+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1mro | ||||||
|---|---|---|---|---|---|---|---|
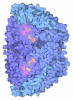
| Title | METHYL-COENZYME M REDUCTASE | ||||||
 Components Components | (METHYL-COENZYME M ...) x 3 | ||||||
 Keywords Keywords | METHANOGENESIS / BIOLOGICAL METHANOGENESIS / NI-ENZYME / OXIDOREDUCTASE | ||||||
| Function / homology |  Function and homology information Function and homology informationcoenzyme-B sulfoethylthiotransferase / coenzyme-B sulfoethylthiotransferase activity / methanogenesis / metal ion binding / cytoplasm Similarity search - Function | ||||||
| Biological species |   Methanothermobacter marburgensis str. Marburg (archaea) Methanothermobacter marburgensis str. Marburg (archaea) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MIR / Resolution: 1.16 Å MIR / Resolution: 1.16 Å | ||||||
 Authors Authors | Ermler, U. / Grabarse, W. | ||||||
 Citation Citation |  Journal: Science / Year: 1997 Journal: Science / Year: 1997Title: Crystal structure of methyl-coenzyme M reductase: the key enzyme of biological methane formation. Authors: Ermler, U. / Grabarse, W. / Shima, S. / Goubeaud, M. / Thauer, R.K. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1mro.cif.gz 1mro.cif.gz | 529 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1mro.ent.gz pdb1mro.ent.gz | 421.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1mro.json.gz 1mro.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/mr/1mro https://data.pdbj.org/pub/pdb/validation_reports/mr/1mro ftp://data.pdbj.org/pub/pdb/validation_reports/mr/1mro ftp://data.pdbj.org/pub/pdb/validation_reports/mr/1mro | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
| ||||||||
| Noncrystallographic symmetry (NCS) | NCS oper: (Code: given Matrix: (0.6268, -0.7779, 0.0458), Vector: |
- Components
Components
-METHYL-COENZYME M ... , 3 types, 6 molecules ADBECF
| #1: Protein | Mass: 60379.289 Da / Num. of mol.: 2 / Source method: isolated from a natural source / Details: MCR-OX-SILENT STATE, ENZYME-SUBSTRATE COMPLEX Source: (natural)   Methanothermobacter marburgensis str. Marburg (archaea) Methanothermobacter marburgensis str. Marburg (archaea)Cellular location: CYTOPLASM / Species: Methanothermobacter marburgensis / Strain: MARBURG References: UniProt: P11558, Oxidoreductases; Acting on a sulfur group of donors #2: Protein | Mass: 47148.477 Da / Num. of mol.: 2 / Source method: isolated from a natural source / Details: MCR-OX-SILENT STATE, ENZYME-SUBSTRATE COMPLEX Source: (natural)   Methanothermobacter marburgensis str. Marburg (archaea) Methanothermobacter marburgensis str. Marburg (archaea)Cellular location: CYTOPLASM / Species: Methanothermobacter marburgensis / Strain: MARBURG References: UniProt: P11560, Oxidoreductases; Acting on a sulfur group of donors #3: Protein | Mass: 28536.926 Da / Num. of mol.: 2 / Source method: isolated from a natural source / Details: MCR-OX-SILENT STATE, ENZYME-SUBSTRATE COMPLEX Source: (natural)   Methanothermobacter marburgensis str. Marburg (archaea) Methanothermobacter marburgensis str. Marburg (archaea)Cellular location: CYTOPLASM / Species: Methanothermobacter marburgensis / Strain: MARBURG References: UniProt: P11562, Oxidoreductases; Acting on a sulfur group of donors |
|---|
-Non-polymers , 7 types, 1726 molecules 









| #4: Chemical | ChemComp-ZN / | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| #5: Chemical | | #6: Chemical | #7: Chemical | #8: Chemical | ChemComp-GOL / #9: Chemical | #10: Water | ChemComp-HOH / | |
-Details
| Has protein modification | Y |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.9 Å3/Da / Density % sol: 31 % |
|---|---|
| Crystal grow | pH: 7.5 Details: 25% PEG 400 0.2 M NACL 20 MG/ML FINAL PROTEIN CONC 0.2 M MGCL 01.M HEPES PH 7.5 |
| Crystal | *PLUS |
| Crystal grow | *PLUS Method: unknown |
| Components of the solutions | *PLUS Conc.: 25 % / Common name: PEG400 |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  EMBL/DESY, HAMBURG EMBL/DESY, HAMBURG  / Beamline: BW7B / Wavelength: 1.08 / Beamline: BW7B / Wavelength: 1.08 |
| Detector | Type: MAR scanner 345 mm plate / Detector: IMAGE PLATE / Date: Apr 1, 1997 |
| Radiation | Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.08 Å / Relative weight: 1 |
| Reflection | Resolution: 1.16→30 Å / Num. obs: 735767 / % possible obs: 93 % / Redundancy: 3 % / Biso Wilson estimate: 10.1 Å2 / Rmerge(I) obs: 0.066 / Rsym value: 0.07 / Net I/σ(I): 18 |
| Reflection shell | Resolution: 1.16→1.23 Å / Redundancy: 2.5 % / Rmerge(I) obs: 0.33 / Mean I/σ(I) obs: 3 / Rsym value: 0.34 / % possible all: 89 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MIR / Resolution: 1.16→30 Å / Rfactor Rfree error: 0.002 / Data cutoff high rms absF: 2798282.65 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0 MIR / Resolution: 1.16→30 Å / Rfactor Rfree error: 0.002 / Data cutoff high rms absF: 2798282.65 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 17.4 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.16→30 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.16→1.23 Å / Rfactor Rfree error: 0.009 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: CNS / Version: 0.3 / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller













 PDBj
PDBj