4L17
 
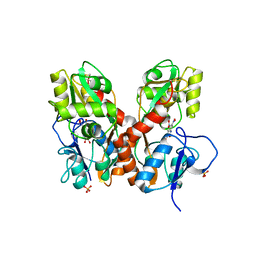 | | GluA2-L483Y-A665C ligand-binding domain in complex with the antagonist DNQX | | Descriptor: | 6,7-DINITROQUINOXALINE-2,3-DIONE, Glutamate receptor 2, SULFATE ION | | Authors: | Lau, A.Y, Blachowicz, L, Roux, B. | | Deposit date: | 2013-06-02 | | Release date: | 2013-08-14 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | A conformational intermediate in glutamate receptor activation.
Neuron, 79, 2013
|
|
1ZWI
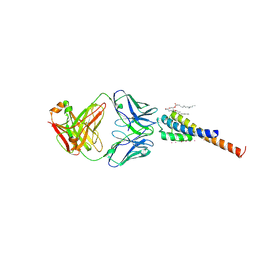 
 | | Structure of mutant KcsA potassium channel | | Descriptor: | DIACYL GLYCEROL, NONAN-1-OL, POTASSIUM ION, ... | | Authors: | Cordero-Morales, J.F, Cuello, L.G, Zhao, Y, Jogini, V, Cortes, D.M, Roux, B, Perozo, E. | | Deposit date: | 2005-06-03 | | Release date: | 2006-03-07 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Molecular determinants of gating at the potassium-channel selectivity filter.
Nat.Struct.Mol.Biol., 13, 2006
|
|
3HWB
 
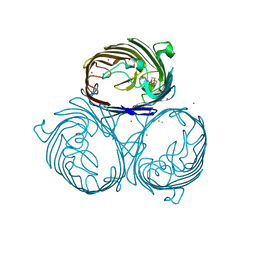 | | Cation selective pathway of OmpF porin revealed by anomalous diffraction | | Descriptor: | HEXAETHYLENE GLYCOL, Outer membrane protein F, RUBIDIUM ION | | Authors: | Balasundaresan, D, Raychaudhury, S, Blachowicz, L, Roux, B. | | Deposit date: | 2009-06-17 | | Release date: | 2009-12-08 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Cation-selective pathway of OmpF porin revealed by anomalous X-ray diffraction.
J.Mol.Biol., 396, 2010
|
|
3HW9
 
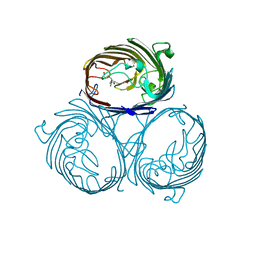 | | Cation selective pathway of OmpF porin revealed by anomalous x-ray diffraction | | Descriptor: | CHLORIDE ION, HEXAETHYLENE GLYCOL, Outer membrane protein F, ... | | Authors: | Balasundaresan, D, Raychaudhury, S, Blachowicz, L, Roux, B. | | Deposit date: | 2009-06-17 | | Release date: | 2009-12-08 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.61 Å) | | Cite: | Cation-selective pathway of OmpF porin revealed by anomalous X-ray diffraction.
J.Mol.Biol., 396, 2010
|
|
6BL8
 
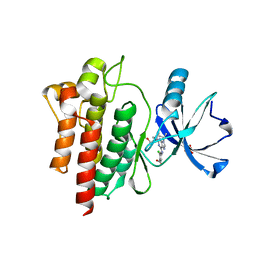 | |
5T88
 
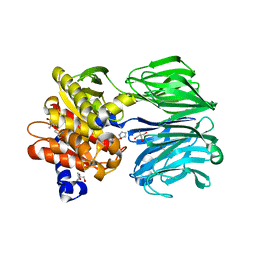 | | Prolyl oligopeptidase from Pyrococcus furiosus | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CHLORIDE ION, PROLINE, ... | | Authors: | Ellis-Guardiola, K, Lewis, J, Sukumar, N. | | Deposit date: | 2016-09-06 | | Release date: | 2017-09-06 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.902 Å) | | Cite: | Crystal Structure and Conformational Dynamics of Pyrococcus furiosus Prolyl Oligopeptidase.
Biochemistry, 58, 2019
|
|
6CAN
 
