2WUH
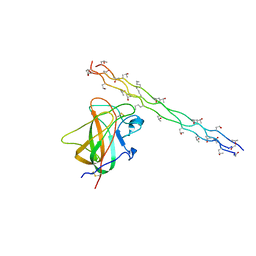 
 | | Crystal structure of the DDR2 discoidin domain bound to a triple- helical collagen peptide | | Descriptor: | COLLAGEN PEPTIDE, DISCOIDIN DOMAIN RECEPTOR 2 | | Authors: | Carafoli, F, Bihan, D, Stathopoulos, S, Konitsiotis, A.D, Kvansakul, M, Farndale, R.W, Leitinger, B, Hohenester, E. | | Deposit date: | 2009-10-05 | | Release date: | 2009-12-29 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystallographic Insight Into Collagen Recognition by Discoidin Domain Receptor 2
Structure, 17, 2009
|
|
6TZC
 
 | | Crystal Structure of African Swine Fever Virus A179L with the Autophagy Regulator Beclin | | Descriptor: | Apoptosis regulator Bcl-2 homolog, Beclin-1, Maltose/maltodextrin-binding periplasmic protein, ... | | Authors: | Banjara, S, Kvansakul, M, Hinds, M.G. | | Deposit date: | 2019-08-12 | | Release date: | 2019-11-20 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.41 Å) | | Cite: | Crystal Structure of African Swine Fever Virus A179L with the Autophagy Regulator Beclin.
Viruses, 11, 2019
|
|
5VYP
 
 | | Crystal structure of the Plant Defensin NsD7 bound to PIP2 | | Descriptor: | Defensin NsD7, GLYCEROL, SULFATE ION, ... | | Authors: | Jarva, M, Lay, F.T, Hulett, M, Kvansakul, M. | | Deposit date: | 2017-05-25 | | Release date: | 2017-08-02 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure of the defensin NsD7 in complex with PIP2 reveals that defensin : lipid oligomer topologies are dependent on lipid type.
FEBS Lett., 591, 2017
|
|
7QS8
 
 | | Structural insight into the Scribble PDZ domains interaction with the oncogenic Human T-cell lymphotrophic virus-1 (HTLV-1) Tax1 | | Descriptor: | Protein Tax-1, Protein scribble homolog | | Authors: | Javorsky, A, Soares da Costa, T.P, Mackie, E.R, Humbert, P.O, Kvansakul, M. | | Deposit date: | 2022-01-13 | | Release date: | 2022-09-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structural insight into the Scribble PDZ domains interaction with the oncogenic Human T-cell lymphotrophic virus-1 (HTLV-1) Tax1 PBM.
Febs J., 290, 2023
|
|
7QRS
 
 | | Structural insight into the Scribble PDZ domains interaction with the oncogenic Human T-cell lymphotrophic virus-1 (HTLV-1) Tax1 | | Descriptor: | ACETATE ION, GLYCEROL, Protein Tax-1, ... | | Authors: | Javorsky, A, Soares da Costa, T.P, Mackie, E.R, Humbert, P.O, Kvansakul, M. | | Deposit date: | 2022-01-12 | | Release date: | 2022-09-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Structural insight into the Scribble PDZ domains interaction with the oncogenic Human T-cell lymphotrophic virus-1 (HTLV-1) Tax1 PBM.
Febs J., 290, 2023
|
|
7QRT
 
 | | Structural insight into the Scribble PDZ domains interaction with the oncogenic Human T-cell lymphotrophic virus-1 (HTLV-1) Tax1 | | Descriptor: | CHLORIDE ION, Protein Tax-1, Protein scribble homolog, ... | | Authors: | Javorsky, A, Soares da Costa, T.P, Mackie, E.R, Humbert, P.O, Kvansakul, M, Maddumage, J.C. | | Deposit date: | 2022-01-12 | | Release date: | 2022-09-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural insight into the Scribble PDZ domains interaction with the oncogenic Human T-cell lymphotrophic virus-1 (HTLV-1) Tax1 PBM.
Febs J., 290, 2023
|
|
7QSA
 
