1SMC
 
 | | Mycobacterium tuberculosis dUTPase complexed with dUTP in the absence of metal ion. | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, DEOXYURIDINE-5'-TRIPHOSPHATE, Deoxyuridine 5'-triphosphate nucleotidohydrolase, ... | | Authors: | Sawaya, M.R, Chan, S, Segelke, B, Lekin, T, Krupka, H, Cho, U.S, Kim, M.-Y, So, M, Kim, C.-Y, Naranjo, C.M, Rogers, Y.C, Park, M.S, Waldo, G.S, Pashkov, I, Cascio, D, Yeates, T.O, Perry, J.L, Terwilliger, T.C, Eisenberg, D, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2004-03-09 | | Release date: | 2004-03-16 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of the Mycobacterium tuberculosis dUTPase: insights into the catalytic mechanism.
J.Mol.Biol., 341, 2004
|
|
7UJI
 
 | |
7VYO
 
 | | The structure of GdmN | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Wei, J, Zheng, J, Zhou, J, Kang, Q, Bai, L. | | Deposit date: | 2021-11-14 | | Release date: | 2022-11-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Endowing homodimeric carbamoyltransferase GdmN with iterative functions through structural characterization and mechanistic studies.
Nat Commun, 13, 2022
|
|
7UPH
 
 | | Structure of a ribosome with tethered subunits | | Descriptor: | 30S ribosomal protein S10, 30S ribosomal protein S11, 30S ribosomal protein S12, ... | | Authors: | Kim, D.S, Watkins, A, Bidstrup, E, Lee, J, Topkar, V.V, Kofman, C, Schwarz, K.J, Liu, Y, Pintilie, G, Roney, E, Das, R, Jewett, M.C. | | Deposit date: | 2022-04-15 | | Release date: | 2022-08-17 | | Last modified: | 2025-03-19 | | Method: | ELECTRON MICROSCOPY (4.18 Å) | | Cite: | Three-dimensional structure-guided evolution of a ribosome with tethered subunits.
Nat.Chem.Biol., 18, 2022
|
|
7UK2
 
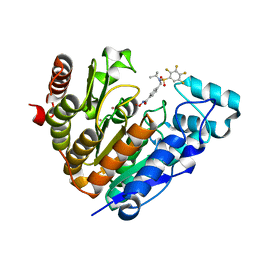 | | Crystal structure of Danio rerio histone deacetylase 6 catalytic domain 2 complexed with NN-390 | | Descriptor: | Hdac6 protein, N-hydroxy-4-{[(propan-2-yl)(2,3,4,5-tetrafluorobenzene-1-sulfonyl)amino]methyl}benzamide, POTASSIUM ION, ... | | Authors: | Erdogan, F, Seo, H.-S, Dhe-Paganon, S. | | Deposit date: | 2022-03-31 | | Release date: | 2022-11-02 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | High Efficacy and Drug Synergy of HDAC6-Selective Inhibitor NN-429 in Natural Killer (NK)/T-Cell Lymphoma.
Pharmaceuticals, 15, 2022
|
|
6OPJ
 
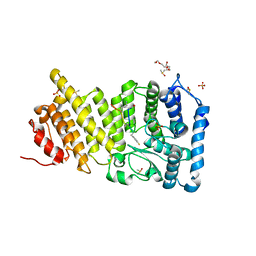 | | Menin in complex with peptide inhibitor 25 | | Descriptor: | DIMETHYL SULFOXIDE, Menin, Peptide inhibitor 25, ... | | Authors: | Linhares, B.M, Fortuna, P, Cierpicki, T, Grembecka, J, Berlicki, L. | | Deposit date: | 2019-04-25 | | Release date: | 2020-09-02 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.5006572 Å) | | Cite: | Covalent and noncovalent constraints yield a figure eight-like conformation of a peptide inhibiting the menin-MLL interaction.
Eur.J.Med.Chem., 207, 2020
|
|
9ONO
 
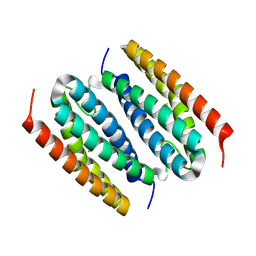 | | Fe-bound B. pseudomallei rubrerythrin | | Descriptor: | FE (III) ION, Rubrerythrin | | Authors: | Budziszewski, G.R, Snell, M.E, Monteiro, D.C.F, Lynch, M.L, Bowman, S.E.J. | | Deposit date: | 2025-05-15 | | Release date: | 2025-06-25 | | Method: | X-RAY DIFFRACTION (1.66 Å) | | Cite: | Burkholderia pseudomallei rubrerythrin promiscuously binds metals in a structurally pre-formed bimetallic binding site.
Biorxiv, 2025
|
|
9NND
 
 | | Structure of the HERV-K (HML-2) spike complex | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Shaked, R, Katz, M, Diskin, R. | | Deposit date: | 2025-03-05 | | Release date: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (2.13 Å) | | Cite: | The prefusion structure of the HERV-K spike complex
To Be Published
|
|
9NYO
 
