1R12
 
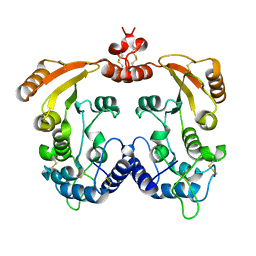 | | Native Aplysia ADP ribosyl cyclase | | Descriptor: | ADP-ribosyl cyclase | | Authors: | Love, M.L, Szebenyi, D.M.E, Kriksunov, I.A, Thiel, D.J, Munshi, C, Graeff, R, Lee, H.C, Hao, Q. | | Deposit date: | 2003-09-23 | | Release date: | 2004-03-09 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | ADP-ribosyl cyclase; crystal structures reveal a covalent intermediate.
Structure, 12, 2004
|
|
1R13
 
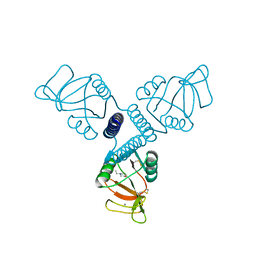 | | Carbohydrate recognition and neck domains of surfactant protein A (SP-A) | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, CALCIUM ION, Pulmonary surfactant-associated protein A, ... | | Authors: | Head, J.F, Mealy, T.R, McCormack, F.X, Seaton, B.A. | | Deposit date: | 2003-09-23 | | Release date: | 2003-11-18 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of trimeric carbohydrate recognition and neck domains of surfactant protein A
J.Biol.Chem., 278, 2003
|
|
1R14
 
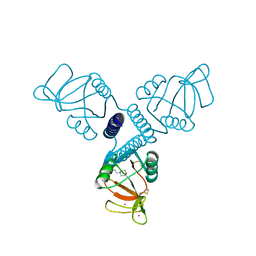 | | Carbohydrate recognition and neck domains of surfactant protein A (Sp-A) containing samarium | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Pulmonary surfactant-associated protein A, SAMARIUM (III) ION | | Authors: | Head, J.F, Mealy, T.R, McCormack, F.X, Seaton, B.A. | | Deposit date: | 2003-09-23 | | Release date: | 2003-11-11 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of trimeric carbohydrate recognition and neck domains of surfactant protein A
J.Biol.Chem., 278, 2003
|
|
1R15
 
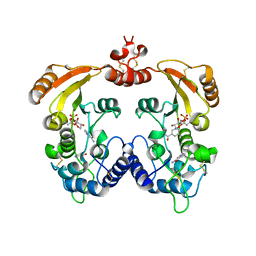 | | Aplysia ADP ribosyl cyclase with bound nicotinamide and R5P | | Descriptor: | ADP-ribosyl cyclase, ANY 5'-MONOPHOSPHATE NUCLEOTIDE, NICOTINAMIDE | | Authors: | Love, M.L, Szebenyi, D.M.E, Kriksunov, I.A, Thiel, D.J, Munshi, C, Graeff, R, Lee, H.C, Hao, Q. | | Deposit date: | 2003-09-23 | | Release date: | 2004-03-09 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | ADP-ribosyl cyclase; crystal structures reveal a covalent intermediate.
Structure, 12, 2004
|
|
1R16
 
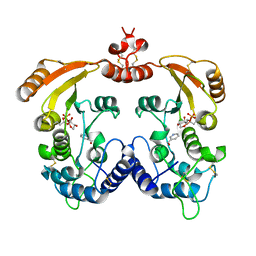 | | Aplysia ADP ribosyl cyclase with bound pyridylcarbinol and R5P | | Descriptor: | 3-PYRIDINYLCARBINOL, ADP-ribosyl cyclase, ANY 5'-MONOPHOSPHATE NUCLEOTIDE | | Authors: | Love, M.L, Szebenyi, D.M.E, Kriksunov, I.A, Thiel, D.J, Munshi, C, Graeff, R, Lee, H.C, Hao, Q. | | Deposit date: | 2003-09-23 | | Release date: | 2004-03-09 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | ADP-ribosyl cyclase; crystal structures reveal a covalent intermediate.
Structure, 12, 2004
|
|
1R17
 
