[English] 日本語
 Yorodumi
Yorodumi- PDB-3pot: Structural analysis of a Ni(III)-methyl species in methyl-coenzym... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3pot | ||||||
|---|---|---|---|---|---|---|---|
| Title | Structural analysis of a Ni(III)-methyl species in methyl-coenzyme M reductase from Methanothermobacter marburgensis | ||||||
 Components Components | (Methyl-coenzyme M reductase I subunit ...) x 3 | ||||||
 Keywords Keywords | TRANSFERASE / METAL-BINDING / NICKEL / METHYL-COENZYME M REDUCTASE / METHANOGENESIS / METHYLATION | ||||||
| Function / homology |  Function and homology information Function and homology informationcoenzyme-B sulfoethylthiotransferase / coenzyme-B sulfoethylthiotransferase activity / methanogenesis / metal ion binding / cytoplasm Similarity search - Function | ||||||
| Biological species |   Methanothermobacter marburgensis (archaea) Methanothermobacter marburgensis (archaea) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  FOURIER SYNTHESIS / Resolution: 1.2 Å FOURIER SYNTHESIS / Resolution: 1.2 Å | ||||||
 Authors Authors | Cedervall, P.E. / Wilmot, C.M. | ||||||
 Citation Citation |  Journal: J.Am.Chem.Soc. / Year: 2011 Journal: J.Am.Chem.Soc. / Year: 2011Title: Structural Analysis of a Ni-Methyl Species in Methyl-Coenzyme M Reductase from Methanothermobacter marburgensis. Authors: Cedervall, P.E. / Dey, M. / Li, X. / Sarangi, R. / Hedman, B. / Ragsdale, S.W. / Wilmot, C.M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3pot.cif.gz 3pot.cif.gz | 1 MB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3pot.ent.gz pdb3pot.ent.gz | 851.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3pot.json.gz 3pot.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/po/3pot https://data.pdbj.org/pub/pdb/validation_reports/po/3pot ftp://data.pdbj.org/pub/pdb/validation_reports/po/3pot ftp://data.pdbj.org/pub/pdb/validation_reports/po/3pot | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  3m2rS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
-Methyl-coenzyme M reductase I subunit ... , 3 types, 6 molecules ADBECF
| #1: Protein | Mass: 60639.664 Da / Num. of mol.: 2 / Source method: isolated from a natural source Source: (natural)   Methanothermobacter marburgensis (archaea) Methanothermobacter marburgensis (archaea)References: UniProt: P11558, coenzyme-B sulfoethylthiotransferase #2: Protein | Mass: 47279.676 Da / Num. of mol.: 2 / Source method: isolated from a natural source Source: (natural)   Methanothermobacter marburgensis (archaea) Methanothermobacter marburgensis (archaea)References: UniProt: P11560, coenzyme-B sulfoethylthiotransferase #3: Protein | Mass: 28797.234 Da / Num. of mol.: 2 / Source method: isolated from a natural source Source: (natural)   Methanothermobacter marburgensis (archaea) Methanothermobacter marburgensis (archaea)References: UniProt: P11562, coenzyme-B sulfoethylthiotransferase |
|---|
-Non-polymers , 9 types, 2626 molecules 
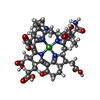
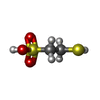
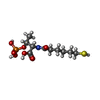











| #4: Chemical | ChemComp-MG / #5: Chemical | #6: Chemical | #7: Chemical | #8: Chemical | #9: Chemical | #10: Chemical | ChemComp-K / | #11: Chemical | ChemComp-EDO / | #12: Water | ChemComp-HOH / | |
|---|
-Details
| Nonpolymer details | 06C HAS REACTED WITH THE ENZYME METAL CENTER TO YIELD A NI-BOUND METHYL GROUP (CH3) (THE NI IS PART ...06C HAS REACTED WITH THE ENZYME METAL CENTER TO YIELD A NI-BOUND METHYL GROUP (CH3) (THE NI IS PART OF COFACTOR F430) AND AN IODIDE (I-) THAT IS SITUATED IN THE ACTIVE SITE AT A DISTANCE OF 4 ANGSTROM FROM CH3 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.16 Å3/Da / Density % sol: 43.17 % |
|---|---|
| Crystal grow | Temperature: 294 K / Method: vapor diffusion, sitting drop / pH: 7.5 Details: 100 mM Hepes-Na pH 7.5, 150 mM magnesium acetate and 20% PEG 400., VAPOR DIFFUSION, SITTING DROP, temperature 294K |
-Data collection
| Diffraction | Mean temperature: 80 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 19-ID / Wavelength: 0.9794 Å / Beamline: 19-ID / Wavelength: 0.9794 Å |
| Detector | Type: ADSC QUANTUM 315r / Detector: CCD / Date: Jun 9, 2009 |
| Radiation | Monochromator: Crystal / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9794 Å / Relative weight: 1 |
| Reflection | Resolution: 1.2→50 Å / Num. all: 721334 / Num. obs: 721334 / % possible obs: 100 % / Observed criterion σ(F): 0 / Observed criterion σ(I): -3 / Redundancy: 3.9 % / Biso Wilson estimate: 13.27 Å2 / Rsym value: 0.075 / Net I/σ(I): 19.5 |
| Reflection shell | Resolution: 1.2→1.24 Å / Redundancy: 3.7 % / Mean I/σ(I) obs: 3.7 / Num. unique all: 72029 / Rsym value: 0.43 / % possible all: 100 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  FOURIER SYNTHESIS FOURIER SYNTHESISStarting model: PDB ID: 3M2R, but with all waters, coenzymes and metal ions removed Resolution: 1.2→50 Å / Cor.coef. Fo:Fc: 0.98 / Cor.coef. Fo:Fc free: 0.972 / SU B: 1.05 / SU ML: 0.022 / Cross valid method: THROUGHOUT / ESU R Free: 0.034 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.4 Å / Solvent model: MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 12.192 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.1779 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.2→50 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.2→1.232 Å / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller











 PDBj
PDBj