[English] 日本語
 Yorodumi
Yorodumi- PDB-5lhz: PB3 Domain of Human PLK4 in Complex with Coiled-Coil Domain of STIL -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5lhz | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | PB3 Domain of Human PLK4 in Complex with Coiled-Coil Domain of STIL | |||||||||
 Components Components |
| |||||||||
 Keywords Keywords | STRUCTURAL PROTEIN / Polo box domain / Centriole / Complex / transferase | |||||||||
| Function / homology |  Function and homology information Function and homology informationfloor plate development / de novo centriole assembly involved in multi-ciliated epithelial cell differentiation / procentriole / deuterosome / regulation of centriole replication / procentriole replication complex / positive regulation of centriole replication / embryonic axis specification / trophoblast giant cell differentiation / notochord development ...floor plate development / de novo centriole assembly involved in multi-ciliated epithelial cell differentiation / procentriole / deuterosome / regulation of centriole replication / procentriole replication complex / positive regulation of centriole replication / embryonic axis specification / trophoblast giant cell differentiation / notochord development / polo kinase / microtubule organizing center organization / positive regulation of spindle assembly / XY body / protein localization to centrosome / determination of left/right symmetry / neural tube development / smoothened signaling pathway / heart looping / forebrain development / centriole replication / cleavage furrow / centrosome duplication / microtubule organizing center / cilium assembly / positive regulation of G1/S transition of mitotic cell cycle / Loss of Nlp from mitotic centrosomes / Loss of proteins required for interphase microtubule organization from the centrosome / Recruitment of mitotic centrosome proteins and complexes / Recruitment of NuMA to mitotic centrosomes / Anchoring of the basal body to the plasma membrane / AURKA Activation by TPX2 / mitotic spindle organization / regulation of mitotic spindle organization / neural tube closure / centriole / multicellular organism growth / Regulation of PLK1 Activity at G2/M Transition / cell cortex / in utero embryonic development / protein phosphorylation / ciliary basal body / protein serine kinase activity / protein serine/threonine kinase activity / centrosome / negative regulation of apoptotic process / nucleolus / nucleoplasm / ATP binding / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.51 Å MOLECULAR REPLACEMENT / Resolution: 2.51 Å | |||||||||
 Authors Authors | Cottee, M.A. / Lea, S.M. | |||||||||
| Funding support |  United Kingdom, 2items United Kingdom, 2items
| |||||||||
 Citation Citation |  Journal: Biol Open / Year: 2017 Journal: Biol Open / Year: 2017Title: A key centriole assembly interaction interface between human PLK4 and STIL appears to not be conserved in flies. Authors: Cottee, M.A. / Johnson, S. / Raff, J.W. / Lea, S.M. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5lhz.cif.gz 5lhz.cif.gz | 121.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5lhz.ent.gz pdb5lhz.ent.gz | 96.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5lhz.json.gz 5lhz.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/lh/5lhz https://data.pdbj.org/pub/pdb/validation_reports/lh/5lhz ftp://data.pdbj.org/pub/pdb/validation_reports/lh/5lhz ftp://data.pdbj.org/pub/pdb/validation_reports/lh/5lhz | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  5lhwC  5lhxC  5lhyC  4yypS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 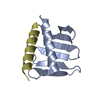
| ||||||||
| 2 | 
| ||||||||
| 3 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 9915.191 Da / Num. of mol.: 3 / Fragment: UNP residues 884-970 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: PLK4, SAK, STK18 / Production host: Homo sapiens (human) / Gene: PLK4, SAK, STK18 / Production host:  #2: Protein/peptide | Mass: 3199.662 Da / Num. of mol.: 3 / Fragment: UNP residues 726-750 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: STIL, SIL / Production host: Homo sapiens (human) / Gene: STIL, SIL / Production host:  #3: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.36 Å3/Da / Density % sol: 47.78 % / Description: Hexagonal rod |
|---|---|
| Crystal grow | Temperature: 292 K / Method: vapor diffusion, sitting drop / pH: 6 Details: 150nl Protein solution (41.87 mg/ml protein in 20 mM Tris pH 7.5, 150 mM NaCl, 2 mM DTT) and 50 nl mother liquor (100mM MES pH 6.0, 191.7 mM Zn acetate, 10% Isopropanol) |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  Diamond Diamond  / Beamline: I02 / Wavelength: 0.9795 Å / Beamline: I02 / Wavelength: 0.9795 Å |
| Detector | Type: DECTRIS PILATUS 6M / Detector: PIXEL / Date: Jan 30, 2016 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9795 Å / Relative weight: 1 |
| Reflection | Resolution: 2.51→36.223 Å / Num. obs: 13265 / % possible obs: 99.7 % / Redundancy: 6.2 % / CC1/2: 0.995 / Net I/σ(I): 12.1 |
| Reflection shell | Highest resolution: 2.51 Å / Rmerge(I) obs: 1.312 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: Model based on 4YYP Resolution: 2.51→36.223 Å / SU ML: 0.43 / Cross valid method: FREE R-VALUE / σ(F): 1.34 / Phase error: 32.21
| ||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å | ||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.51→36.223 Å
| ||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller












 PDBj
PDBj
