[English] 日本語
 Yorodumi
Yorodumi- PDB-2onx: NNQQ peptide corresponding to residues 8-11 of yeast prion sup35 ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2onx | ||||||
|---|---|---|---|---|---|---|---|
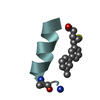
| Title | NNQQ peptide corresponding to residues 8-11 of yeast prion sup35 (alternate crystal form) | ||||||
 Components Components | peptide corresponding to residues 8-11 of yeast prion sup35 | ||||||
 Keywords Keywords | PROTEIN FIBRIL / steric zipper / beta sheets | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.52 Å MOLECULAR REPLACEMENT / Resolution: 1.52 Å | ||||||
 Authors Authors | Sawaya, M.R. / Sambashivan, S. / Nelson, R. / Ivanova, M. / Sievers, S.A. / Apostol, M.I. / Thompson, M.J. / Balbirnie, M. / Wiltzius, J.J. / McFarlane, H. ...Sawaya, M.R. / Sambashivan, S. / Nelson, R. / Ivanova, M. / Sievers, S.A. / Apostol, M.I. / Thompson, M.J. / Balbirnie, M. / Wiltzius, J.J. / McFarlane, H. / Madsen, A.O. / Riekel, C. / Eisenberg, D. | ||||||
 Citation Citation |  Journal: Nature / Year: 2007 Journal: Nature / Year: 2007Title: Atomic structures of amyloid cross-beta spines reveal varied steric zippers. Authors: Sawaya, M.R. / Sambashivan, S. / Nelson, R. / Ivanova, M.I. / Sievers, S.A. / Apostol, M.I. / Thompson, M.J. / Balbirnie, M. / Wiltzius, J.J. / McFarlane, H.T. / Madsen, A.O. / Riekel, C. / Eisenberg, D. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2onx.cif.gz 2onx.cif.gz | 7.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2onx.ent.gz pdb2onx.ent.gz | 4.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2onx.json.gz 2onx.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/on/2onx https://data.pdbj.org/pub/pdb/validation_reports/on/2onx ftp://data.pdbj.org/pub/pdb/validation_reports/on/2onx ftp://data.pdbj.org/pub/pdb/validation_reports/on/2onx | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2okzC  2ol9C  2olxC  2ommC  2ompC  2omqC  2on9C  2onaC  2onvC  2onwC  1yjpS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| Unit cell |
| ||||||||
| Details | One sheet of the steric zipper can be generated by repeated application of the crystallographic unit cell translation along the "a" unit cell dimension. The second sheet of the steric zipper can be generated by application of the crystallographic operator -X,1/2+Y,1-Z, and repeated unit cell translations of this strand along the "a" unit cell dimension. |
- Components
Components
| #1: Protein/peptide | Mass: 502.478 Da / Num. of mol.: 1 / Fragment: residues 8-11 / Source method: obtained synthetically |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, hanging drop / pH: 5.6 Details: 30-50 mg/mL peptide disolved in water and mixed with an equal volume of reservoir solution consisting of 100 mM trisodium citrate, 20% polyethylene glycol 4000 and 20% isopropanol, pH 5.6, ...Details: 30-50 mg/mL peptide disolved in water and mixed with an equal volume of reservoir solution consisting of 100 mM trisodium citrate, 20% polyethylene glycol 4000 and 20% isopropanol, pH 5.6, VAPOR DIFFUSION, HANGING DROP, temperature 298K |
|---|
-Data collection
| Diffraction | Mean temperature: 100 K | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID13 / Wavelength: 0.9466 / Beamline: ID13 / Wavelength: 0.9466 | ||||||||||||||||||||||||||||||||||||
| Detector | Type: MAR CCD 165 mm / Detector: CCD / Date: Jul 16, 2005 | ||||||||||||||||||||||||||||||||||||
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 0.9466 Å / Relative weight: 1 | ||||||||||||||||||||||||||||||||||||
| Reflection | Resolution: 1.5→90 Å / Num. obs: 259 / % possible obs: 63.8 % / Observed criterion σ(I): 0 / Redundancy: 1.5 % / Biso Wilson estimate: 20.4 Å2 / Rmerge(I) obs: 0.152 / Χ2: 1.15 / Net I/σ(I): 10.1 | ||||||||||||||||||||||||||||||||||||
| Reflection shell |
|
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: pdb entry 1yjp Resolution: 1.52→15.43 Å / Cor.coef. Fo:Fc: 0.968 / Cor.coef. Fo:Fc free: 0.962 / SU B: 2.275 / SU ML: 0.072 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.203 / ESU R Free: 0.146 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.4 Å / Solvent model: MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 12.236 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.52→15.43 Å /
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.52→1.563 Å / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller








 PDBj
PDBj