8Q3S
 
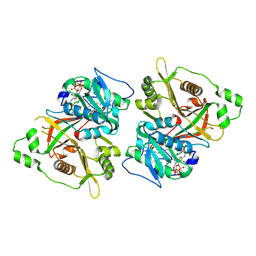 | | HsNMT1 in complex with both MyrCoA and GNCFSKAR inhibitor peptide | | Descriptor: | CHLORIDE ION, COENZYME A, GLY-ASN-CYS-PHE-SER-LYS-ALA-ARG, ... | | Authors: | Dian, C, Giglione, C, Meinnel, T. | | Deposit date: | 2023-08-04 | | Release date: | 2024-08-14 | | Last modified: | 2025-02-26 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | Novel, tightly structurally related N-myristoyltransferase inhibitors display equally potent yet distinct inhibitory mechanisms.
Structure, 32, 2024
|
|
8Q23
 
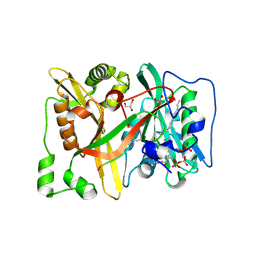 | | HsNMT1 in complex with both MyrCoA and Ac-D-ORN-SFSKPR inhibitor peptide | | Descriptor: | ACE-ORD-SER-PHE-SER-LYS-PRO-ARG, CHLORIDE ION, GLYCEROL, ... | | Authors: | Dian, C, Giglione, C, Meinnel, T. | | Deposit date: | 2023-08-01 | | Release date: | 2024-08-14 | | Last modified: | 2025-02-26 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Novel, tightly structurally related N-myristoyltransferase inhibitors display equally potent yet distinct inhibitory mechanisms.
Structure, 32, 2024
|
|
8Q3T
 
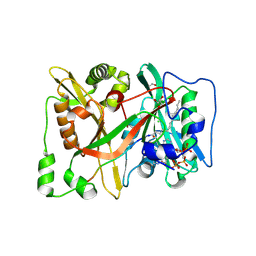 | | HsNMT1 in complex with both MyrCoA and GNCFSKPRVPTK inhibitor peptide | | Descriptor: | CHLORIDE ION, COENZYME A, GLYCEROL, ... | | Authors: | Dian, C, Giglione, C, Meinnel, T. | | Deposit date: | 2023-08-04 | | Release date: | 2024-08-14 | | Last modified: | 2025-02-26 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Novel, tightly structurally related N-myristoyltransferase inhibitors display equally potent yet distinct inhibitory mechanisms.
Structure, 32, 2024
|
|
8Q2Z
 
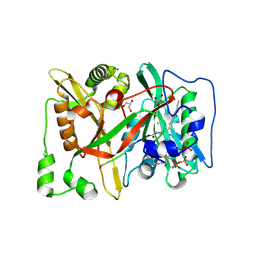 | | HsNMT1 in complex with both MyrCoA and GNLLSKFR peptide | | Descriptor: | CHLORIDE ION, COENZYME A, GLYCEROL, ... | | Authors: | Dian, C, Giglione, C, Meinnel, T. | | Deposit date: | 2023-08-03 | | Release date: | 2024-08-14 | | Last modified: | 2025-02-26 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Novel, tightly structurally related N-myristoyltransferase inhibitors display equally potent yet distinct inhibitory mechanisms.
Structure, 32, 2024
|
|
8Q24
 
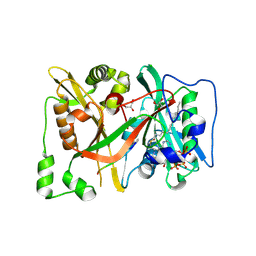 | | HsNMT1 in complex with both MyrCoA and (DAB)SFSKPR inhibitor peptide | | Descriptor: | CHLORIDE ION, DAB-SER-PHE-SER-LYS-PRO-ARG, GLYCEROL, ... | | Authors: | Dian, C, Giglione, C, Meinnel, T. | | Deposit date: | 2023-08-01 | | Release date: | 2024-08-14 | | Last modified: | 2025-02-26 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Novel, tightly structurally related N-myristoyltransferase inhibitors display equally potent yet distinct inhibitory mechanisms.
Structure, 32, 2024
|
|
3IQS
 
