+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3sk7 | ||||||
|---|---|---|---|---|---|---|---|
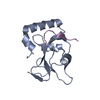
| Title | Crystal Structure of V. cholerae SeqA | ||||||
 Components Components | Protein SeqA | ||||||
 Keywords Keywords | REPLICATION INHIBITOR / Sequestration / Negative regulator / DNA replication initiation / DNA binding | ||||||
| Function / homology | Replication modulator SeqA, C-terminal DNA-binding domain / Replication modulator SeqA, C-terminal DNA-binding domain / Up-down Bundle / Mainly Alpha / :  Function and homology information Function and homology information | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.5 Å MOLECULAR REPLACEMENT / Resolution: 1.5 Å | ||||||
 Authors Authors | Chung, Y.S. / Guarne, A. | ||||||
 Citation Citation |  Journal: To be Published Journal: To be PublishedTitle: Investigating the aggregational property of SeqAvibrio in the absence of the oligomerization domain Authors: Chung, Y.S. / Guarne, A. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3sk7.cif.gz 3sk7.cif.gz | 173.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3sk7.ent.gz pdb3sk7.ent.gz | 136.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3sk7.json.gz 3sk7.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/sk/3sk7 https://data.pdbj.org/pub/pdb/validation_reports/sk/3sk7 ftp://data.pdbj.org/pub/pdb/validation_reports/sk/3sk7 ftp://data.pdbj.org/pub/pdb/validation_reports/sk/3sk7 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  3fmtS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
| ||||||||
| Details | AUTHORS STATE THAT NO QUATERNARY STRUCTURES WERE DETECTED. |
- Components
Components
| #1: Protein | Mass: 13285.413 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   #2: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 2 X-RAY DIFFRACTION / Number of used crystals: 2 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.04 Å3/Da / Density % sol: 39.58 % |
|---|---|
| Crystal grow | Temperature: 277 K / Method: vapor diffusion, hanging drop Details: 30% PEG 4000, VAPOR DIFFUSION, HANGING DROP, temperature 277K |
-Data collection
| Diffraction |
| ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source |
| ||||||||||||||||||
| Detector |
| ||||||||||||||||||
| Radiation |
| ||||||||||||||||||
| Radiation wavelength |
| ||||||||||||||||||
| Reflection | Resolution: 1.46→50 Å / Num. obs: 35671 / % possible obs: 99.6 % / Observed criterion σ(F): 0 / Observed criterion σ(I): -3 / Redundancy: 6.9 % / Rmerge(I) obs: 0.082 / Net I/σ(I): 21.266 | ||||||||||||||||||
| Reflection shell | Resolution: 1.46→1.49 Å / Redundancy: 4.2 % / Rmerge(I) obs: 0.462 / Mean I/σ(I) obs: 4.403 / Rsym value: 0.462 / % possible all: 98.3 |
- Processing
Processing
| Software | Name: PHENIX / Version: (phenix.refine: 1.7_650) / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB Entry 3FMT Resolution: 1.5→34.918 Å / SU ML: 0.18 / Cross valid method: THROUGHOUT / σ(F): 1.33 / Phase error: 17.81 / Stereochemistry target values: Engh & Huber
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 1.01 Å / VDW probe radii: 1.1 Å / Solvent model: FLAT BULK SOLVENT MODEL / Bsol: 29.972 Å2 / ksol: 0.42 e/Å3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.5→34.918 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller
















 PDBj
PDBj

