[English] 日本語
 Yorodumi
Yorodumi- PDB-5n18: Second Bromodomain (BD2) from Candida albicans Bdf1 bound to an i... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5n18 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
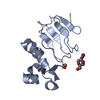
| Title | Second Bromodomain (BD2) from Candida albicans Bdf1 bound to an imidazopyridine (compound 2) | |||||||||
 Components Components | Bromodomain-containing factor 1 | |||||||||
 Keywords Keywords | TRANSCRIPTION / Bromodomain | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of heterochromatin formation / sporulation resulting in formation of a cellular spore / TFIID-class transcription factor complex binding / histone reader activity / core promoter sequence-specific DNA binding / chromatin remodeling / DNA repair / regulation of DNA-templated transcription / chromatin / nucleus Similarity search - Function | |||||||||
| Biological species |  Candida albicans (yeast) Candida albicans (yeast) | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.45 Å MOLECULAR REPLACEMENT / Resolution: 1.45 Å | |||||||||
 Authors Authors | Mietton, F. / Ferri, E. / Champleboux, M. / Zala, N. / Maubon, D. / Zhou, Y. / Harbut, M. / Spittler, D. / Garnaud, C. / Courcon, M. ...Mietton, F. / Ferri, E. / Champleboux, M. / Zala, N. / Maubon, D. / Zhou, Y. / Harbut, M. / Spittler, D. / Garnaud, C. / Courcon, M. / Chauvel, M. / d'Enfert, C. / Kashemirov, B.A. / Hull, M. / Cornet, M. / McKenna, C.E. / Govin, J. / Petosa, C. | |||||||||
| Funding support |  France, 1items France, 1items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2017 Journal: Nat Commun / Year: 2017Title: Selective BET bromodomain inhibition as an antifungal therapeutic strategy. Authors: Mietton, F. / Ferri, E. / Champleboux, M. / Zala, N. / Maubon, D. / Zhou, Y. / Harbut, M. / Spittler, D. / Garnaud, C. / Courcon, M. / Chauvel, M. / d'Enfert, C. / Kashemirov, B.A. / Hull, M. ...Authors: Mietton, F. / Ferri, E. / Champleboux, M. / Zala, N. / Maubon, D. / Zhou, Y. / Harbut, M. / Spittler, D. / Garnaud, C. / Courcon, M. / Chauvel, M. / d'Enfert, C. / Kashemirov, B.A. / Hull, M. / Cornet, M. / McKenna, C.E. / Govin, J. / Petosa, C. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5n18.cif.gz 5n18.cif.gz | 105 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5n18.ent.gz pdb5n18.ent.gz | 79.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5n18.json.gz 5n18.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/n1/5n18 https://data.pdbj.org/pub/pdb/validation_reports/n1/5n18 ftp://data.pdbj.org/pub/pdb/validation_reports/n1/5n18 ftp://data.pdbj.org/pub/pdb/validation_reports/n1/5n18 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  5n13SC  5n15C  5n16C  5n17C S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 12626.337 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Candida albicans (strain SC5314 / ATCC MYA-2876) (yeast) Candida albicans (strain SC5314 / ATCC MYA-2876) (yeast)Strain: SC5314 / ATCC MYA-2876 / Gene: BDF1, CaO19.8593, CaO19.978 / Production host:  #2: Chemical | ChemComp-8HZ / | #3: Chemical | ChemComp-GOL / | #4: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.04 Å3/Da / Density % sol: 39.61 % |
|---|---|
| Crystal grow | Temperature: 277 K / Method: vapor diffusion, hanging drop / pH: 4.6 Details: Crystals were obtained by mixing a solution of 22 mg/mL protein and 1.5 mM inhibitor with 0.1 M sodium acetate (pH 4.6) and 25% (w/v) PEG 3000. |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: MASSIF-1 / Wavelength: 0.966 Å / Beamline: MASSIF-1 / Wavelength: 0.966 Å |
| Detector | Type: DECTRIS PILATUS3 2M / Detector: PIXEL / Date: Apr 1, 2016 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.966 Å / Relative weight: 1 |
| Reflection | Resolution: 1.45→44.307 Å / Num. obs: 34661 / % possible obs: 95.2 % / Redundancy: 2.45 % / CC1/2: 0.998 / Rmerge(I) obs: 0.047 / Rpim(I) all: 0.035 / Net I/σ(I): 12.1 |
| Reflection shell | Resolution: 1.45→1.5 Å / Redundancy: 2.4 % / Rmerge(I) obs: 0.442 / Num. unique obs: 3549 / CC1/2: 0.805 / Rpim(I) all: 0.321 / % possible all: 97.3 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 5N13 Resolution: 1.45→44.307 Å / SU ML: 0.17 / Cross valid method: FREE R-VALUE / σ(F): 1.35 / Phase error: 20.29
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.45→44.307 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller











 PDBj
PDBj




