[English] 日本語
 Yorodumi
Yorodumi- EMDB-0392: Structure of bacteriophage T7 leading-strand DNA polymerase (D5A/... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0392 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of bacteriophage T7 leading-strand DNA polymerase (D5A/E7A)/Trx and of gp4 (E343Q) bound to a DNA fork (Lead2) | |||||||||
 Map data Map data | T7 DNA polymerase/Trx interacting with C-terminal tail of helicase domains E of T7 helicase (gp4) in complex with a fork DNA substrate and dTTP (from gp4-gp5 leading-strand complex Lead2) | |||||||||
 Sample Sample |
| |||||||||
| Biological species |   Enterobacteria phage T7 (virus) / Enterobacteria phage T7 (virus) /  | |||||||||
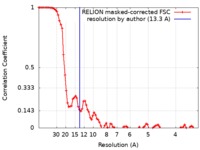
| Method | single particle reconstruction / cryo EM / Resolution: 13.3 Å | |||||||||
 Authors Authors | Gao Y / Fox T / Val N / Yang W | |||||||||
 Citation Citation |  Journal: Science / Year: 2019 Journal: Science / Year: 2019Title: Structures and operating principles of the replisome. Authors: Yang Gao / Yanxiang Cui / Tara Fox / Shiqiang Lin / Huaibin Wang / Natalia de Val / Z Hong Zhou / Wei Yang /  Abstract: Visualization in atomic detail of the replisome that performs concerted leading- and lagging-DNA strand synthesis at a replication fork has not been reported. Using bacteriophage T7 as a model ...Visualization in atomic detail of the replisome that performs concerted leading- and lagging-DNA strand synthesis at a replication fork has not been reported. Using bacteriophage T7 as a model system, we determined cryo-electron microscopy structures up to 3.2-angstroms resolution of helicase translocating along DNA and of helicase-polymerase-primase complexes engaging in synthesis of both DNA strands. Each domain of the spiral-shaped hexameric helicase translocates sequentially hand-over-hand along a single-stranded DNA coil, akin to the way AAA+ ATPases (adenosine triphosphatases) unfold peptides. Two lagging-strand polymerases are attached to the primase, ready for Okazaki fragment synthesis in tandem. A β hairpin from the leading-strand polymerase separates two parental DNA strands into a T-shaped fork, thus enabling the closely coupled helicase to advance perpendicular to the downstream DNA duplex. These structures reveal the molecular organization and operating principles of a replisome. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0392.map.gz emd_0392.map.gz | 4.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0392-v30.xml emd-0392-v30.xml emd-0392.xml emd-0392.xml | 18 KB 18 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_0392_fsc.xml emd_0392_fsc.xml | 8.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_0392.png emd_0392.png | 73.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0392 http://ftp.pdbj.org/pub/emdb/structures/EMD-0392 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0392 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0392 | HTTPS FTP |
-Related structure data
| Related structure data |  0357C  0359C  0362C  0363C  0364C  0365C  0379C  0380C  0381C  0382C  0386C  0387C  0388C  0389C  0390C  0391C  0393C  0394C  0395C  6n7iC  6n7nC  6n7sC  6n7tC  6n7vC  6n7wC  6n9uC  6n9vC  6n9wC  6n9xC C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_0392.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0392.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | T7 DNA polymerase/Trx interacting with C-terminal tail of helicase domains E of T7 helicase (gp4) in complex with a fork DNA substrate and dTTP (from gp4-gp5 leading-strand complex Lead2) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.72 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : T7 DNA polymerase/Trx interacting with C-terminal tail of helicas...
| Entire | Name: T7 DNA polymerase/Trx interacting with C-terminal tail of helicase domains E of T7 helicase (gp4) in complex with a fork DNA substrate and dTTP (from gp4-gp5 leading-strand complex Lead2) |
|---|---|
| Components |
|
-Supramolecule #1: T7 DNA polymerase/Trx interacting with C-terminal tail of helicas...
