8XWR
 
 | |
8XWT
 
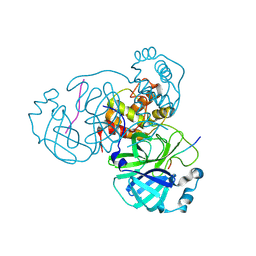 | |
8XY9
 
 | | Crystal structure of SARS-CoV-2 BF.7 RBD and human ACE2 complex | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Processed angiotensin-converting enzyme 2, ... | | Authors: | Lan, J, Wang, C.H. | | Deposit date: | 2024-01-19 | | Release date: | 2025-01-22 | | Last modified: | 2025-04-16 | | Method: | X-RAY DIFFRACTION (3.64 Å) | | Cite: | Receptor binding mechanism and immune evasion capacity of SARS-CoV-2 BQ.1.1 lineage.
Virology, 600, 2024
|
|
8XYE
 
 | | Crystal structure of SARS-CoV-2 BA.4 RBD and human ACE2 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Processed angiotensin-converting enzyme 2, ... | | Authors: | Lan, J, Wang, C.H. | | Deposit date: | 2024-01-19 | | Release date: | 2025-01-22 | | Last modified: | 2025-04-16 | | Method: | X-RAY DIFFRACTION (3.32 Å) | | Cite: | Receptor binding mechanism and immune evasion capacity of SARS-CoV-2 BQ.1.1 lineage.
Virology, 600, 2024
|
|
8XYG
 
 | | Crystal structure of SARS-CoV-2 BQ.1.1 RBD and human ACE2 | | Descriptor: | Processed angiotensin-converting enzyme 2, Spike protein S1 | | Authors: | Lan, J, Wang, C.H. | | Deposit date: | 2024-01-19 | | Release date: | 2025-01-22 | | Last modified: | 2025-04-16 | | Method: | X-RAY DIFFRACTION (3.64 Å) | | Cite: | Receptor binding mechanism and immune evasion capacity of SARS-CoV-2 BQ.1.1 lineage.
Virology, 600, 2024
|
|
8XYO
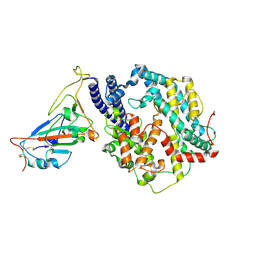 
 | | Cryo-EM structure of CX1 receptor binding domain in complex with human ACE2 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Angiotensin-converting enzyme 2, Spike glycoprotein, ... | | Authors: | Xu, Z.P, Li, L.J, Gu, Y.H, Qi, J.X, Gao, G.F. | | Deposit date: | 2024-01-20 | | Release date: | 2025-01-22 | | Last modified: | 2025-09-03 | | Method: | ELECTRON MICROSCOPY (3.04 Å) | | Cite: | CX1/BtSY2 and BANAL-20-52 exhibit broader receptor binding and higher affinities to multiple animal ACE2 orthologs than SARS-CoV-2 prototype.
J.Virol., 99, 2025
|
|
8YZC
 
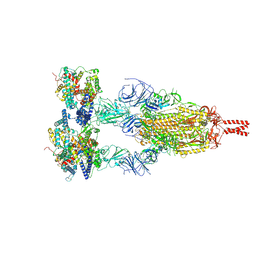 | | Structure of BA.2.86 spike protein in complex with ACE2. | | Descriptor: | Angiotensin-converting enzyme 2, Spike glycoprotein,Fibritin,Expression Tag | | Authors: | Wang, Y.J, Zang, X, Sun, L. | | Deposit date: | 2024-04-06 | | Release date: | 2025-01-22 | | Last modified: | 2025-07-16 | | Method: | ELECTRON MICROSCOPY (2.7 Å) | | Cite: | Lineage-specific pathogenicity, immune evasion, and virological features of SARS-CoV-2 BA.2.86/JN.1 and EG.5.1/HK.3.
Nat Commun, 15, 2024
|
|
8YZE
 
 | | The JN.1 spike protein (S) in complex with ACE2. | | Descriptor: | Angiotensin-converting enzyme 2, Spike glycoprotein,Fibritin,Expression Tag | | Authors: | Wang, Y.J, Zhang, X, Sun, L. | | Deposit date: | 2024-04-06 | | Release date: | 2025-01-22 | | Last modified: | 2025-07-23 | | Method: | ELECTRON MICROSCOPY (3.06 Å) | | Cite: | Lineage-specific pathogenicity, immune evasion, and virological features of SARS-CoV-2 BA.2.86/JN.1 and EG.5.1/HK.3.
Nat Commun, 15, 2024
|
|
9EXA
 | |
9JS4
 