 | |
9R2L
 | | De novo designed N37 protein fold | | Descriptor: | ACETATE ION, ETHANOLAMINE, N37, ... | | Authors: | Pacesa, M, Miao, Y, Georgeon, S, Schmidt, J, Correia, B.E. | | Deposit date: | 2025-04-30 | | Release date: | 2025-05-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Reimagine accessibility of protein space with generative model
To Be Published
|
|
9R2R
 | |
9R2V
 | | De novo designed M16 protein fold | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CALCIUM ION, FORMIC ACID, ... | | Authors: | Pacesa, M, Miao, Y, Georgeon, S, Schmidt, J, Correia, B.E. | | Deposit date: | 2025-05-01 | | Release date: | 2025-05-14 | | Method: | X-RAY DIFFRACTION (2.49 Å) | | Cite: | Reimagine accessibility of protein space with generative model
To Be Published
|
|
9ONR
 | | rubrerythrin- E53A E124A | | Descriptor: | DI(HYDROXYETHYL)ETHER, Rubrerythrin, THIOCYANATE ION | | Authors: | Budziszewski, G.R, Snell, M.E, Monteiro, D.C.F, Lynch, M.L, Bowman, S.E.J. | | Deposit date: | 2025-05-15 | | Release date: | 2025-06-25 | | Method: | X-RAY DIFFRACTION (1.38 Å) | | Cite: | Burkholderia pseudomallei rubrerythrin promiscuously binds metals in a structurally pre-formed bimetallic binding site.
Biorxiv, 2025
|
|
9ONN
 
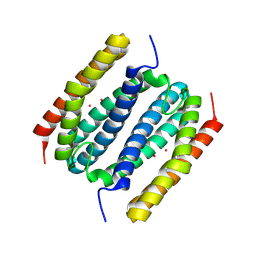 | | Co-bound B. pseudomallei Rubrerythrin | | Descriptor: | COBALT (II) ION, Rubrerythrin | | Authors: | Budziszewski, G.R, Snell, M.E, Monteiro, D.C.F, Lynch, M.L, Bowman, S.E.J. | | Deposit date: | 2025-05-15 | | Release date: | 2025-06-25 | | Method: | X-RAY DIFFRACTION (1.53 Å) | | Cite: | Burkholderia pseudomallei rubrerythrin promiscuously binds metals in a structurally pre-formed bimetallic binding site.
Biorxiv, 2025
|
|
9ONM
 
 | | near-apo rubrerythrin | | Descriptor: | DI(HYDROXYETHYL)ETHER, FE (III) ION, Rubrerythrin | | Authors: | Budziszewski, G.R, Snell, M.E, Monteiro, D.C.F, Lynch, M.L, Bowman, S.E.J. | | Deposit date: | 2025-05-15 | | Release date: | 2025-06-25 | | Method: | X-RAY DIFFRACTION (1.57 Å) | | Cite: | Burkholderia pseudomallei rubrerythrin promiscuously binds metals in a structurally pre-formed bimetallic binding site.
Biorxiv, 2025
|
|
9ONQ
 | | Mn-bound B. pseudomallei rubrerythrin | | Descriptor: | MANGANESE (II) ION, Rubrerythrin | | Authors: | Budziszewski, G.R, Snell, M.E, Monteiro, D.C.F, Lynch, M.L, Bowman, S.E.J. | | Deposit date: | 2025-05-15 | | Release date: | 2025-06-25 | | Method: | X-RAY DIFFRACTION (1.47 Å) | | Cite: | Burkholderia pseudomallei rubrerythrin promiscuously binds metals in a structurally pre-formed bimetallic binding site.
Biorxiv, 2025
|
|
9NA5
 
 | | IRAK4 in Complex with Compound 24 | | Descriptor: | (6P)-6-[(8R)-3-cyanopyrrolo[1,2-b]pyridazin-7-yl]-4-({(1s,4S)-4-[1-(difluoromethyl)-1H-pyrazol-4-yl]cyclohexyl}amino)-N-[(2S)-2-fluoro-3-hydroxy-3-methylbutyl]pyridine-3-carboxamide, Interleukin-1 receptor-associated kinase 4, SULFATE ION | | Authors: | Ferrao, R, Lansdon, E.B. | | Deposit date: | 2025-02-11 | | Release date: | 2025-05-28 | | Last modified: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (1.73 Å) | | Cite: | Discovery of Edecesertib (GS-5718): A Potent, Selective Inhibitor of IRAK4.
J.Med.Chem., 68, 2025
|
|
9R2O
 
 | | De novo designed N5 protein fold | | Descriptor: | N5, POTASSIUM ION, THIOCYANATE ION | | Authors: | Pacesa, M, Miao, Y, Georgeon, S, Schmidt, J, Correia, B.E. | | Deposit date: | 2025-04-30 | | Release date: | 2025-05-14 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Reimagine accessibility of protein space with generative model
To Be Published
|
|
9NA2
 
 | | IRAK4 in Complex with Compound 9 | | Descriptor: | (6P)-6-[(8R)-3-cyanopyrrolo[1,2-b]pyridazin-7-yl]-N-[(2R)-2-fluoro-3-hydroxy-3-methylbutyl]-4-(methylamino)pyridine-3-carboxamide, DIMETHYL SULFOXIDE, Interleukin-1 receptor-associated kinase 4 | | Authors: | Ferrao, R, Lansdon, E.B. | | Deposit date: | 2025-02-11 | | Release date: | 2025-05-28 | | Last modified: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Discovery of Edecesertib (GS-5718): A Potent, Selective Inhibitor of IRAK4.
J.Med.Chem., 68, 2025
|
|
9NA3
 
