9IS4
 
 | | Cryo-EM structure of a TEF30-associated intermediate C2S-type PSII-LHCII supercomplex from Chlamydomonas reinhardtii | | Descriptor: | (1R,3R)-6-{(3E,5E,7E,9E,11E,13E,15E,17E)-18-[(1S,4R,6R)-4-HYDROXY-2,2,6-TRIMETHYL-7-OXABICYCLO[4.1.0]HEPT-1-YL]-3,7,12,16-TETRAMETHYLOCTADECA-1,3,5,7,9,11,13,15,17-NONAENYLIDENE}-1,5,5-TRIMETHYLCYCLOHEXANE-1,3-DIOL, (3R,3'R,6S)-4,5-DIDEHYDRO-5,6-DIHYDRO-BETA,BETA-CAROTENE-3,3'-DIOL, (3S,5R,6S,3'S,5'R,6'S)-5,6,5',6'-DIEPOXY-5,6,5',6'- TETRAHYDRO-BETA,BETA-CAROTENE-3,3'-DIOL, ... | | Authors: | Wang, Y, Wang, C, Li, A, Liu, Z. | | Deposit date: | 2024-07-16 | | Release date: | 2025-05-21 | | Last modified: | 2025-07-30 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Roles of multiple TEF30-associated intermediate complexes in the repair and reassembly of photosystem II in Chlamydomonas reinhardtii.
Nat.Plants, 11, 2025
|
|
9ISM
 
 | | Cryo-EM structure of MxaF/MxaJ complex | | Descriptor: | Methanol dehydrogenase, alpha subunit, Methanol oxidation protein MxaJ | | Authors: | Sun, J.Q, Gao, F. | | Deposit date: | 2024-07-18 | | Release date: | 2025-07-23 | | Last modified: | 2025-07-30 | | Method: | ELECTRON MICROSCOPY (2.78 Å) | | Cite: | Deciphering the assembly process of PQQ dependent methanol dehydrogenase.
Nat Commun, 16, 2025
|
|
9ISO
 
 | | Cryo-EM structure of MxaF/MxaJ/PQQ complex | | Descriptor: | Methanol dehydrogenase, alpha subunit, Methanol oxidation protein MxaJ, ... | | Authors: | Sun, J.Q, Gao, F. | | Deposit date: | 2024-07-18 | | Release date: | 2025-07-23 | | Last modified: | 2025-07-30 | | Method: | ELECTRON MICROSCOPY (2.81 Å) | | Cite: | Deciphering the assembly process of PQQ dependent methanol dehydrogenase.
Nat Commun, 16, 2025
|
|
9ISZ
 
 | | Structure of Clr4 catalyzing K14-ubiquitinated histone H3 K9 methylation | | Descriptor: | Histone H3.1/H3.2, Histone-lysine N-methyltransferase, H3 lysine-9 specific, ... | | Authors: | Du, Y.X, Liu, L. | | Deposit date: | 2024-07-19 | | Release date: | 2025-04-30 | | Last modified: | 2025-07-30 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Mechanistic insights into the stimulation of the histone H3K9 methyltransferase Clr4 by proximal H3K14 ubiquitination.
Sci Adv, 11, 2025
|
|
9IT4
 
 | | Structure of Clr4 catalyzing histone H3 K9 methylation | | Descriptor: | Histone H3.1/H3.2, Histone-lysine N-methyltransferase, H3 lysine-9 specific, ... | | Authors: | Du, Y.X, Liu, L. | | Deposit date: | 2024-07-19 | | Release date: | 2025-04-30 | | Last modified: | 2025-07-30 | | Method: | X-RAY DIFFRACTION (2.39 Å) | | Cite: | Mechanistic insights into the stimulation of the histone H3K9 methyltransferase Clr4 by proximal H3K14 ubiquitination.
Sci Adv, 11, 2025
|
|
9J6Z
 
 | |
9J7K
 
 | |
9J7L
 
 | |
9JGZ
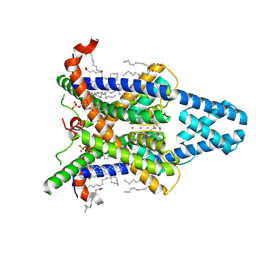 
 | | Cryo-EM Structure of Human Kcnk13. | | Descriptor: | (2R)-1-(hexadecanoyloxy)-3-(phosphonooxy)propan-2-yl (9Z)-octadec-9-enoate, (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(trimethylammonio)ethyl phosphate, ARACHIDONIC ACID, ... | | Authors: | Baobin, L, Ran, Z, Jin, W. | | Deposit date: | 2024-09-08 | | Release date: | 2025-05-07 | | Last modified: | 2025-07-30 | | Method: | ELECTRON MICROSCOPY (2.74 Å) | | Cite: | Gating mechanism of the two-pore-domain potassium channel THIK1.
Nat.Struct.Mol.Biol., 32, 2025
|
|
9JH1
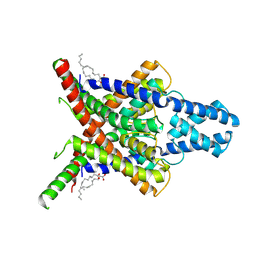 
 | | The Cryo-EM structure of Kcnk13-S136P | | Descriptor: | (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(trimethylammonio)ethyl phosphate, LINOLEIC ACID, POTASSIUM ION, ... | | Authors: | Xinagyun, F, Haichao, J, Jin, W, Ran, Z, Baobin, L. | | Deposit date: | 2024-09-08 | | Release date: | 2025-05-07 | | Last modified: | 2025-07-30 | | Method: | ELECTRON MICROSCOPY (3.07 Å) | | Cite: | Gating mechanism of the two-pore-domain potassium channel THIK1.
Nat.Struct.Mol.Biol., 32, 2025
|
|
9JL0
 
