+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7c2p | ||||||
|---|---|---|---|---|---|---|---|
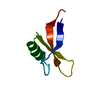
| Title | Structure of Egk Peptide | ||||||
 Components Components | Plant defensing Egk | ||||||
 Keywords Keywords | PROTEIN BINDING / Plant Defensin | ||||||
| Biological species |  Elaeis guineensis (African oil palm) Elaeis guineensis (African oil palm) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.1 Å MOLECULAR REPLACEMENT / Resolution: 2.1 Å | ||||||
 Authors Authors | El Sahili, A. | ||||||
 Citation Citation |  Journal: Acs Pharmacol Transl Sci / Year: 2020 Journal: Acs Pharmacol Transl Sci / Year: 2020Title: Modulation of Lymphocyte Potassium Channel KV1.3 by Membrane-Penetrating, Joint-Targeting Immunomodulatory Plant Defensin. Authors: Ong, S.T. / Bajaj, S. / Tanner, M.R. / Chang, S.C. / Krishnarjuna, B. / Ng, X.R. / Morales, R.A.V. / Chen, M.W. / Luo, D. / Patel, D. / Yasmin, S. / Ng, J.J.H. / Zhuang, Z. / Nguyen, H.M. / ...Authors: Ong, S.T. / Bajaj, S. / Tanner, M.R. / Chang, S.C. / Krishnarjuna, B. / Ng, X.R. / Morales, R.A.V. / Chen, M.W. / Luo, D. / Patel, D. / Yasmin, S. / Ng, J.J.H. / Zhuang, Z. / Nguyen, H.M. / El Sahili, A. / Lescar, J. / Patil, R. / Charman, S.A. / Robins, E.G. / Goggi, J.L. / Tan, P.W. / Sadasivam, P. / Ramasamy, B. / Hartimath, S.V. / Dhawan, V. / Bednenko, J. / Colussi, P. / Wulff, H. / Pennington, M.W. / Kuyucak, S. / Norton, R.S. / Beeton, C. / Chandy, K.G. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7c2p.cif.gz 7c2p.cif.gz | 85.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7c2p.ent.gz pdb7c2p.ent.gz | 67 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7c2p.json.gz 7c2p.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/c2/7c2p https://data.pdbj.org/pub/pdb/validation_reports/c2/7c2p ftp://data.pdbj.org/pub/pdb/validation_reports/c2/7c2p ftp://data.pdbj.org/pub/pdb/validation_reports/c2/7c2p | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  7c31C  4uj0S S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| 3 | 
| ||||||||
| 4 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein/peptide | Mass: 5351.280 Da / Num. of mol.: 4 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Elaeis guineensis (African oil palm) / Production host: Elaeis guineensis (African oil palm) / Production host:  #2: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.22 Å3/Da / Density % sol: 44.57 % |
|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop Details: 0.2 M sodium tartrate dibasic dehydrate, 20% w/v PEG 3350 |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SOLEIL SOLEIL  / Beamline: PROXIMA 2 / Wavelength: 1.000034 Å / Beamline: PROXIMA 2 / Wavelength: 1.000034 Å |
| Detector | Type: DECTRIS EIGER X 9M / Detector: PIXEL / Date: Sep 23, 2017 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.000034 Å / Relative weight: 1 |
| Reflection | Resolution: 2.1→65 Å / Num. obs: 9535 / % possible obs: 88.3 % / Redundancy: 1.7 % / CC1/2: 0.902 / Rmerge(I) obs: 0.16 / Net I/σ(I): 3.11 |
| Reflection shell | Resolution: 2.1→2.22 Å / Num. unique obs: 1392 / CC1/2: 0.67 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 4UJ0 Resolution: 2.1→35.65 Å / Cor.coef. Fo:Fc: 0.927 / Cor.coef. Fo:Fc free: 0.9 / SU R Cruickshank DPI: 0.326 / Cross valid method: THROUGHOUT / SU R Blow DPI: 0.312 / SU Rfree Blow DPI: 0.221 / SU Rfree Cruickshank DPI: 0.226
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 52.06 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.39 Å | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.1→35.65 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.1→2.13 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Refine-ID: X-RAY DIFFRACTION
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
|
 Movie
Movie Controller
Controller










 PDBj
PDBj