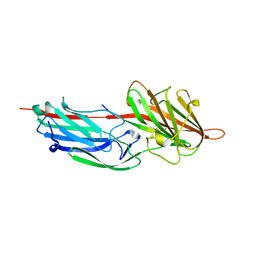 | | Crystal Structure Analysis of S.epidermidis adhesin SdrG binding to Fibrinogen (adhesin-ligand complex) | | Descriptor: | CALCIUM ION, fibrinogen-binding protein SdrG, fibrinopeptide B | | Authors: | Ponnuraj, K, Bowden, M.G, Davis, S, Gurusiddappa, S, Moore, D, Choe, D, Xu, Y, Hook, M, Narayana, S.V.L. | | Deposit date: | 2003-09-23 | | Release date: | 2003-10-28 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | A "dock, lock and latch" Structural Model for a Staphylococcal Adhesin Binding to Fibrinogen
Cell(Cambridge,Mass.), 115, 2003
|
|
1R18
 
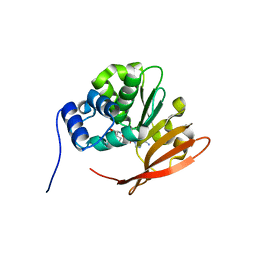 | | Drosophila protein isoaspartyl methyltransferase with S-adenosyl-L-homocysteine | | Descriptor: | Protein-L-isoaspartate(D-aspartate)-O-methyltransferase, S-ADENOSYL-L-HOMOCYSTEINE | | Authors: | Bennett, E.J, Bjerregaard, J, Knapp, J.E, Chavous, D.A, Friedman, A.M, Royer Jr, W.E, O'Connor, C.M. | | Deposit date: | 2003-09-23 | | Release date: | 2003-12-09 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Catalytic implications from the Drosophila protein L-isoaspartyl methyltransferase structure and site-directed mutagenesis.
Biochemistry, 42, 2003
|
|
1R19
 | | Crystal Structure Analysis of S.epidermidis adhesin SdrG binding to Fibrinogen (Apo structure) | | Descriptor: | fibrinogen-binding protein SdrG | | Authors: | Ponnuraj, K, Bowden, M.G, Davis, S, Gurusiddappa, S, Moore, D, Choe, D, Xu, Y, Hook, M, Narayana, S.V.L. | | Deposit date: | 2003-09-23 | | Release date: | 2003-10-28 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (3.51 Å) | | Cite: | A "dock, lock and latch" Structural Model for a Staphylococcal Adhesin Binding to Fibrinogen
Cell(Cambridge,Mass.), 115, 2003
|
|
1R1D
 
 | |
1R1G
 
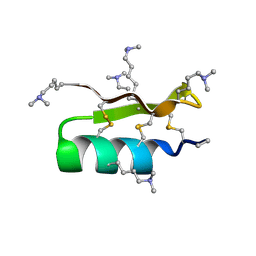 | |
1R1H
 
 | | STRUCTURAL ANALYSIS OF NEPRILYSIN WITH VARIOUS SPECIFIC AND POTENT INHIBITORS | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, N-[3-[(1-AMINOETHYL)(HYDROXY)PHOSPHORYL]-2-(1,1'-BIPHENYL-4-YLMETHYL)PROPANOYL]ALANINE, Neprilysin, ... | | Authors: | Oefner, C, Roques, B.P, Fournie-Zaluski, M.C, Dale, G.E. | | Deposit date: | 2003-09-24 | | Release date: | 2004-09-28 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structural analysis of neprilysin with various specific and potent inhibitors.
Acta Crystallogr.,Sect.D, 60, 2004
|
|
1R1I
 
 | | STRUCTURAL ANALYSIS OF NEPRILYSIN WITH VARIOUS SPECIFIC AND POTENT INHIBITORS | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Neprilysin, ZINC ION, ... | | Authors: | Oefner, C, Roques, B.P, Fournie-Zaluski, M.C, Dale, G.E. | | Deposit date: | 2003-09-24 | | Release date: | 2004-09-28 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural analysis of neprilysin with various specific and potent inhibitors.
Acta Crystallogr.,Sect.D, 60, 2004
|
|
1R1J
 
