+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2mxe | ||||||
|---|---|---|---|---|---|---|---|
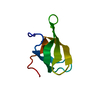
| Title | Solution structure of the C-terminal domain of MvaT | ||||||
 Components Components | MvaT | ||||||
 Keywords Keywords | TRANSCRIPTION REGULATOR | ||||||
| Function / homology | MvaT, DNA-binding domain / Bacterial xenogeneic silencer MvaT, C-terminal domain / negative regulation of single-species biofilm formation on inanimate substrate / negative regulation of secondary metabolite biosynthetic process / DNA-binding transcription activator activity / protein-DNA complex / transcription cis-regulatory region binding / positive regulation of DNA-templated transcription / MvaT Function and homology information Function and homology information | ||||||
| Biological species |  Pseudomonas aeruginosa PAO1 (bacteria) Pseudomonas aeruginosa PAO1 (bacteria) | ||||||
| Method | SOLUTION NMR / simulated annealing | ||||||
| Model details | lowest energy, model1 | ||||||
 Authors Authors | Ding, P. / Xia, B. | ||||||
 Citation Citation |  Journal: Plos Pathog. / Year: 2015 Journal: Plos Pathog. / Year: 2015Title: A Novel AT-Rich DNA Recognition Mechanism for Bacterial Xenogeneic Silencer MvaT. Authors: Ding, P. / McFarland, K.A. / Jin, S. / Tong, G. / Duan, B. / Yang, A. / Hughes, T.R. / Liu, J. / Dove, S.L. / Navarre, W.W. / Xia, B. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2mxe.cif.gz 2mxe.cif.gz | 297.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2mxe.ent.gz pdb2mxe.ent.gz | 248 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2mxe.json.gz 2mxe.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/mx/2mxe https://data.pdbj.org/pub/pdb/validation_reports/mx/2mxe ftp://data.pdbj.org/pub/pdb/validation_reports/mx/2mxe ftp://data.pdbj.org/pub/pdb/validation_reports/mx/2mxe | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 6460.413 Da / Num. of mol.: 1 / Fragment: C-terminal domain (UNP residues 77-124) Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Pseudomonas aeruginosa PAO1 (bacteria) / Strain: PAO1 / Gene: mvaT, PA4315 / Production host: Pseudomonas aeruginosa PAO1 (bacteria) / Strain: PAO1 / Gene: mvaT, PA4315 / Production host:  |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details | Contents: 1 mM [U-100% 13C; U-100% 15N] MvaT, 50 mM sodium phosphate, 50 mM sodium chloride, 90% H2O/10% D2O Solvent system: 90% H2O/10% D2O | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| ||||||||||||||||
| Sample conditions | Ionic strength: 0.1 / pH: 6 / Pressure: ambient / Temperature: 298 K |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software | Name:  Amber AmberDeveloper: Case, Darden, Cheatham, III, Simmerling, Wang, Duke, Luo, ... and Kollman Classification: refinement |
|---|---|
| Refinement | Method: simulated annealing / Software ordinal: 1 |
| NMR constraints | NOE constraints total: 1820 / NOE intraresidue total count: 557 / NOE long range total count: 242 / NOE medium range total count: 177 / NOE sequential total count: 299 / Protein chi angle constraints total count: 7 / Protein other angle constraints total count: 8 / Protein phi angle constraints total count: 32 / Protein psi angle constraints total count: 32 |
| NMR representative | Selection criteria: lowest energy |
| NMR ensemble | Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 100 / Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller










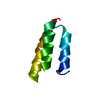


 PDBj
PDBj