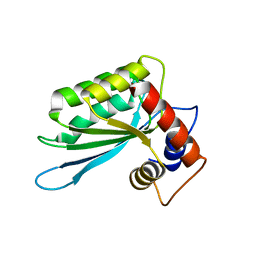 | | Crystal structure of the anti-viral APOBEC3G catalytic domain | | Descriptor: | DNA dC->dU-editing enzyme APOBEC-3G, ZINC ION | | Authors: | Holden, L.G, Prochnow, C, Chang, Y.P, Bransteitter, R, Chelico, L, Sen, U, Stevens, R.C, Goodman, R.F, Chen, X.S. | | Deposit date: | 2009-08-20 | | Release date: | 2009-11-10 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of the anti-viral APOBEC3G catalytic domain and functional implications.
Nature, 456, 2008
|
|
8QUS
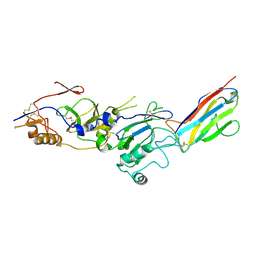 
 | |
8QU7
 
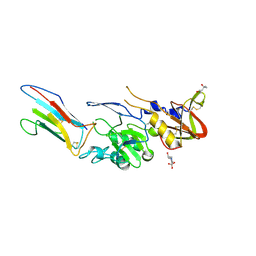 | |
8QOC
 
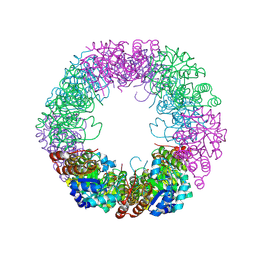 | | Crystal structure of Staphylococcus aureus PLP Synthase (Pdx1) | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, MAGNESIUM ION, ... | | Authors: | Ullah, N, Wrenger, C, Betzel, C. | | Deposit date: | 2023-09-28 | | Release date: | 2024-10-02 | | Method: | X-RAY DIFFRACTION (2.83 Å) | | Cite: | Structure and dynamics of the staphylococcal pyridoxal 5-phosphate synthase complex reveal transient interactions at the enzyme interface.
J.Biol.Chem., 300, 2024
|
|
8R1M
 
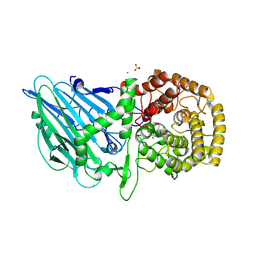 | | Structure of TxGH116 with covalently bound N-azido-octyl aziridine | | Descriptor: | (1R,2R,3R,4S)-4-[8-(azanyldiazenyl)octylamino]-3-(hydroxymethyl)cyclopentane-1,2-diol, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Offen, W.A, Davies, G.J. | | Deposit date: | 2023-11-02 | | Release date: | 2024-10-02 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Selective labelling of GBA2 in cells with fluorescent beta-d-arabinofuranosyl cyclitol aziridines.
Chem Sci, 15, 2024
|
|
8QAT
 
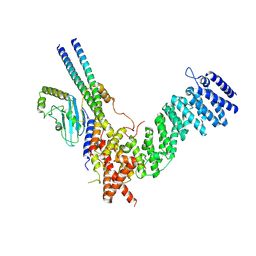 | | Cryo-EM structure of Fts-Hook3-FHIP1B at 3.2 A resolution. | | Descriptor: | AKT-interacting protein, FHF complex subunit HOOK-interacting protein 1B, Protein Hook homolog 3 | | Authors: | Abid Ali, F, Carter, A.P. | | Deposit date: | 2023-08-23 | | Release date: | 2024-09-04 | | Last modified: | 2025-04-23 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | KIF1C activates and extends dynein movement through the FHF cargo adapter.
Nat.Struct.Mol.Biol., 32, 2025
|
|
8QQL
 
 | |
1XEK
 
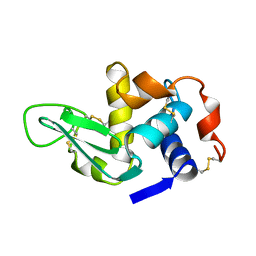 | |
1XEI
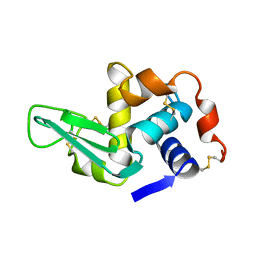 
 | |
1XEJ
 
