4XQ9
 
 | |
4XQA
 
 | | CRYSTAL STRUCTURE OF AD37 FIBER KNOB IN COMPLEX WITH TRIVALENT SIALIC ACID INHIBITOR ME0462 | | Descriptor: | (1-{2-[bis(2-{4-[({(6R)-5-(acetylamino)-3,5-dideoxy-6-[(1R,2R)-1,2,3-trihydroxypropyl]-beta-L-threo-hex-2-ulopyranonosyl}oxy)methyl]-1H-1,2,3-triazol-1-yl}ethyl)amino]ethyl}-1H-1,2,3-triazol-4-yl)methyl (6R)-5-(acetylamino)-3,5-dideoxy-6-[(1R,2R)-1,2,3-trihydroxypropyl]-beta-L-threo-hex-2-ulopyranosidonic acid, ACETATE ION, Fiber, ... | | Authors: | Stehle, T, Liaci, A.M. | | Deposit date: | 2015-01-19 | | Release date: | 2015-07-29 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.4072 Å) | | Cite: | Triazole linker-based trivalent sialic acid inhibitors of adenovirus type 37 infection of human corneal epithelial cells.
Org.Biomol.Chem., 13, 2015
|
|
4XQB
 
 | | CRYSTAL STRUCTURE OF AD37 FIBER KNOB IN COMPLEX WITH TRIVALENT SIALIC ACID INHIBITOR ME0461 | | Descriptor: | 2-(1-{2-[bis(2-{4-[2-({(6R)-5-(acetylamino)-3,5-dideoxy-6-[(1R,2R)-1,2,3-trihydroxypropyl]-beta-L-threo-hex-2-ulopyranonosyl}oxy)ethyl]-1H-1,2,3-triazol-1-yl}ethyl)amino]ethyl}-1H-1,2,3-triazol-4-yl)ethyl (6R)-5-(acetylamino)-3,5-dideoxy-6-[(1R,2R)-1,2,3-trihydroxypropyl]-beta-L-threo-hex-2-ulopyranosidonic acid, ACETATE ION, Fiber, ... | | Authors: | Stehle, T, Liaci, A.M. | | Deposit date: | 2015-01-19 | | Release date: | 2015-07-29 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.597 Å) | | Cite: | Triazole linker-based trivalent sialic acid inhibitors of adenovirus type 37 infection of human corneal epithelial cells.
Org.Biomol.Chem., 13, 2015
|
|
4XQC
 
 | |
4XQD
 
 | | X-ray structure analysis of xylanase-WT at pH4.0 | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Endo-1,4-beta-xylanase 2, IODIDE ION | | Authors: | Wan, Q, Park, J.M, Riccardi, D.M, Hanson, L.B, Fisher, Z, Smith, J.C, Ostermann, A, Schrader, T, Graham, D.E, Coates, L, Langan, P, Kovalevsky, A.Y. | | Deposit date: | 2015-01-19 | | Release date: | 2015-09-23 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Direct determination of protonation states and visualization of hydrogen bonding in a glycoside hydrolase with neutron crystallography.
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
4XQE
 
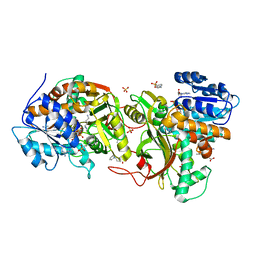 | |
4XQF
 
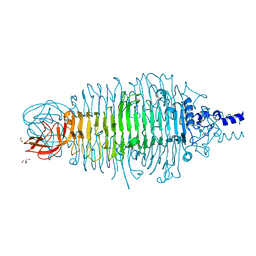 | | Tailspike protein mutant E372Q (delta D470/N471) of E. coli bacteriophage HK620 | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, FORMIC ACID, SODIUM ION, ... | | Authors: | Gohlke, U, Broeker, N.K, Heinemann, U, Seckler, R, Barbirz, S. | | Deposit date: | 2015-01-19 | | Release date: | 2016-01-27 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Enthalpic cost of water removal from a hydrophobic glucose binding cavity on HK620 tailspike protein.
to be published
|
|
4XQG
 
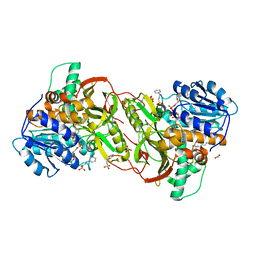 | |
4XQJ
 
 | | Crystal structure of AgrA LytTR domain in complex with promoters | | Descriptor: | 1,2-ETHANEDIOL, Accessory gene regulator A, CALCIUM ION, ... | | Authors: | Gopal, B, Rajasree, K. | | Deposit date: | 2015-01-19 | | Release date: | 2016-04-06 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Conformational features of theStaphylococcus aureusAgrA-promoter interactions rationalize quorum-sensing triggered gene expression.
Biochem Biophys Rep, 6, 2016
|
|
4XQK
 