 | |
7QS9
 
 | |
7QSB
 
 | |
7QTX
 
 | | Kaposi sarcoma associated herpes virus (KSHV) encoded apoptosis inhibitor, KsBcl-2 in complex with Puma BH3 | | Descriptor: | 1,2-ETHANEDIOL, BROMIDE ION, Bcl-2, ... | | Authors: | Suraweera, C.D, Hinds, M.G, Kvansakul, M. | | Deposit date: | 2022-01-17 | | Release date: | 2022-11-23 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.11598849 Å) | | Cite: | Structural Insight into KsBcl-2 Mediated Apoptosis Inhibition by Kaposi Sarcoma Associated Herpes Virus.
Viruses, 14, 2022
|
|
7QTW
 
 | | Kaposi sarcoma associated herpes virus(KSHV) encoded apoptosis inhibitor, KsBcl-2 in complex with Bid BH3 | | Descriptor: | 1,2-ETHANEDIOL, BH3-interacting domain death agonist p15, Bcl-2 | | Authors: | Suraweera, C.D, Hinds, M.G, Kvansakul, M. | | Deposit date: | 2022-01-17 | | Release date: | 2022-11-23 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.41 Å) | | Cite: | Structural Insight into KsBcl-2 Mediated Apoptosis Inhibition by Kaposi Sarcoma Associated Herpes Virus.
Viruses, 14, 2022
|
|
5WOU
 
 | |
4B4S
 
 | |
7P0U
 
 | | ORF virus encoded Bcl-2 homolog ORFV125 in complex with Puma BH3 peptide | | Descriptor: | 1,2-ETHANEDIOL, Activator of apoptosis harakiri, Apoptosis inhibitor, ... | | Authors: | Suraweera, C.D, Hinds, M.G, Kvansakul, M. | | Deposit date: | 2021-06-30 | | Release date: | 2022-07-13 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.99374259 Å) | | Cite: | Structural Investigation of Orf Virus Bcl-2 Homolog ORFV125 Interactions with BH3-Motifs from BH3-Only Proteins Puma and Hrk.
Viruses, 13, 2021
|
|
7P0S
 
 | |
7P33
 
 | | Epstein-Barr virus encoded Bcl-2 homolog BHRF-1 in complex with Bid BH3 peptide | | Descriptor: | 1,2-ETHANEDIOL, Apoptosis regulator BHRF1, BH3-interacting domain death agonist p15, ... | | Authors: | Suraweera, C.D, Hinds, M.G, Kvansakul, M. | | Deposit date: | 2021-07-07 | | Release date: | 2022-07-20 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.78542733 Å) | | Cite: | Crystal Structures of Epstein-Barr Virus Bcl-2 Homolog BHRF1 Bound to Bid and Puma BH3 Motif Peptides.
Viruses, 14, 2022
|
|
7P9W
 
 | | Epstein-Barr virus encoded apoptosis regulator BHRF1 in complex with Puma BH3 | | Descriptor: | 1,2-ETHANEDIOL, 1,3-PROPANDIOL, AMMONIUM ION, ... | | Authors: | Suraweera, C.D, Hinds, M.G, Kvansakul, M. | | Deposit date: | 2021-07-28 | | Release date: | 2022-08-10 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.00010061 Å) | | Cite: | Crystal Structures of Epstein-Barr Virus Bcl-2 Homolog BHRF1 Bound to Bid and Puma BH3 Motif Peptides.
Viruses, 14, 2022
|
|
6XA8
 | |
6XA6
 
 | |
6XA7
 | |
6XY4
 
 | |
6XY6
 | |
5VWK
 
 | | Crystal structure of human Scribble PDZ1:Beta-PIX complex | | Descriptor: | Beta-PIX, Protein scribble homolog, SULFATE ION | | Authors: | Lim, K.Y.B, Kvansakul, M. | | Deposit date: | 2017-05-22 | | Release date: | 2017-11-08 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Structural basis for the differential interaction of Scribble PDZ domains with the guanine nucleotide exchange factor beta-PIX.
J. Biol. Chem., 292, 2017
|
|
5VWI
 
 | |
5WOS
 
 | |
