+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20390 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Activated Class III PI-3 Kinase by NRBF2 | |||||||||
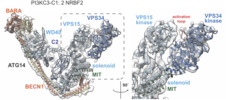
 Map data Map data | Full length NRBF2 pushes PI3KC3-C1 to an activate conformation in which peripheral membrane binding protein VPS34 is posed to phosphorylate substrate phosphatidylinositol. | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
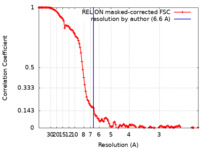
| Method | single particle reconstruction / cryo EM / Resolution: 6.6 Å | |||||||||
 Authors Authors | Young LN / Goerdeler F / Hurley JH | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2019 Journal: Proc Natl Acad Sci U S A / Year: 2019Title: Structural pathway for allosteric activation of the autophagic PI 3-kinase complex I. Authors: Lindsey N Young / Felix Goerdeler / James H Hurley /  Abstract: Autophagy induction by starvation and stress involves the enzymatic activation of the class III phosphatidylinositol (PI) 3-kinase complex I (PI3KC3-C1). The inactive basal state of PI3KC3-C1 is ...Autophagy induction by starvation and stress involves the enzymatic activation of the class III phosphatidylinositol (PI) 3-kinase complex I (PI3KC3-C1). The inactive basal state of PI3KC3-C1 is maintained by inhibitory contacts between the VPS15 protein kinase and VPS34 lipid kinase domains that restrict the conformation of the VPS34 activation loop. Here, the proautophagic MIT domain-containing protein NRBF2 was used to map the structural changes leading to activation. Cryoelectron microscopy was used to visualize a 2-step PI3KC3-C1 activation pathway driven by NRFB2 MIT domain binding. Binding of a single NRBF2 MIT domain bends the helical solenoid of the VPS15 scaffold, displaces the protein kinase domain of VPS15, and releases the VPS34 kinase domain from the inhibited conformation. Binding of a second MIT stabilizes the VPS34 lipid kinase domain in an active conformation that has an unrestricted activation loop and is poised for access to membranes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20390.map.gz emd_20390.map.gz | 12.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20390-v30.xml emd-20390-v30.xml emd-20390.xml emd-20390.xml | 13.2 KB 13.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_20390_fsc.xml emd_20390_fsc.xml | 12.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_20390.png emd_20390.png | 414.4 KB | ||
| Masks |  emd_20390_msk_1.map emd_20390_msk_1.map | 166.4 MB |  Mask map Mask map | |
| Others |  emd_20390_additional.map.gz emd_20390_additional.map.gz emd_20390_additional_1.map.gz emd_20390_additional_1.map.gz | 155.6 MB 155.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20390 http://ftp.pdbj.org/pub/emdb/structures/EMD-20390 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20390 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20390 | HTTPS FTP |
-Validation report
| Summary document |  emd_20390_validation.pdf.gz emd_20390_validation.pdf.gz | 78.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_20390_full_validation.pdf.gz emd_20390_full_validation.pdf.gz | 77.2 KB | Display | |
| Data in XML |  emd_20390_validation.xml.gz emd_20390_validation.xml.gz | 494 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20390 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20390 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20390 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20390 | HTTPS FTP |
-Related structure data
| Related structure data | C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_20390.map.gz / Format: CCP4 / Size: 166.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20390.map.gz / Format: CCP4 / Size: 166.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Full length NRBF2 pushes PI3KC3-C1 to an activate conformation in which peripheral membrane binding protein VPS34 is posed to phosphorylate substrate phosphatidylinositol. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.137 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_20390_msk_1.map emd_20390_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
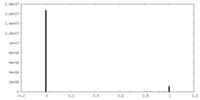
| Density Histograms |
-Additional map: Unsharpened map.
| File | emd_20390_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map. | ||||||||||||
| Projections & Slices |
| ||||||||||||
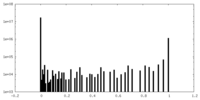
| Density Histograms |
-Additional map: Unsharpened map.
| File | emd_20390_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map. | ||||||||||||
| Projections & Slices |
| ||||||||||||
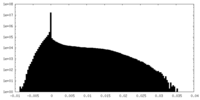
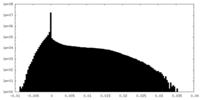
| Density Histograms |
- Sample components
Sample components
-Entire : Class III PI 3-Kinase Complex 1 containing NRBF2-MIT-linker-BECN1...
| Entire | Name: Class III PI 3-Kinase Complex 1 containing NRBF2-MIT-linker-BECN1, ATG14, VPS34, VPS15 |
|---|---|
| Components |
|
-Supramolecule #1: Class III PI 3-Kinase Complex 1 containing NRBF2-MIT-linker-BECN1...
| Supramolecule | Name: Class III PI 3-Kinase Complex 1 containing NRBF2-MIT-linker-BECN1, ATG14, VPS34, VPS15 type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant strain: HEK 293 GNTI / Recombinant cell: Kidney cells / Recombinant plasmid: pCAG vectors Homo sapiens (human) / Recombinant strain: HEK 293 GNTI / Recombinant cell: Kidney cells / Recombinant plasmid: pCAG vectors |
| Molecular weight | Theoretical: 400 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.2 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 88 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average exposure time: 6.4 sec. / Average electron dose: 0.98 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)