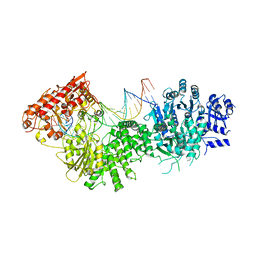 | |
4XQM
 | |
4XQN
 
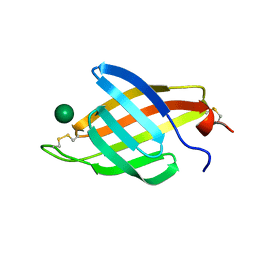
 | | Crystal structure of AgrA LytTR domain in complex with promoters | | Descriptor: | 1,2-ETHANEDIOL, Accessory gene regulator A, DNA (5'-D(*AP*CP*AP*GP*TP*TP*AP*AP*GP*AP*AP*T)-3'), ... | | Authors: | Gopal, B, Rajasree, K. | | Deposit date: | 2015-01-19 | | Release date: | 2016-04-06 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Conformational features of theStaphylococcus aureusAgrA-promoter interactions rationalize quorum-sensing triggered gene expression.
Biochem Biophys Rep, 6, 2016
|
|
4XQO
 
 | | Crystal structure of hemagglutinin from Jiangxi-Donghu (2013) H10N8 influenza virus in complex with 6'-SLN | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Hemagglutinin HA1 chain, ... | | Authors: | Tzarum, N, Zhang, H, Zhu, X, Wilson, I.A. | | Deposit date: | 2015-01-19 | | Release date: | 2015-04-01 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | A Human-Infecting H10N8 Influenza Virus Retains a Strong Preference for Avian-type Receptors.
Cell Host Microbe, 17, 2015
|
|
4XQQ
 
 | | Crystal structure of AgrA LytTR domain in complex with promoters | | Descriptor: | Accessory gene regulator A, DNA (5'-D(*AP*TP*TP*TP*CP*TP*TP*AP*AP*CP*TP*AP*GP*TP*CP*G)-3'), DNA (5'-D(*TP*CP*GP*AP*CP*TP*AP*GP*TP*TP*AP*AP*GP*AP*AP*A)-3') | | Authors: | Gopal, B, Rajasree, K. | | Deposit date: | 2015-01-19 | | Release date: | 2016-04-27 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (3.05 Å) | | Cite: | Conformational features of theStaphylococcus aureusAgrA-promoter interactions rationalize quorum-sensing triggered gene expression.
Biochem Biophys Rep, 6, 2016
|
|
4XQR
 
 | |
4XQS
 
 | |
4XQT
 
 | |
4XQU
 
 | | Crystal structure of hemagglutinin from Jiangxi-Donghu (2013) H10N8 influenza virus in complex with 3'-SLN | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Hemagglutinin HA1, ... | | Authors: | Tzarum, N, Zhang, H, Zhu, X, Wilson, I.A. | | Deposit date: | 2015-01-20 | | Release date: | 2015-04-01 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (3.25 Å) | | Cite: | A Human-Infecting H10N8 Influenza Virus Retains a Strong Preference for Avian-type Receptors.
Cell Host Microbe, 17, 2015
|
|
4XQW
 
 | | X-ray structure analysis of xylanase-N44E with MES at pH6.0 | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Endo-1,4-beta-xylanase 2, IODIDE ION | | Authors: | Wan, Q, Park, J.M, Riccardi, D.M, Hanson, L.B, Fisher, Z, Smith, J.C, Ostermann, A, Schrader, T, Graham, D.E, Coates, L, Langan, P, Kovalevsky, A.Y. | | Deposit date: | 2015-01-20 | | Release date: | 2015-09-23 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Direct determination of protonation states and visualization of hydrogen bonding in a glycoside hydrolase with neutron crystallography.
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
4XQZ
 
 | | Calcium(II) and copper(II) bound to the Z-DNA form of d(CGCGCG), complexed by chloride and MES | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Rohner, M, Medina-Molner, A, Spingler, B. | | Deposit date: | 2015-01-20 | | Release date: | 2016-01-27 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.151 Å) | | Cite: | N,N,O and N,O,N Meridional cis Coordination of Two Guanines to Copper(II) by d(CGCGCG)2.
Inorg.Chem., 55, 2016
|
|
4XR0
 
 | |
4XR1
 
 | |
4XR2
 
 | |
4XR3
 | |
4XR4
 
 | |