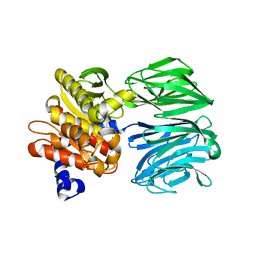 | |
8FZO
 
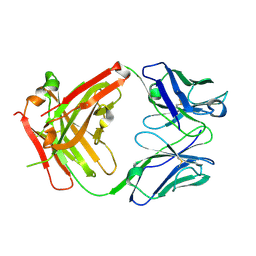 | |
8G1B
 
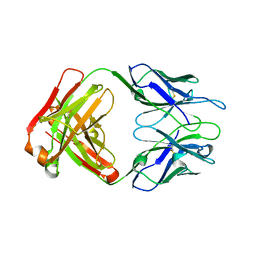 | | Crystal structure of polyreactive 2G02 human Fab | | Descriptor: | The heavy chain of human 2G02 Fab fragment, The light chain of human 2G02 Fab fragment | | Authors: | Borowska, M.T, Adams, E.J. | | Deposit date: | 2023-02-01 | | Release date: | 2023-10-18 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.67 Å) | | Cite: | Biochemical and biophysical characterization of natural polyreactivity in antibodies.
Cell Rep, 42, 2023
|
|
8G19
 
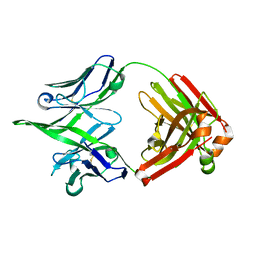 | | Crystal structure of monoreactive 4C05 human Fab | | Descriptor: | Heavy chain of anti-influenza human 4C05 Fab fragment, Light chain of anti-influenza human 4C05 Fab fragment | | Authors: | Borowska, M.T, Adams, E.J. | | Deposit date: | 2023-02-01 | | Release date: | 2023-10-18 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Biochemical and biophysical characterization of natural polyreactivity in antibodies.
Cell Rep, 42, 2023
|
|
8FZP
 
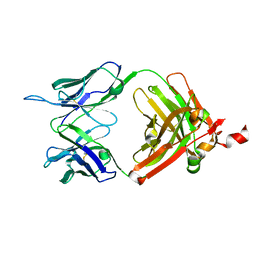 | |
8G1C
 
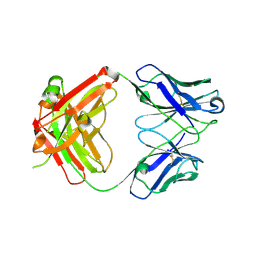 | | Crystal structure of polyreactive 3B03 human Fab | | Descriptor: | Heavy chain of monoreactive 3B03 human Fab fragment, Light chain of monoreactive 3B03 human Fab fragment | | Authors: | Borowska, M.T, Adams, E.J. | | Deposit date: | 2023-02-01 | | Release date: | 2023-10-18 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.63 Å) | | Cite: | Biochemical and biophysical characterization of natural polyreactivity in antibodies.
Cell Rep, 42, 2023
|
|
5HM7
 
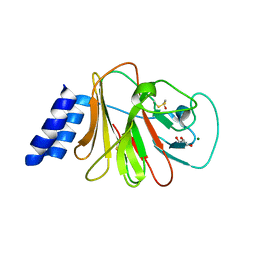 | | The Intracellular domain of Butyrophilin 3A1 protein | | Descriptor: | BETA-MERCAPTOETHANOL, Butyrophilin subfamily 3 member A1, GLYCEROL, ... | | Authors: | Adams, E.J, Gu, S. | | Deposit date: | 2016-01-15 | | Release date: | 2017-07-26 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Phosphoantigen-induced conformational change of butyrophilin 3A1 (BTN3A1) and its implication on V gamma 9V delta 2 T cell activation.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
3CK6
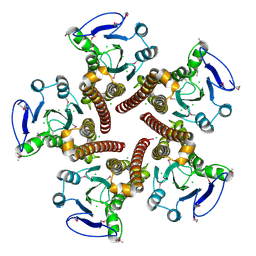 
 | | Crystal structure of ZntB cytoplasmic domain from Vibrio parahaemolyticus RIMD 2210633 | | Descriptor: | CHLORIDE ION, Putative membrane transport protein | | Authors: | Tan, K, Sather, A, Moy, S, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2008-03-14 | | Release date: | 2008-03-25 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure and electrostatic property of cytoplasmic domain of ZntB transporter.
Protein Sci., 18, 2009
|
|
3OR6
 
 | | On the structural basis of modal gating behavior in K+channels - E71Q | | Descriptor: | POTASSIUM ION, Voltage-gated potassium channel, antibody fab fragment heavy chain, ... | | Authors: | Chakrapani, S, Cordero-Morales, J.F, Jogini, V, Perozo, E. | | Deposit date: | 2010-09-06 | | Release date: | 2011-01-05 | | Last modified: | 2024-11-27 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | On the structural basis of modal gating behavior in K(+) channels.
Nat.Struct.Mol.Biol., 18, 2011
|
|
6W0B
 