 | | STRUCTURAL ANALYSIS OF NEPRILYSIN WITH VARIOUS SPECIFIC AND POTENT INHIBITORS | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, N-(3-PHENYL-2-SULFANYLPROPANOYL)PHENYLALANYLALANINE, Neprilysin, ... | | Authors: | Oefner, C, Roques, B.P, Fournie-Zaluski, M.C, Dale, G.E. | | Deposit date: | 2003-09-24 | | Release date: | 2004-09-28 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Structural analysis of neprilysin with various specific and potent inhibitors.
Acta Crystallogr.,Sect.D, 60, 2004
|
|
1R1K
 
 | | Crystal structure of the ligand-binding domains of the heterodimer EcR/USP bound to ponasterone A | | Descriptor: | 2,3,14,20,22-PENTAHYDROXYCHOLEST-7-EN-6-ONE, Ecdysone receptor, L-ALPHA-PHOSPHATIDYL-BETA-OLEOYL-GAMMA-PALMITOYL-PHOSPHATIDYLETHANOLAMINE, ... | | Authors: | Billas, I.M.L, Iwema, T, Garnier, J.-M, Mitschler, A, Rochel, N, Moras, D, Structural Proteomics in Europe (SPINE) | | Deposit date: | 2003-09-24 | | Release date: | 2003-11-18 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structural adaptability in the ligand-binding pocket of the ecdysone hormone receptor.
Nature, 426, 2003
|
|
1R1L
 
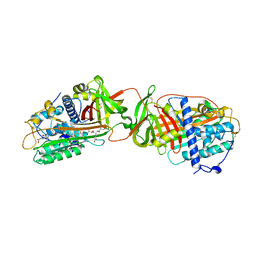 | | Structure of dimeric antithrombin complexed with a P14-P9 reactive loop peptide and an exogenous tripeptide (formyl-norleucine-LF) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Antithrombin P14-P9 peptide, Antithrombin-III, ... | | Authors: | Zhou, A, Huntington, J.A, Lomas, D.A, Stein, P.E, Carrell, R.W. | | Deposit date: | 2003-09-24 | | Release date: | 2004-10-05 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Serpins and the design of peptides to block intermolecular beta-linkages
To be Published
|
|
1R1M
 
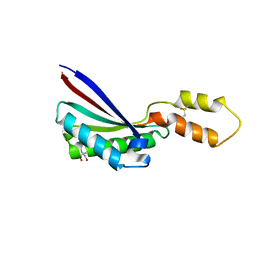 | |
1R1R
 
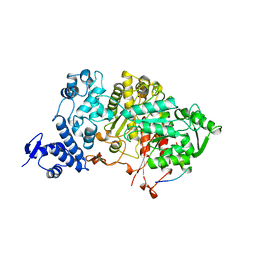 | |
1R1X
 | | Crystal structure of oxy-human hemoglobin Bassett at 2.15 angstrom | | Descriptor: | CARBON MONOXIDE, Hemoglobin alpha chain, Hemoglobin beta chain, ... | | Authors: | Abdulmalik, O, Safo, M.K, Lerner, N.B, Ochotorena, J, Dhaikin, Y, Abraham, D.J, Asakura, T. | | Deposit date: | 2003-09-25 | | Release date: | 2003-10-07 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Characterization of hemoglobin bassett (alpha94Asp-->Ala), a variant with very low oxygen affinity
Am.J.Hematol., 77, 2004
|
|
1R1Y
 
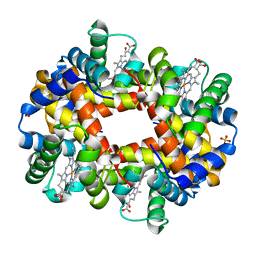 | | Crystal structure of deoxy-human hemoglobin Bassett at 1.8 angstrom | | Descriptor: | Hemoglobin alpha chain, Hemoglobin beta chain, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Abdulmalik, O, Safo, M.K, Lerner, N.B, Ochotorena, J, Dhaikin, Y, Abraham, D.J, Asakura, T. | | Deposit date: | 2003-09-25 | | Release date: | 2003-10-07 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Characterization of hemoglobin bassett (alpha94Asp-->Ala), a variant with very low oxygen affinity
Am.J.Hematol., 77, 2004
|
|
1R1Z
 