 | | Crystal structure of feline CD8aa homodimer | | Descriptor: | T-cell surface glycoprotein CD8 alpha chain | | Authors: | Li, Z, Liang, R. | | Deposit date: | 2024-09-17 | | Release date: | 2025-07-02 | | Last modified: | 2025-07-30 | | Method: | X-RAY DIFFRACTION (2.48 Å) | | Cite: | Insights into the structural features of a feline CD8 alpha alpha homodimer.
Dev.Comp.Immunol., 169, 2025
|
|
9K2G
 
 | | Human RNA Polymerase III de novo transcribing complex 13 (TC13) | | Descriptor: | 3'-DEOXYADENOSINE-5'-TRIPHOSPHATE, DNA (104-MER), DNA-directed RNA polymerase III subunit RPC1, ... | | Authors: | Wang, Q, Ren, Y, Jin, Q, Chen, X, Xu, Y. | | Deposit date: | 2024-10-17 | | Release date: | 2025-04-16 | | Last modified: | 2025-07-30 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Structural insights into human Pol III transcription initiation in action.
Nature, 643, 2025
|
|
9K36
 
 | | Human RNA Polymerase III de novo transcribing complex 12 (TC12) | | Descriptor: | 3'-DEOXYADENOSINE-5'-TRIPHOSPHATE, DNA (54-MER), DNA-directed RNA polymerase III subunit RPC1, ... | | Authors: | Wang, Q, Ren, Y, Jin, Q, Chen, X, Xu, Y. | | Deposit date: | 2024-10-18 | | Release date: | 2025-04-16 | | Last modified: | 2025-07-30 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Structural insights into human Pol III transcription initiation in action.
Nature, 643, 2025
|
|
9K38
 
 | | Human RNA Polymerase III de novo transcribing complex 6 (TC6) | | Descriptor: | 3'-DEOXYADENOSINE-5'-TRIPHOSPHATE, DNA (51-MER), DNA-directed RNA polymerase III subunit RPC1, ... | | Authors: | Wang, Q, Ren, Y, Jin, Q, Chen, X, Xu, Y. | | Deposit date: | 2024-10-18 | | Release date: | 2025-04-16 | | Last modified: | 2025-07-30 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Structural insights into human Pol III transcription initiation in action.
Nature, 643, 2025
|
|
9K39
 
 | | Human RNA Polymerase III de novo transcribing complex 8 (TC8) | | Descriptor: | 3'-DEOXYADENOSINE-5'-TRIPHOSPHATE, DNA (50-MER), DNA-directed RNA polymerase III subunit RPC1, ... | | Authors: | Wang, Q, Ren, Y, Jin, Q, Chen, X, Xu, Y. | | Deposit date: | 2024-10-18 | | Release date: | 2025-04-16 | | Last modified: | 2025-07-30 | | Method: | ELECTRON MICROSCOPY (2.8 Å) | | Cite: | Structural insights into human Pol III transcription initiation in action.
Nature, 643, 2025
|
|
9K3B
 
 | | Human RNA Polymerase III de novo transcribing complex 10 overall (TC10-overall) | | Descriptor: | 3'-DEOXYADENOSINE-5'-TRIPHOSPHATE, DNA (102-MER), DNA-directed RNA polymerase III subunit RPC1, ... | | Authors: | Wang, Q, Ren, Y, Jin, Q, Chen, X, Xu, Y. | | Deposit date: | 2024-10-18 | | Release date: | 2025-04-16 | | Last modified: | 2025-07-30 | | Method: | ELECTRON MICROSCOPY (4.8 Å) | | Cite: | Structural insights into human Pol III transcription initiation in action.
Nature, 643, 2025
|
|
9K3U
 
