3EW1
 
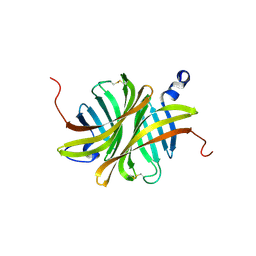 | | Crystal structure of rhizavidin | | Descriptor: | rhizavidin | | Authors: | Livnah, O, Meir, A. | | Deposit date: | 2008-10-14 | | Release date: | 2008-12-23 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystal structure of rhizavidin: insights into the enigmatic high-affinity interaction of an innate biotin-binding protein dimer.
J.Mol.Biol., 386, 2009
|
|
3EW2
 
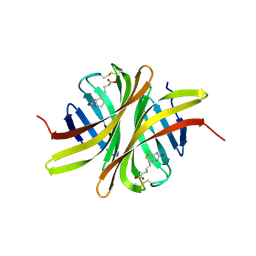 | | Crystal structure of rhizavidin-biotin complex | | Descriptor: | BIOTIN, rhizavidin | | Authors: | Livnah, O, Meir, A. | | Deposit date: | 2008-10-14 | | Release date: | 2008-12-23 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of rhizavidin: insights into the enigmatic high-affinity interaction of an innate biotin-binding protein dimer.
J.Mol.Biol., 386, 2009
|
|
4Z2P
 
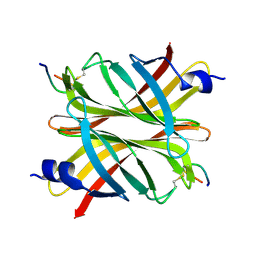 | |
4Z27
 
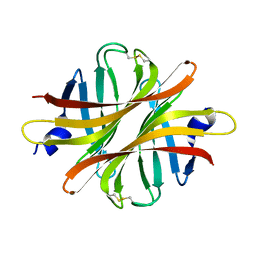 | |
4Z2V
 
 | |
4Z2O
 
 | |
4Z28
 | | Crystal structure of short hoefavidin biotin complex | | Descriptor: | 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Avidin family, BIOTIN | | Authors: | Livnah, O, Avraham, O. | | Deposit date: | 2015-03-29 | | Release date: | 2015-07-15 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.66 Å) | | Cite: | Hoefavidin: A dimeric bacterial avidin with a C-terminal binding tail.
J.Struct.Biol., 191, 2015
|
|
4Z6J
 
 | |
6RFP
 
 | | ERK2 MAP kinase with mutations at Helix-G | | Descriptor: | Mitogen-activated protein kinase 1 | | Authors: | Livnah, O, Eitan-Wexler, M. | | Deposit date: | 2019-04-16 | | Release date: | 2020-05-13 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | The bacterial metalloprotease NleD selectively cleaves mitogen-activated protein kinases that have high flexibility in their activation loop.
J.Biol.Chem., 295, 2020
|
|
3P63
 
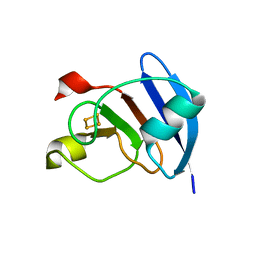 | | Structure of M. laminosus Ferredoxin with a shorter L1,2 loop | | Descriptor: | FE2/S2 (INORGANIC) CLUSTER, Ferredoxin | | Authors: | Livnah, O, Nechushtai, R, Eisenberg-Domovich, Y, Michaeli, D. | | Deposit date: | 2010-10-11 | | Release date: | 2011-02-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Allostery in the ferredoxin protein motif does not involve a conformational switch.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
6QTY
 
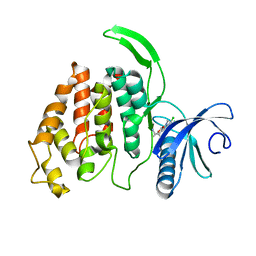 | |
6QYX
 
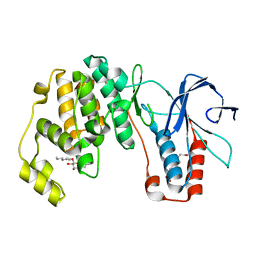 | | p38(alpha) MAP kinase with the activation loop of ERK2 | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Mitogen-activated protein kinase 14,Mitogen-activated protein kinase 1,Mitogen-activated protein kinase 14, octyl beta-D-glucopyranoside | | Authors: | Livnah, O, Eitan-Wexler, M, Vinograd, N. | | Deposit date: | 2019-03-10 | | Release date: | 2020-04-01 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.66 Å) | | Cite: | The bacterial metalloprotease NleD selectively cleaves mitogen-activated protein kinases that have high flexibility in their activation loop.
J.Biol.Chem., 295, 2020
|
|
8ASU
 