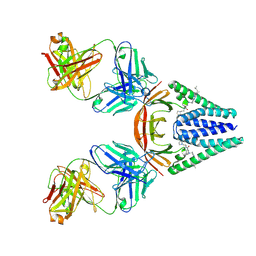
 | | Cryo-EM structure of neutralizing antibody 8G3 in complex with BA.1 RBD | | Descriptor: | Heavy chain of 8G3, Light chain of 8G3, Spike glycoprotein | | Authors: | Li, J, Li, H. | | Deposit date: | 2024-09-30 | | Release date: | 2025-01-22 | | Last modified: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (3.8 Å) | | Cite: | Rapid restoration of potent neutralization activity against the latest Omicron variant JN.1 via AI rational design and antibody engineering.
Proc.Natl.Acad.Sci.USA, 122, 2025
|
|
8YZR
 
 | |
8YZW
 
 | |
8YZZ
 
 | | The structure of HLA-A*2402 complex with peptide from SARS-CoV-2 S448-456 NYNYLYRLF(Prototype) | | Descriptor: | Beta-2-microglobulin, MHC class I antigen, Spike protein S1 | | Authors: | Shang, B.L, Zhang, J.N, Tian, J.M, Liu, J. | | Deposit date: | 2024-04-08 | | Release date: | 2025-01-29 | | Last modified: | 2025-05-28 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | T cell immune evasion by SARS-CoV-2 JN.1 escapees targeting two cytotoxic T cell epitope hotspots.
Nat.Immunol., 26, 2025
|
|
8Z05
 
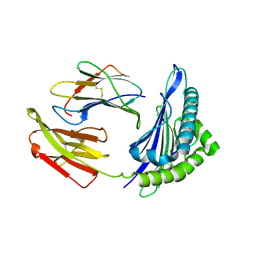 | | The structure of HLA-A*0201 complex with peptide from SARS-CoV-2 N222-230 LLLDRLNKL(BA.2.86/JN.1) | | Descriptor: | Beta-2-microglobulin, HLA class I histocompatibility antigen, A alpha chain, ... | | Authors: | Zhang, J.N, Tian, J.M, Liu, J. | | Deposit date: | 2024-04-09 | | Release date: | 2025-01-29 | | Last modified: | 2025-05-28 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | T cell immune evasion by SARS-CoV-2 JN.1 escapees targeting two cytotoxic T cell epitope hotspots.
Nat.Immunol., 26, 2025
|
|
8Z06
 
 | |
8Z07
 
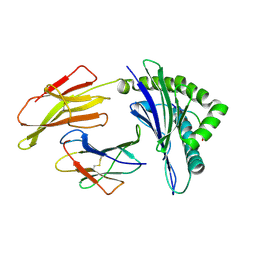 | |
8Z08
 
 | |
8Z6R
 
 | | Structure of XBB.1.16 S trimer with 3 down-RBDs complex with antibody CYFN1006-1. | | Descriptor: | CYFN1006-1 heavy chain, CYFN1006-1 light chain, Spike glycoprotein,Fibritin,Spike glycoprotein,Fibritin,Spike glycoprotein,Fibritin,Expression Tag | | Authors: | Wang, Y.J, Sun, L. | | Deposit date: | 2024-04-19 | | Release date: | 2025-01-29 | | Last modified: | 2025-07-23 | | Method: | ELECTRON MICROSCOPY (2.87 Å) | | Cite: | Structure of XBB.1.16 S trimer with 3 down-RBDs complex with antibody CYFN1006-1.
To Be Published
|
|
8Z6X
 
 | |
9ARU
 
 | | COVA2-15 fragment antigen binding in complex with SARS-CoV-2 6P-mut7 S protein | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, COVA2-15 heavy chain variable region, ... | | Authors: | Ozorowski, G, Turner, H.L, Ward, A.B. | | Deposit date: | 2024-02-23 | | Release date: | 2025-01-29 | | Last modified: | 2025-05-28 | | Method: | ELECTRON MICROSCOPY (3.9 Å) | | Cite: | Plant-produced SARS-CoV-2 antibody engineered towards enhanced potency and in vivo efficacy.
Plant Biotechnol J, 23, 2025
|
|
9G9I
 
 | |
9G9K
 
 | |
8VUO
 
 | |
8Z6A
 
 | |
8Z7B
 
 | |








