+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1mhn | ||||||
|---|---|---|---|---|---|---|---|
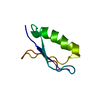
| Title | High resolution crystal structure of the SMN Tudor domain | ||||||
 Components Components | Survival motor neuron protein | ||||||
 Keywords Keywords | RNA BINDING PROTEIN / SMN / SMA / spinal muscular atrophy | ||||||
| Function / homology |  Function and homology information Function and homology informationGemini of Cajal bodies / SMN complex / SMN-Sm protein complex / spliceosomal complex assembly / spliceosomal snRNP assembly / Cajal body / DNA-templated transcription termination / Z disc / cytoplasmic ribonucleoprotein granule / nervous system development ...Gemini of Cajal bodies / SMN complex / SMN-Sm protein complex / spliceosomal complex assembly / spliceosomal snRNP assembly / Cajal body / DNA-templated transcription termination / Z disc / cytoplasmic ribonucleoprotein granule / nervous system development / snRNP Assembly / SARS-CoV-2 modulates host translation machinery / perikaryon / neuron projection / nuclear body / axon / RNA binding / nucleoplasm / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.8 Å MOLECULAR REPLACEMENT / Resolution: 1.8 Å | ||||||
 Authors Authors | Sprangers, R. / Groves, M.R. / Sinning, I. / Sattler, M. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2003 Journal: J.Mol.Biol. / Year: 2003Title: High Resolution X-ray and NMR Structures of the SMN Tudor Domain: conformational variation in the binding site for symmetrically dimethylated arginine residues Authors: Sprangers, R. / Groves, M.R. / Sinning, I. / Sattler, M. #1:  Journal: To be Published Journal: To be PublishedTitle: Definition of domain boundaries and crystallization of the SMN Tudor domain Authors: Sprangers, R. / Selenko, P. / Sattler, M. / Sinning, I. / Groves, M.R. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1mhn.cif.gz 1mhn.cif.gz | 23.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1mhn.ent.gz pdb1mhn.ent.gz | 14.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1mhn.json.gz 1mhn.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/mh/1mhn https://data.pdbj.org/pub/pdb/validation_reports/mh/1mhn ftp://data.pdbj.org/pub/pdb/validation_reports/mh/1mhn ftp://data.pdbj.org/pub/pdb/validation_reports/mh/1mhn | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 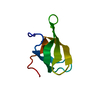
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 6649.454 Da / Num. of mol.: 1 / Fragment: Tudor domain Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: smn1 / Species (production host): Escherichia coli / Production host: Homo sapiens (human) / Gene: smn1 / Species (production host): Escherichia coli / Production host:  |
|---|---|
| #2: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.83 Å3/Da / Density % sol: 32.79 % |
|---|---|
| Crystal grow | Temperature: 285 K / Method: vapor diffusion, hanging drop / pH: 4.5 Details: Amonium Sulphate, pH 4.5, VAPOR DIFFUSION, HANGING DROP, temperature 285K |
| Crystal grow | *PLUS Details: Sprangers, R., (2003) Acta Crystallogr., D59, 366. |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID29 / Wavelength: 0.934 Å / Beamline: ID29 / Wavelength: 0.934 Å |
| Detector | Type: ADSC QUANTUM 4 / Detector: CCD / Date: Dec 1, 2001 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.934 Å / Relative weight: 1 |
| Reflection | Resolution: 1.8→23.95 Å / Num. all: 4489 / Num. obs: 4499 / % possible obs: 100 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 1 / Redundancy: 11.4 % |
| Reflection shell | Resolution: 1.8→1.9 Å / % possible all: 100 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT / Resolution: 1.8→23.947 Å / Isotropic thermal model: Isotropic / σ(F): 0 / Stereochemistry target values: Engh & Huber MOLECULAR REPLACEMENT / Resolution: 1.8→23.947 Å / Isotropic thermal model: Isotropic / σ(F): 0 / Stereochemistry target values: Engh & Huber
| |||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.8→23.947 Å
| |||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.8→1.9 Å | |||||||||||||||||||||||||
| Refinement | *PLUS | |||||||||||||||||||||||||
| Solvent computation | *PLUS | |||||||||||||||||||||||||
| Displacement parameters | *PLUS | |||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller







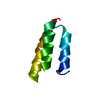



 PDBj
PDBj