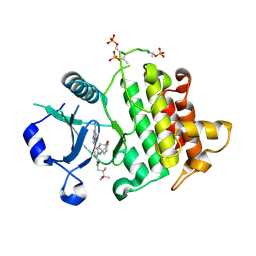 | | IRAK4 in Complex with Compound 15 | | Descriptor: | (6P)-6-[(8R)-3-cyanopyrrolo[1,2-b]pyridazin-7-yl]-N-[(2S)-2-fluoro-3-hydroxy-3-methylbutyl]-4-[(4-hydroxybicyclo[2.2.2]octan-1-yl)amino]pyridine-3-carboxamide, Interleukin-1 receptor-associated kinase 4 | | Authors: | Ferrao, R, Lansdon, E.B. | | Deposit date: | 2025-02-11 | | Release date: | 2025-05-28 | | Last modified: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Discovery of Edecesertib (GS-5718): A Potent, Selective Inhibitor of IRAK4.
J.Med.Chem., 68, 2025
|
|
9NA4
 
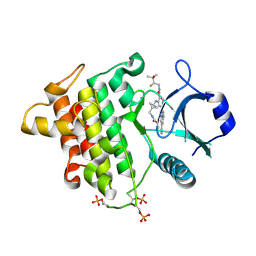 | | IRAK4 in Complex with Compound 18 | | Descriptor: | Interleukin-1 receptor-associated kinase 4, methyl {[(1S,3s)-3-{[(2P)-2-[(8R)-3-cyanopyrrolo[1,2-b]pyridazin-7-yl]-5-{[(2S)-2-fluoro-3-hydroxy-3-methylbutyl]carbamoyl}pyridin-4-yl]amino}cyclobutyl]methyl}carbamate | | Authors: | Ferrao, R, Lansdon, E.B. | | Deposit date: | 2025-02-11 | | Release date: | 2025-05-28 | | Last modified: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (2.33 Å) | | Cite: | Discovery of Edecesertib (GS-5718): A Potent, Selective Inhibitor of IRAK4.
J.Med.Chem., 68, 2025
|
|
9NA6
 
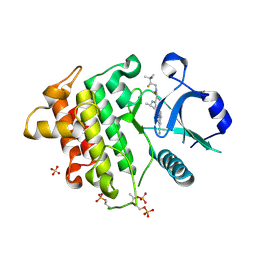 | | IRAK4 in Complex with Compound 34 | | Descriptor: | (6P)-4-{[(1S)-1-cyanoethyl]amino}-6-[(8S)-3-cyanopyrrolo[1,2-b]pyridazin-7-yl]-N-[(2S)-2-fluoro-3-hydroxy-3-methylbutyl]pyridine-3-carboxamide, Interleukin-1 receptor-associated kinase 4, SULFATE ION | | Authors: | Ferrao, R, Lansdon, E.B. | | Deposit date: | 2025-02-11 | | Release date: | 2025-05-28 | | Last modified: | 2025-06-18 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | Discovery of Edecesertib (GS-5718): A Potent, Selective Inhibitor of IRAK4.
J.Med.Chem., 68, 2025
|
|
9QMP
 
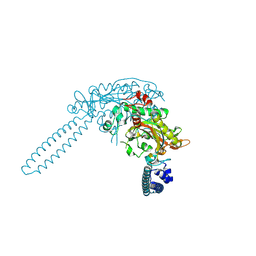 | | E.coli seryl-tRNA synthetase | | Descriptor: | Serine--tRNA ligase | | Authors: | Cusack, S. | | Deposit date: | 2025-03-24 | | Release date: | 2025-05-14 | | Method: | X-RAY DIFFRACTION (2.48 Å) | | Cite: | A second class of synthetase structure revealed by X-ray analysis of Escherichia coli seryl-tRNA synthetase at 2.5 A.
Nature, 347, 1990
|
|
4FE1
 
 | | Improving the Accuracy of Macromolecular Structure Refinement at 7 A Resolution | | Descriptor: | 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE, 1,2-DISTEAROYL-MONOGALACTOSYL-DIGLYCERIDE, BETA-CAROTENE, ... | | Authors: | Fromme, R, Adams, P.D, Fromme, P, Levitt, M, Schroeder, G.F, Brunger, A.T. | | Deposit date: | 2012-05-29 | | Release date: | 2012-08-15 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (4.9228 Å) | | Cite: | Improving the accuracy of macromolecular structure refinement at 7 A resolution.
Structure, 20, 2012
|
|
7SBG
 
 | |