 | | Open-gate KcsA soaked in 2 mM BaCl2 | | Descriptor: | BARIUM ION, Fab Heavy Chain, Fab Light Chain, ... | | Authors: | Rohaim, A, Gong, L, Li, J. | | Deposit date: | 2020-02-29 | | Release date: | 2020-07-08 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (3.604 Å) | | Cite: | Open and Closed Structures of a Barium-Blocked Potassium Channel.
J.Mol.Biol., 432, 2020
|
|
6W0H
 
 | | Closed-gate KcsA soaked in 5mM KCl/5mM BaCl2 | | Descriptor: | Fab Heavy Chain, Fab Light Chain, POTASSIUM ION, ... | | Authors: | Rohaim, A, Gong, L, Li, J. | | Deposit date: | 2020-02-29 | | Release date: | 2020-07-08 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Open and Closed Structures of a Barium-Blocked Potassium Channel.
J.Mol.Biol., 432, 2020
|
|
6W0G
 
 | | Closed-gate KcsA soaked in 1mM KCl/5mM BaCl2 | | Descriptor: | Fab Heavy Chain, Fab Light Chain, POTASSIUM ION, ... | | Authors: | Rohaim, A, Gong, L, Li, J. | | Deposit date: | 2020-02-29 | | Release date: | 2020-07-08 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Open and Closed Structures of a Barium-Blocked Potassium Channel.
J.Mol.Biol., 432, 2020
|
|
6W0C
 
 | | Open-gate KcsA soaked in 4 mM BaCl2 | | Descriptor: | BARIUM ION, Fab Heavy Chain, Fab Light Chain, ... | | Authors: | Rohaim, A, Gong, L, Li, J. | | Deposit date: | 2020-02-29 | | Release date: | 2020-07-08 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (3.556 Å) | | Cite: | Open and Closed Structures of a Barium-Blocked Potassium Channel.
J.Mol.Biol., 432, 2020
|
|
6W0D
 
 | | Open-gate KcsA soaked in 5 mM BaCl2 | | Descriptor: | BARIUM ION, Fab Heavy Chain, Fab Light Chain, ... | | Authors: | Rohaim, A, Gong, L, Li, J. | | Deposit date: | 2020-02-29 | | Release date: | 2020-07-08 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (3.639 Å) | | Cite: | Open and Closed Structures of a Barium-Blocked Potassium Channel.
J.Mol.Biol., 432, 2020
|
|
6W0J
 
 | | Closed-gate KcsA incubated in BaCl2/NaCl | | Descriptor: | BARIUM ION, Fab Heavy Chain, Fab Light Chain, ... | | Authors: | Rohaim, A, Gong, L, Li, J. | | Deposit date: | 2020-02-29 | | Release date: | 2020-07-08 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Open and Closed Structures of a Barium-Blocked Potassium Channel.
J.Mol.Biol., 432, 2020
|
|
6W0A
 
 | | Open-gate KcsA soaked in 1 mM BaCl2 | | Descriptor: | BARIUM ION, Fab Heavy Chain, Fab Light Chain, ... | | Authors: | Rohaim, A, Gong, L, Li, J. | | Deposit date: | 2020-02-29 | | Release date: | 2020-07-08 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (3.237 Å) | | Cite: | Open and Closed Structures of a Barium-Blocked Potassium Channel.
J.Mol.Biol., 432, 2020
|
|
6W0I
 
 | | Closed-gate KcsA soaked in 10mM KCl/5mM BaCl2 | | Descriptor: | Fab Heavy Chain, Fab Light Chain, POTASSIUM ION, ... | | Authors: | Rohaim, A, Gong, L, Li, J. | | Deposit date: | 2020-02-29 | | Release date: | 2020-07-08 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.328 Å) | | Cite: | Open and Closed Structures of a Barium-Blocked Potassium Channel.
J.Mol.Biol., 432, 2020
|
|
6W0E
 
 | | Open-gate KcsA soaked in 10 mM BaCl2 | | Descriptor: | BARIUM ION, Fab Heavy Chain, Fab Light Chain, ... | | Authors: | Rohaim, A, Gong, L, Li, J. | | Deposit date: | 2020-02-29 | | Release date: | 2020-07-08 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (3.512 Å) | | Cite: | Open and Closed Structures of a Barium-Blocked Potassium Channel.
J.Mol.Biol., 432, 2020
|
|
6W0F
 
 | | Closed-gate KcsA soaked in 0mM KCl/5mM BaCl2 | | Descriptor: | Fab Heavy Chain, Fab Light Chain, POTASSIUM ION, ... | | Authors: | Rohaim, A, Gong, L, Li, J. | | Deposit date: | 2020-02-29 | | Release date: | 2020-07-08 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Open and Closed Structures of a Barium-Blocked Potassium Channel.
J.Mol.Biol., 432, 2020
|
|
