+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1ejg | ||||||
|---|---|---|---|---|---|---|---|
| Title | CRAMBIN AT ULTRA-HIGH RESOLUTION: VALENCE ELECTRON DENSITY. | ||||||
 Components Components | CRAMBIN (PRO22,SER22/LEU25,ILE25) | ||||||
 Keywords Keywords | PLANT PROTEIN / VALENCE ELECTRON DENSITY / MULTI-SUBSTATE / MULTIPOLE REFINEMENT | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |  Crambe hispanica subsp. abyssinica (Abyssinian crambe) Crambe hispanica subsp. abyssinica (Abyssinian crambe) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / Resolution: 0.54 Å SYNCHROTRON / Resolution: 0.54 Å | ||||||
 Authors Authors | Jelsch, C. / Teeter, M.M. / Lamzin, V. / Pichon-Lesme, V. / Blessing, B. / Lecomte, C. | ||||||
 Citation Citation |  Journal: Proc.Natl.Acad.Sci.USA / Year: 2000 Journal: Proc.Natl.Acad.Sci.USA / Year: 2000Title: Accurate protein crystallography at ultra-high resolution: valence electron distribution in crambin. Authors: Jelsch, C. / Teeter, M.M. / Lamzin, V. / Pichon-Pesme, V. / Blessing, R.H. / Lecomte, C. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1ejg.cif.gz 1ejg.cif.gz | 37.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1ejg.ent.gz pdb1ejg.ent.gz | 26.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1ejg.json.gz 1ejg.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ej/1ejg https://data.pdbj.org/pub/pdb/validation_reports/ej/1ejg ftp://data.pdbj.org/pub/pdb/validation_reports/ej/1ejg ftp://data.pdbj.org/pub/pdb/validation_reports/ej/1ejg | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 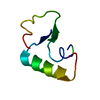
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein/peptide | Mass: 4738.447 Da / Num. of mol.: 1 / Fragment: CRAMBIN / Source method: isolated from a natural source Source: (natural)  Crambe hispanica subsp. abyssinica (Abyssinian crambe) Crambe hispanica subsp. abyssinica (Abyssinian crambe)Species: Crambe hispanica / Strain: subsp. abyssinica / References: UniProt: P01542 |
|---|---|
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.4 Å3/Da / Density % sol: 30 % | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, sitting drop / pH: 7 Details: The protein (40 mg/ml in 80% ethanol) is equilibrated against 60% ethanol. Not buffered., pH 7, VAPOR DIFFUSION, SITTING DROP, temperature 293K | ||||||||||||||||||||
| Crystal grow | *PLUS Method: vapor diffusion / Details: Teeter, M.M., (1993) J. Mol. Biol., 230, 292. | ||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  EMBL/DESY, HAMBURG EMBL/DESY, HAMBURG  / Beamline: BW7A / Wavelength: 0.54 / Beamline: BW7A / Wavelength: 0.54 |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: May 31, 1997 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.54 Å / Relative weight: 1 |
| Reflection | Resolution: 0.54→50 Å / Num. obs: 79868 / % possible obs: 97.6 % / Observed criterion σ(I): 3 / Redundancy: 4.5 % / Biso Wilson estimate: 2.8 Å2 / Rmerge(I) obs: 0.055 / Net I/σ(I): 8.2 |
| Reflection shell | Resolution: 0.54→0.55 Å / Redundancy: 3.8 % / Rmerge(I) obs: 0.1476 / % possible all: 100 |
| Reflection | *PLUS Highest resolution: 0.54 Å / Lowest resolution: 50 Å / Num. all: 112293 / Num. obs: 79869 / Observed criterion σ(I): 3 / Redundancy: 4.5 % / Num. measured all: 489969 |
| Reflection shell | *PLUS Redundancy: 3.8 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 0.54→22.37 Å / σ(F): 0 Details: SHELX97 FOLLOWED BY MOLLY (N.K.HANSEN & P.COPPENS ACTA CRYSTALLOGR. A34, 909-921) REFINEMENT OF ELECTRON DENSITY.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 0.54→22.37 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 0.54→0.57 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file | Serial no: 1 / Param file: SHELXL97 DICTIONARY | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: MOLLY / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS % reflection Rfree: 5 % / Rfactor obs: 0.09 / Rfactor Rwork: 0.09 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS Type: x_angle_deg / Dev ideal: 2.7 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | *PLUS Rfactor Rfree: 0.205 / % reflection Rfree: 5 % / Rfactor Rwork: 0.197 |
 Movie
Movie Controller
Controller










 PDBj
PDBj