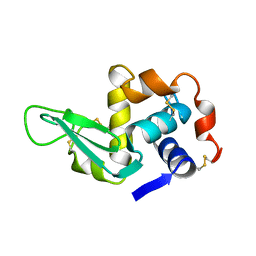 | |
1LMB
 
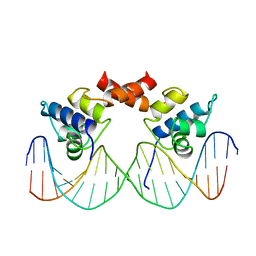 | | REFINED 1.8 ANGSTROM CRYSTAL STRUCTURE OF THE LAMBDA REPRESSOR-OPERATOR COMPLEX | | Descriptor: | DNA (5'-D(*AP*AP*TP*AP*CP*CP*AP*CP*TP*GP*GP*CP*GP*GP*TP*GP*A P*TP*AP*T)-3'), DNA (5'-D(*TP*AP*TP*AP*TP*CP*AP*CP*CP*GP*CP*CP*AP*GP*TP*GP*G P*TP*AP*T)-3'), PROTEIN (LAMBDA REPRESSOR) | | Authors: | Beamer, L.J, Pabo, C.O. | | Deposit date: | 1991-11-05 | | Release date: | 1991-11-05 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Refined 1.8 A crystal structure of the lambda repressor-operator complex.
J.Mol.Biol., 227, 1992
|
|
1LRP
 
 | |
1MSC
 
 | |
1RL0
 
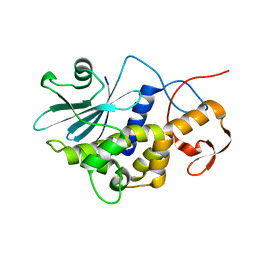 | | Crystal structure of a new ribosome-inactivating protein (RIP): dianthin 30 | | Descriptor: | Antiviral protein DAP-30 | | Authors: | Fermani, S, Falini, G, Ripamonti, A, Bolognesi, A, Polito, L, Stirpe, F. | | Deposit date: | 2003-11-24 | | Release date: | 2004-12-07 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | The 1.4A structure of dianthin 30 indicates a role of surface potential at the active site of type 1 ribosome inactivating proteins
J.Struct.Biol., 149, 2005
|
|
1QIO
 
 | |
1PMK
 
 | |
1POZ
 
 | | SOLUTION STRUCTURE OF THE HYALURONAN BINDING DOMAIN OF HUMAN CD44 | | Descriptor: | CD44 antigen | | Authors: | Teriete, P, Banerji, S, Blundell, C.D, Kahmann, J.D, Pickford, A.R, Wright, A.J, Campbell, I.D, Jackson, D.G, Day, A.J. | | Deposit date: | 2003-06-16 | | Release date: | 2004-03-16 | | Last modified: | 2024-10-30 | | Method: | SOLUTION NMR | | Cite: | Structure of the Regulatory Hyaluronan Binding Domain in the Inflammatory Leukocyte Homing Receptor CD44.
Mol.Cell, 13, 2004
|
|
1PK2
 
 | |
1PML
 
 | |
1PBK
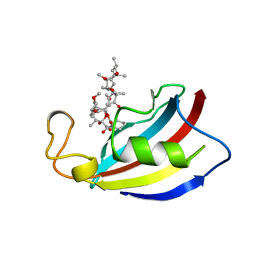 
 | | HOMOLOGOUS DOMAIN OF HUMAN FKBP25 | | Descriptor: | FKBP25, RAPAMYCIN IMMUNOSUPPRESSANT DRUG | | Authors: | Liang, J, Hung, D.T, Schreiber, S.L, Clardy, J. | | Deposit date: | 1995-09-01 | | Release date: | 1996-10-14 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure of the human 25 kDa FK506 binding protein complexed with rapamycin.
J.Am.Chem.Soc., 118, 1996
|
|