 | | The Crystal structure of the Carbohydrate recognition domain of the glycoprotein sorting receptor p58/ERGIC-53 reveals a novel metal binding site and conformational changes associated with calcium ion binding | | Descriptor: | CALCIUM ION, ERGIC-53 protein | | Authors: | Velloso, L.M, Svensson, K, Pettersson, R.F, Lindqvist, Y. | | Deposit date: | 2003-09-25 | | Release date: | 2003-12-16 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | The Crystal Structure of the Carbohydrate-recognition Domain of the Glycoprotein Sorting Receptor p58/ERGIC-53 Reveals an Unpredicted Metal-binding Site and Conformational Changes Associated with Calcium Ion Binding.
J.Mol.Biol., 334, 2003
|
|
1R20
 
 | | Crystal structure of the ligand-binding domains of the heterodimer EcR/USP bound to the synthetic agonist BYI06830 | | Descriptor: | ECDYSONE RECEPTOR, L-ALPHA-PHOSPHATIDYL-BETA-OLEOYL-GAMMA-PALMITOYL-PHOSPHATIDYLETHANOLAMINE, N-(TERT-BUTYL)-3,5-DIMETHYL-N'-[(5-METHYL-2,3-DIHYDRO-1,4-BENZODIOXIN-6-YL)CARBONYL]BENZOHYDRAZIDE, ... | | Authors: | Billas, I.M.L, Iwema, T, Garnier, J.M, Mitschler, A, Rochel, N, Moras, D, Structural Proteomics in Europe (SPINE) | | Deposit date: | 2003-09-25 | | Release date: | 2003-11-18 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural adaptability in the ligand-binding pocket of the ecdysone hormone receptor.
Nature, 426, 2003
|
|
1R24
 
 | | FAB FROM MURINE IGG3 KAPPA | | Descriptor: | PROTEIN (IGG3-KAPPA ANTIBODY (HEAVY CHAIN)), PROTEIN (IGG3-KAPPA ANTIBODY (LIGHT CHAIN)) | | Authors: | Evans, S.V. | | Deposit date: | 1998-11-05 | | Release date: | 1999-11-08 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | The role of homophilic binding in anti-tumor antibody R24 recognition of molecular surfaces. Demonstration of an intermolecular beta-sheet interaction between vh domains.
J.Biol.Chem., 274, 1999
|
|
1R26
 
 | | Crystal structure of thioredoxin from Trypanosoma brucei brucei | | Descriptor: | Thioredoxin | | Authors: | Friemann, R, Schmidt, H, Ramaswamy, S, Forstner, M, Krauth-Siegel, R.L, Eklund, H. | | Deposit date: | 2003-09-26 | | Release date: | 2003-12-09 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structure of thioredoxin from Trypanosoma brucei brucei
FEBS Lett., 554, 2003
|
|
1R28
 
 | | Crystal Structure of the B-Cell Lymphoma 6 (BCL6) BTB domain to 2.2 Angstrom | | Descriptor: | B-cell lymphoma 6 protein | | Authors: | Ahmad, K.F, Melnick, A, Lax, S.A, Bouchard, D, Liu, J, Kiang, C.L, Mayer, S, Licht, J.D, Prive, G.G. | | Deposit date: | 2003-09-26 | | Release date: | 2003-12-23 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Mechanism of SMRT corepressor recruitment by the BCL6 BTB domain.
Mol.Cell, 12, 2003
|
|
1R29
 
 | | Crystal Structure of the B-Cell Lymphoma 6 (BCL6) BTB Domain to 1.3 Angstrom | | Descriptor: | B-cell lymphoma 6 protein | | Authors: | Ahmad, K.F, Melnick, A, Lax, S.A, Bouchard, D, Liu, J, Kiang, C.L, Mayer, S, Licht, J.D, Prive, G.G. | | Deposit date: | 2003-09-26 | | Release date: | 2003-12-23 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Mechanism of SMRT corepressor recruitment by the BCL6 BTB domain.
Mol.Cell, 12, 2003
|
|