| Supramolecule | Name: T7 DNA polymerase/Trx interacting with C-terminal tail of helicase domains E of T7 helicase (gp4) in complex with a fork DNA substrate and dTTP (from gp4-gp5 leading-strand complex Lead2) type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Enterobacteria phage T7 (virus) Enterobacteria phage T7 (virus) |
| Recombinant expression | Organism:  |
-Macromolecule #1: gene product 4 helicase-primase
| Macromolecule | Name: gene product 4 helicase-primase / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Enterobacteria phage T7 (virus) Enterobacteria phage T7 (virus) |
| Sequence | String: MDNSHDSDSV FLYHIPCDNC GSSDGNSLFS DGHTFCYVCE KWTAGNEDTK ERASKRKPSG GKPMTYNVW NFGESNGRYS ALTARGISKE TCQKAGYWIA KVDGVMYQVA DYRDQNGNIV S QKVRDKDK NFKTTGSHKS DALFGKHLWN GGKKIVVTEG EIDMLTVMEL ...String: MDNSHDSDSV FLYHIPCDNC GSSDGNSLFS DGHTFCYVCE KWTAGNEDTK ERASKRKPSG GKPMTYNVW NFGESNGRYS ALTARGISKE TCQKAGYWIA KVDGVMYQVA DYRDQNGNIV S QKVRDKDK NFKTTGSHKS DALFGKHLWN GGKKIVVTEG EIDMLTVMEL QDCKYPVVSL GH GASAAKK TCAANYEYFD QFEQIILMFD MDEAGRKAVE EAAQVLPAGK VRVAVLPCKD ANE CHLNGH DREIMEQVWN AGPWIPDGVV SALSLRERIR EHLSSEESVG LLFSGCTGIN DKTL GARGG EVIMVTSGSG MGKSTFVRQQ ALQWGTAMGK KVGLAMLQES VEETAEDLIG LHNRV RLRQ SDSLKREIIE NGKFDQWFDE LFGNDTFHLY DSFAEAETDR LLAKLAYMRS GLGCDV IIL DHISIVVSAS GESDERKMID NLMTKLKGFA KSTGVVLVVI CHLKNPDKGK AHEEGRP VS ITDLRGSGAL RQLSDTIIAL ERNQQGDMPN LVLVRILKCR FTGDTGIAGY MEYNKETG W LEPSSYSGEE ESHSESTDWS NDTDF |
-Macromolecule #2: gene product 5 DNA polymerase
| Macromolecule | Name: gene product 5 DNA polymerase / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Enterobacteria phage T7 (virus) Enterobacteria phage T7 (virus) |
| Sequence | String: MIVSAIAANA LLESVTKFHC GVIYDYSTAE YVSYRPSDFG AYLDALEAEV ARGGLIVFHN GHKYDVPAL TKLAKLQLNR EFHLPRENCI DTLVLSRLIH SNLKDTDMGL LRSGKLPGKR F GSHALEAW GYRLGEMKGE YKDDFKRMLE EQGEEYVDGM EWWNFNEEMM ...String: MIVSAIAANA LLESVTKFHC GVIYDYSTAE YVSYRPSDFG AYLDALEAEV ARGGLIVFHN GHKYDVPAL TKLAKLQLNR EFHLPRENCI DTLVLSRLIH SNLKDTDMGL LRSGKLPGKR F GSHALEAW GYRLGEMKGE YKDDFKRMLE EQGEEYVDGM EWWNFNEEMM DYNVQDVVVT KA LLEKLLS DKHYFPPEID FTDVGYTTFW SESLEAVDIE HRAAWLLAKQ ERNGFPFDTK AIE ELYVEL AARRSELLRK LTETFGSWYQ PKGGTEMFCH PRTGKPLPKY PRIKTPKVGG IFKK PKNKA QREGREPCEL DTREYVAGAP YTPVEHVVFN PSSRDHIQKK LQEAGWVPTK YTDKG APVV DDEVLEGVRV DDPEKQAAID LIKEYLMIQK RIGQSAEGDK AWLRYVAEDG KIHGSV NPN GAVTGRATHA FPNLAQIPGV RSPYGEQCRA AFGAEHHLDG ITGKPWVQAG IDASGLE LR CLAHFMARFD NGEYAHEILN GDIHTKNQIA AELPTRDNAK TFIYGFLYGA GDEKIGQI V GAGKERGKEL KKKFLENTPA IAALRESIQQ TLVESSQWVA GEQQVKWKRR WIKGLDGRK VHVRSPHAAL NTLLQSAGAL ICKLWIIKTE EMLVEKGLKH GWDGDFAYMA WVHDEIQVGC RTEEIAQVV IETAQEAMRW VGDHWNFRCL LDTEGKMGPN WAICH |
-Macromolecule #3: Thioredoxin 1
| Macromolecule | Name: Thioredoxin 1 / type: protein_or_peptide / ID: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: MSDKIIHLTD DSFDTDVLKA DGAILVDFWA EWCGPCKMIA PILDEIADEY QGKLTVAKLN IDQNPGTAP KYGIRGIPTL LLFKNGEVAA TKVGALSKGQ LKEFLDANLA |
-Macromolecule #4: Primer
| Macromolecule | Name: Primer / type: dna / ID: 4 / Classification: DNA |
|---|---|
| Source (natural) | Organism:   Enterobacteria phage T7 (virus) Enterobacteria phage T7 (virus) |
| Sequence | String: (DG)(DG)(DT)(DA)(DC)(DA)(DA)(DC)(DT)(DT) (DG)(DA)(DC)(DG)(DA)(DC)(DA)(DT)(DA)(DG) (DC)(DG)(DT)(DG)(DOA) |
-Macromolecule #5: Template
| Macromolecule | Name: Template / type: dna / ID: 5 / Classification: DNA |
|---|---|
| Source (natural) | Organism:   Enterobacteria phage T7 (virus) Enterobacteria phage T7 (virus) |
| Sequence | String: (DT)(DT)(DT)(DG)(DG)(DT)(DC)(DA)(DT)(DT) (DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT) (DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DA)(DC) (DG)(DG)(DA)(DG)(DT)(DC)(DG)(DT)(DT)(DT) (DC)(DG)(DA)(DC)(DT)(DC)(DC) ...String: (DT)(DT)(DT)(DG)(DG)(DT)(DC)(DA)(DT)(DT) (DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT) (DT)(DT)(DT)(DT)(DT)(DT)(DT)(DT)(DA)(DC) (DG)(DG)(DA)(DG)(DT)(DC)(DG)(DT)(DT)(DT) (DC)(DG)(DA)(DC)(DT)(DC)(DC)(DG)(DT)(DT) (DA)(DT)(DC)(DA)(DC)(DG)(DC)(DT)(DA)(DT) (DG)(DT)(DC)(DG)(DT)(DC)(DA)(DA)(DG)(DT) (DT)(DG)(DT)(DA)(DC)(DC) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Details: unspecified | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK I |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)