8X40
 | |
8X4F
 | |
8X8T
 
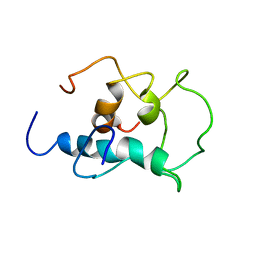 | |
8XEQ
 
 | |
8XGW
 
 | |
8XID
 
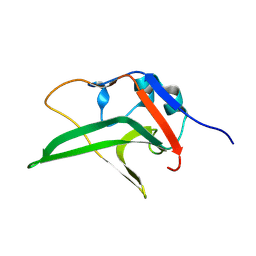 | |
8XTH
 
 | |
8XZ2
 
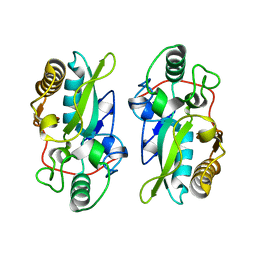 | | The structural model of a homodimeric D-Ala-D-Ala metallopeptidase, VanX, from vancomycin-resistant bacteria | | Descriptor: | D-alanyl-D-alanine dipeptidase | | Authors: | Konuma, T, Takai, T, Tsuchiya, C, Nishida, M, Hashiba, M, Yamada, Y, Shirai, H, Motoda, Y, Nagadoi, A, Chikaishi, E, Akagi, K, Akashi, S, Yamazaki, T, Akutsu, H, Oe, A, Ikegami, T. | | Deposit date: | 2024-01-20 | | Release date: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Analysis of the homodimeric structure of a D-Ala-D-Ala metallopeptidase, VanX, from vancomycin-resistant bacteria.
Protein Sci., 33, 2024
|
|
8Y0F
 
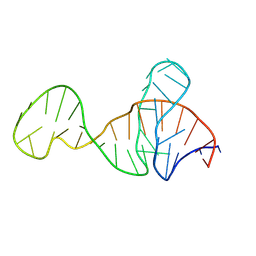 | |
8Y4Z
 
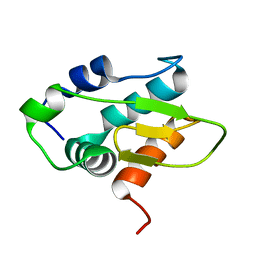 | | Monomeric HERC5 HECT c-lobe structure in solution | | Descriptor: | E3 ISG15--protein ligase HERC5 | | Authors: | Dag, C, Lambert, M, Kahraman, K, Lohn, F, Lee, W, Gocenler, O, Guntert, P, Dotsch, V. | | Deposit date: | 2024-01-31 | | Release date: | 2024-02-14 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Monomeric HERC5 HECT c-lobe structure in solution
To Be Published
|
|
8YSY
 
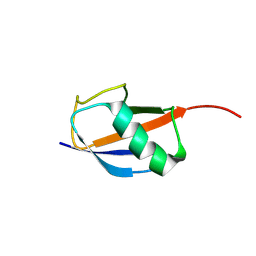 | |
8YUM
 
 | |
8ZUG
 | |
9ATN
 
 | |
9AZI
 | |
9B4V
 
 | | 4F-Trp labeled Oscillatoria Agardhii Agglutinin (OAA) | | Descriptor: | Lectin | | Authors: | Runge, B.R, Zadorozhnyi, R.R, Quinn, C.M, Russell, R.W, Lu, M, Antolinez, S, Struppe, J, Schwieters, C.D, Byeon, I.L, Hadden-Perilla, J, Gronenborn, A.M, Polenova, T. | | Deposit date: | 2024-03-21 | | Release date: | 2024-11-06 | | Method: | SOLID-STATE NMR | | Cite: | Integrating 19 F Distance Restraints for Accurate Protein Structure Determination by Magic Angle Spinning NMR Spectroscopy.
J.Am.Chem.Soc., 2024
|
|
9B4W
 
 | | 4F-Trp labeled Oscillatoria Agardhii Agglutinin (OAA) | | Descriptor: | Lectin | | Authors: | Runge, B.R, Zadorozhnyi, R.R, Quinn, C.M, Russell, R.W, Lu, M, Antolinez, S, Struppe, J, Schwieters, C.D, Byeon, I.L, Hadden-Perilla, J, Gronenborn, A.M, Polenova, T. | | Deposit date: | 2024-03-21 | | Release date: | 2024-11-06 | | Method: | SOLID-STATE NMR | | Cite: | Integrating 19 F Distance Restraints for Accurate Protein Structure Determination by Magic Angle Spinning NMR Spectroscopy.
J.Am.Chem.Soc., 2024
|
|
9BA1
 
 | |
9BAF
 
 | | Solution NMR structure of conofurin-Delta | | Descriptor: | Alpha-conotoxin LvIA | | Authors: | Harvey, P.J, Craik, D.J, Hone, A.J, McIntosh, J.M. | | Deposit date: | 2024-04-04 | | Release date: | 2024-07-03 | | Last modified: | 2024-10-16 | | Method: | SOLUTION NMR | | Cite: | Design, Synthesis, and Structure-Activity Relationships of Novel Peptide Derivatives of the Severe Acute Respiratory Syndrome-Coronavirus-2 Spike-Protein that Potently Inhibit Nicotinic Acetylcholine Receptors.
J.Med.Chem., 67, 2024
|
|
9BB1
 | |
9BB2
 
 | |
9BB3
 
 | |
9BB4
 | |
9BB5
 | |
9BB6
 | |