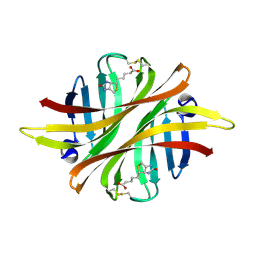 | |
8AVP
 
 | |
8AMH
 
 | |
8AN6
 
 | |
8ASR
 
 | |
8AST
 
 | |
8ASS
 
 | |
8AVJ
 
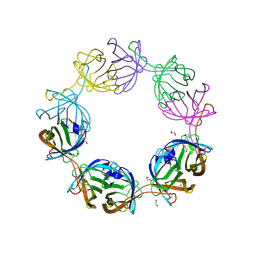 | |
7P0O
 
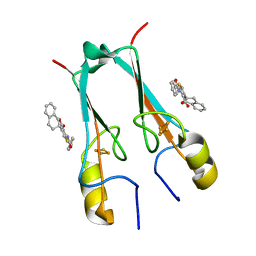 | | mitoNEET bound to M1 molecule | | Descriptor: | 2-benzamido-4-[(2~{R})-1,2,3,4-tetrahydronaphthalen-2-yl]thiophene-3-carboxylic acid, 2-benzamido-4-[(2~{S})-1,2,3,4-tetrahydronaphthalen-2-yl]thiophene-3-carboxylic acid, CDGSH iron-sulfur domain-containing protein 1, ... | | Authors: | Livnah, O, Eisenberg-Domovich, Y, Marjault, H.B, Nechushtai, R. | | Deposit date: | 2021-06-30 | | Release date: | 2022-05-25 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | An anti-diabetic drug targets NEET (CISD) proteins through destabilization of their [2Fe-2S] clusters.
Commun Biol, 5, 2022
|
|
7P0P
 
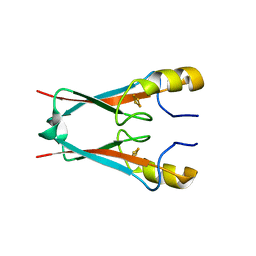 | | NAF-1 bound to M1 molecule | | Descriptor: | 2-benzamido-4-[(2~{R})-1,2,3,4-tetrahydronaphthalen-2-yl]thiophene-3-carboxylic acid, CDGSH iron-sulfur domain-containing protein 2, FE2/S2 (INORGANIC) CLUSTER | | Authors: | Livnah, O, Eisenberg-Domovich, Y, Marjault, H.B, Nechushtai, R. | | Deposit date: | 2021-06-30 | | Release date: | 2022-05-25 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | An anti-diabetic drug targets NEET (CISD) proteins through destabilization of their [2Fe-2S] clusters.
Commun Biol, 5, 2022
|
|
2OFA
 
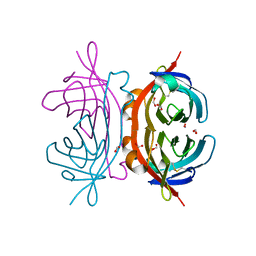 | | Crystal structure of apo AVR4 (R112L,C122S) | | Descriptor: | Avidin-related protein 4/5, FORMIC ACID | | Authors: | Livnah, O, Hayouka, R, Eisenberg-Domovich, Y. | | Deposit date: | 2007-01-03 | | Release date: | 2007-12-25 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Critical importance of loop conformation to avidin-enhanced hydrolysis of an active biotin ester.
Acta Crystallogr.,Sect.D, 64, 2008
|
|
2OF8
 
 | | Crystal structure of AVR4 (D39A/C122S)-BNA complex | | Descriptor: | 5-(2-OXO-HEXAHYDRO-THIENO[3,4-D]IMIDAZOL-6-YL)-PENTANOIC ACID (4-NITRO-PHENYL)-AMIDE, Avidin-related protein 4/5, FORMIC ACID | | Authors: | Livnah, O, Hayouka, R, Eisenberg-Domovich, Y. | | Deposit date: | 2007-01-03 | | Release date: | 2007-12-25 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.05 Å) | | Cite: | Critical importance of loop conformation to avidin-enhanced hydrolysis of an active biotin ester.
Acta Crystallogr.,Sect.D, 64, 2008
|
|
2OFB
 
 | | Crystal structure of AVR4 (R112L/C122S)-BNA complex | | Descriptor: | 5-(2-OXO-HEXAHYDRO-THIENO[3,4-D]IMIDAZOL-6-YL)-PENTANOIC ACID (4-NITRO-PHENYL)-AMIDE, Avidin-related protein 4/5, FORMIC ACID | | Authors: | Livnah, O, Hayouka, R, Eisenberg-Domovich, Y. | | Deposit date: | 2007-01-03 | | Release date: | 2007-12-25 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.16 Å) | | Cite: | Critical importance of loop conformation to avidin-enhanced hydrolysis of an active biotin ester.
Acta Crystallogr.,Sect.D, 64, 2008
|
|