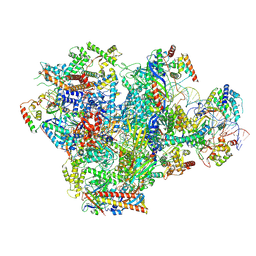 | | Human RNA Polymerase III de novo transcribing complex 5 (TC5) | | Descriptor: | DNA (97-MER), DNA-directed RNA polymerase III subunit RPC1, DNA-directed RNA polymerase III subunit RPC10, ... | | Authors: | Wang, Q, Ren, Y, Jin, Q, Chen, X, Xu, Y. | | Deposit date: | 2024-10-20 | | Release date: | 2025-04-16 | | Last modified: | 2025-07-30 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Structural insights into human Pol III transcription initiation in action.
Nature, 643, 2025
|
|
9K3V
 
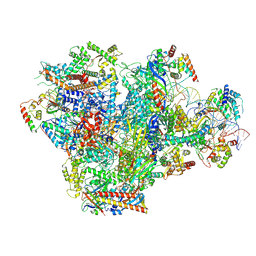 | | Human RNA Polymerase III de novo transcribing complex 4 (TC4) | | Descriptor: | 3'-DEOXYADENOSINE-5'-TRIPHOSPHATE, DNA (96-MER), DNA-directed RNA polymerase III subunit RPC1, ... | | Authors: | Wang, Q, Ren, Y, Jin, Q, Chen, X, Xu, Y. | | Deposit date: | 2024-10-20 | | Release date: | 2025-04-16 | | Last modified: | 2025-07-30 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Structural insights into human Pol III transcription initiation in action.
Nature, 643, 2025
|
|
9K49
 
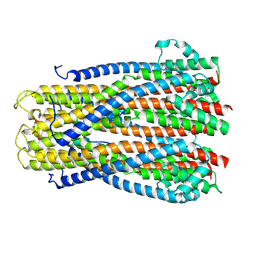 | |
9KCH
 
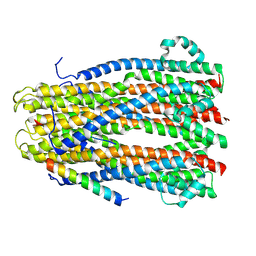 | |
9KDP
 
 | | Crystal structure of monooxygenase PenE | | Descriptor: | Anthrone monooxygenase | | Authors: | Chi, C.B, Ma, M. | | Deposit date: | 2024-11-03 | | Release date: | 2025-07-09 | | Last modified: | 2025-07-30 | | Method: | X-RAY DIFFRACTION (2.592 Å) | | Cite: | Functional Conservation and Divergence of AlpJ-Family Oxygenases Catalyzing C-C Bond Cleavage in Atypical Angucycline Biosynthesis.
Acs Chem.Biol., 20, 2025
|
|
9KN3
 
 | |
9KUF
 
 | | Cryo-EM structure of HsClpP bound to CLPP-2068 | | Descriptor: | 3-[[(7~{R})-2-[(4-bromophenyl)methylamino]-7-methyl-4-oxidanylidene-3,5,7,8-tetrahydropyrido[4,3-d]pyrimidin-6-yl]methyl]benzenecarbonitrile, ATP-dependent Clp protease proteolytic subunit, mitochondrial | | Authors: | Zhao, H, Yuan, Q, Yin, W. | | Deposit date: | 2024-12-03 | | Release date: | 2025-07-16 | | Last modified: | 2025-07-30 | | Method: | ELECTRON MICROSCOPY (2.45 Å) | | Cite: | Harnessing the Magic Methyl Effect: Discovery of CLPP-2068 as a Novel HsClpP Activator for the Treatment of Diffuse Large B-Cell Lymphoma.
J.Med.Chem., 68, 2025
|
|
9L14
 
 | | Crystal structure of the monobody CL-1 in complex with the Escherichia coli adenylate kinase | | Descriptor: | Adenylate kinase, BIS(ADENOSINE)-5'-PENTAPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Nakamura, I, Amesaka, H, Tanaka, S.I, Matsuo, T. | | Deposit date: | 2024-12-13 | | Release date: | 2025-06-04 | | Last modified: | 2025-07-30 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Binding mechanism of adenylate kinase-specific monobodies.
Febs Lett., 599, 2025
|
|
9L2N
 
 | | Crystal structure of Cytochalasin D bound to a filamentous conformation actin | | Descriptor: | (3S,3aR,4S,6S,6aR,7E,10S,12R,13E,15R,15aR)-3-benzyl-6,12-dihydroxy-4,10,12-trimethyl-5-methylidene-1,11-dioxo-2,3,3a,4,5,6,6a,9,10,11,12,15-dodecahydro-1H-cycloundeca[d]isoindol-15-yl acetate, 1,2-ETHANEDIOL, ADENOSINE-5'-DIPHOSPHATE, ... | | Authors: | Takeda, S, Maeda, Y, Fujiwara, I. | | Deposit date: | 2024-12-17 | | Release date: | 2025-07-02 | | Last modified: | 2025-07-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Microscopic and structural observations of actin filament capping and severing by cytochalasin D.
Proc.Natl.Acad.Sci.USA, 122, 2025
|
|
