: | Dehydrogenase/reductase SDR family member 9, MALONATE ION, NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | Authors: | Pakhomova, S, Belyaeva, O.V, Boeglin, W.E, Kedishvili, N.Y, Brash, A.R, Newcomer, M.E, Popov, K.M. | | Deposit date: | 2024-03-15 | | Release date: | 2025-03-19 | | Method: | X-RAY DIFFRACTION (1.662 Å) | | Cite: | The large substrate binding pocket of dehydrogenase reductase 9 underlies its ability to oxidize diverse pro-inflammatory and pro-resolving oxylipins.
To Be Published
|
|
6NS2
 
 | | Crystal structure of fungal lipoxygenase from Fusarium graminearum. P212121 crystal form. | | Descriptor: | FE (II) ION, lipoxygenase | | Authors: | Pakhomova, S, Boeglin, W.E, Neau, D.B, Bartlett, S.G, Brash, A.R, Newcomer, M.E. | | Deposit date: | 2019-01-24 | | Release date: | 2019-03-27 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.79 Å) | | Cite: | An ensemble of lipoxygenase structures reveals novel conformations of the Fe coordination sphere.
Protein Sci., 28, 2019
|
|
6NS3
 | | Crystal structure of fungal lipoxygenase from Fusarium graminearum. I222 crystal form. | | Descriptor: | FE (II) ION, lipoxygenase | | Authors: | Pakhomova, S, Boeglin, W.E, Neau, D.B, Bartlett, S.G, Brash, A.R, Newcomer, M.E. | | Deposit date: | 2019-01-24 | | Release date: | 2019-03-27 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.84 Å) | | Cite: | An ensemble of lipoxygenase structures reveals novel conformations of the Fe coordination sphere.
Protein Sci., 28, 2019
|
|
6NS4
 
 | | Crystal structure of fungal lipoxygenase from Fusarium graminearum. C2 crystal form. | | Descriptor: | ACETATE ION, FE (II) ION, GLYCEROL, ... | | Authors: | Pakhomova, S, Boeglin, W.E, Neau, D.B, Bartlett, S.G, Brash, A.R, Newcomer, M.E. | | Deposit date: | 2019-01-24 | | Release date: | 2019-03-27 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | An ensemble of lipoxygenase structures reveals novel conformations of the Fe coordination sphere.
Protein Sci., 28, 2019
|
|
6NS6
 
 | | Crystal structure of fungal lipoxygenase from Fusarium graminearum. P21 crystal form. | | Descriptor: | FE (II) ION, lipoxygenase | | Authors: | Pakhomova, S, Boeglin, W.E, Neau, D.B, Bartlett, S.G, Brash, A.R, Newcomer, M.E. | | Deposit date: | 2019-01-24 | | Release date: | 2019-03-27 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | An ensemble of lipoxygenase structures reveals novel conformations of the Fe coordination sphere.
Protein Sci., 28, 2019
|
|
4H7B
 
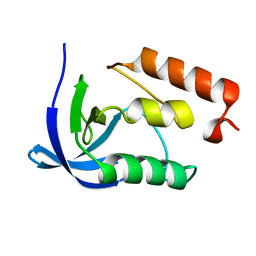 | |
1NWI
 
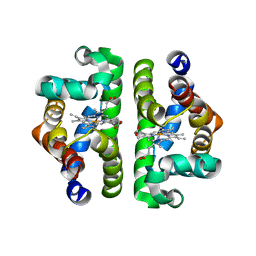 | |
1NWN
 | |
1J6X
 | | CRYSTAL STRUCTURE OF HELICOBACTER PYLORI LUXS | | Descriptor: | AUTOINDUCER-2 PRODUCTION PROTEIN LUXS, METHIONINE, ZINC ION | | Authors: | Lewis, H.A, Furlong, E.B, Bergseid, M.G, Sanderson, W.E, Buchanan, S.G. | | Deposit date: | 2001-05-14 | | Release date: | 2001-06-08 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.38 Å) | | Cite: | A structural genomics approach to the study of quorum sensing: crystal structures of three LuxS orthologs.
Structure, 9, 2001
|
|
1J6W
 
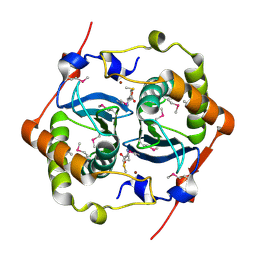 | | CRYSTAL STRUCTURE OF HAEMOPHILUS INFLUENZAE LUXS | | Descriptor: | AUTOINDUCER-2 PRODUCTION PROTEIN LUXS, METHIONINE, ZINC ION | | Authors: | Lewis, H.A, Furlong, E.B, Bergseid, M.G, Sanderson, W.E, Buchanan, S.G. | | Deposit date: | 2001-05-14 | | Release date: | 2001-06-08 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | A structural genomics approach to the study of quorum sensing: crystal structures of three LuxS orthologs.
Structure, 9, 2001
|
|
1INN
 | | CRYSTAL STRUCTURE OF D. RADIODURANS LUXS, P21 | | Descriptor: | AUTOINDUCER-2 PRODUCTION PROTEIN LUXS, METHIONINE, ZINC ION | | Authors: | Lewis, H.A, Furlong, E.B, Bergseid, M.G, Sanderson, W.E, Buchanan, S.G. | | Deposit date: | 2001-05-14 | | Release date: | 2001-06-08 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | A structural genomics approach to the study of quorum sensing: crystal structures of three LuxS orthologs.
Structure, 9, 2001
|
|
1GND
 
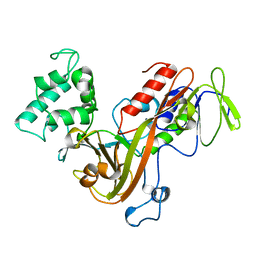 | | GUANINE NUCLEOTIDE DISSOCIATION INHIBITOR, ALPHA-ISOFORM | | Descriptor: | GUANINE NUCLEOTIDE DISSOCIATION INHIBITOR | | Authors: | Schalk, I, Zeng, K, Wu, S.-K, Stura, E.A, Metteson, J, Huang, M, Tandon, A, Wilson, I.A, Balch, W.E. | | Deposit date: | 1996-07-10 | | Release date: | 1997-02-12 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Structure and mutational analysis of Rab GDP-dissociation inhibitor.
Nature, 381, 1996
|
|
1J6V
 
 | | CRYSTAL STRUCTURE OF D. RADIODURANS LUXS, C2 | | Descriptor: | AUTOINDUCER-2 PRODUCTION PROTEIN LUXS, ZINC ION | | Authors: | Lewis, H.A, Furlong, E.B, Bergseid, M.G, Sanderson, W.E, Buchanan, S.G. | | Deposit date: | 2001-05-14 | | Release date: | 2001-06-08 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | A structural genomics approach to the study of quorum sensing: crystal structures of three LuxS orthologs.
Structure, 9, 2001
|
|
5JRN
 
 | | Crystal Structure of a Xylanase in Complex with a Monosaccharide at 2.84 Angstroem resolution | | Descriptor: | Endo-1,4-beta-xylanase, GLYCEROL, methyl beta-D-xylopyranoside | | Authors: | Gomez, S, Payne, A.M, Savko, M, Fox, G.C, Shepard, W.E, Fernandez, F.J, Vega, M.C. | | Deposit date: | 2016-05-06 | | Release date: | 2017-05-24 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.841 Å) | | Cite: | Structural and functional characterization of a highly stable endo-beta-1,4-xylanase from Fusarium oxysporum and its development as an efficient immobilized biocatalyst.
Biotechnol Biofuels, 9, 2016
|
|
