+ Open data
Open data
- Basic information
Basic information
| Entry | Database: SASBDB / ID: SASDCW3 |
|---|---|
 Sample Sample | Proline utilization A from Legionella pneumophila 8 mg/mL
|
| Function / homology |  Function and homology information Function and homology informationproline dehydrogenase / proline dehydrogenase activity / L-glutamate gamma-semialdehyde dehydrogenase / L-glutamate gamma-semialdehyde dehydrogenase activity / : / cytoplasmic side of plasma membrane / DNA-binding transcription factor activity / DNA binding Similarity search - Function |
| Biological species |  Legionella pneumophila subsp. pneumophila (strain Philadelphia 1 / ATCC 33152 / DSM 7513) (bacteria) Legionella pneumophila subsp. pneumophila (strain Philadelphia 1 / ATCC 33152 / DSM 7513) (bacteria) |
 Citation Citation |  Journal: FEBS J / Year: 2017 Journal: FEBS J / Year: 2017Title: Biophysical investigation of type A PutAs reveals a conserved core oligomeric structure. Authors: David A Korasick / Harkewal Singh / Travis A Pemberton / Min Luo / Richa Dhatwalia / John J Tanner /  Abstract: Many enzymes form homooligomers, yet the functional significance of self-association is seldom obvious. Herein, we examine the connection between oligomerization and catalytic function for proline ...Many enzymes form homooligomers, yet the functional significance of self-association is seldom obvious. Herein, we examine the connection between oligomerization and catalytic function for proline utilization A (PutA) enzymes. PutAs are bifunctional enzymes that catalyze both reactions of proline catabolism. Type A PutAs are the smallest members of the family, possessing a minimal domain architecture consisting of N-terminal proline dehydrogenase and C-terminal l-glutamate-γ-semialdehyde dehydrogenase modules. Type A PutAs form domain-swapped dimers, and in one case (Bradyrhizobium japonicum PutA), two of the dimers assemble into a ring-shaped tetramer. Whereas the dimer has a clear role in substrate channeling, the functional significance of the tetramer is unknown. To address this question, we performed structural studies of four-type A PutAs from two clades of the PutA tree. The crystal structure of Bdellovibrio bacteriovorus PutA covalently inactivated by N-propargylglycine revealed a fold and substrate-channeling tunnel similar to other PutAs. Small-angle X-ray scattering (SAXS) and analytical ultracentrifugation indicated that Bdellovibrio PutA is dimeric in solution, in contrast to the prediction from crystal packing of a stable tetrameric assembly. SAXS studies of two other type A PutAs from separate clades also suggested that the dimer predominates in solution. To assess whether the tetramer of B. japonicum PutA is necessary for catalytic function, a hot spot disruption mutant that cleanly produces dimeric protein was generated. The dimeric variant exhibited kinetic parameters similar to the wild-type enzyme. These results implicate the domain-swapped dimer as the core structural and functional unit of type A PutAs. ENZYMES: Proline dehydrogenase (EC 1.5.5.2); l-glutamate-γ-semialdehyde dehydrogenase (EC 1.2.1.88). DATABASES: The atomic coordinates and structure factor amplitudes have been deposited in the Protein Data Bank under accession number 5UR2. The SAXS data have been deposited in the SASBDB under the ...DATABASES: The atomic coordinates and structure factor amplitudes have been deposited in the Protein Data Bank under accession number 5UR2. The SAXS data have been deposited in the SASBDB under the following accession codes: SASDCP3 (BbPutA), SASDCQ3 (DvPutA 1.5 mg·mL ), SASDCX3 (DvPutA 3.0 mg·mL ), SASDCY3 (DvPutA 4.5 mg·mL ), SASDCR3 (LpPutA 3.0 mg·mL ), SASDCV3 (LpPutA 5.0 mg·mL ), SASDCW3 (LpPutA 8.0 mg·mL ), SASDCS3 (BjPutA 2.3 mg·mL ), SASDCT3 (BjPutA 4.7 mg·mL ), SASDCU3 (BjPutA 7.0 mg·mL ), SASDCZ3 (R51E 2.3 mg·mL ), SASDC24 (R51E 4.7 mg·mL ), SASDC34 (R51E 7.0 mg·mL ). |
 Contact author Contact author |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
-Data source
| SASBDB page |  SASDCW3 SASDCW3 |
|---|
-Related structure data
- External links
External links
| Related items in Molecule of the Month |
|---|
-Models
| Model #1427 |  Type: dummy / Radius of dummy atoms: 3.30 A / Symmetry: P2 / Chi-square value: 4.222  Search similar-shape structures of this assembly by Omokage search (details) Search similar-shape structures of this assembly by Omokage search (details) |
|---|---|
| Model #1429 |  Type: dummy / Software: (5.0 r6293) / Radius of dummy atoms: 5.00 A / Chi-square value: 4.222  Search similar-shape structures of this assembly by Omokage search (details) Search similar-shape structures of this assembly by Omokage search (details) |
- Sample
Sample
 Sample Sample | Name: Proline utilization A from Legionella pneumophila 8 mg/mL Specimen concentration: 8 mg/ml |
|---|---|
| Buffer | Name: 50 mM Tris-HCl, 50 mM NaCl, 0.5 mM EDTA, and 0.5 mM THP at pH 7.5. pH: 7.5 |
| Entity #680 | Name: LpPutA / Type: protein / Description: Bifunctional protein PutA / Formula weight: 119.178 / Num. of mol.: 2 Source: Legionella pneumophila subsp. pneumophila (strain Philadelphia 1 / ATCC 33152 / DSM 7513) References: UniProt: Q5ZUU6 Sequence: HHHHHHGKPI PNPLLGLDST ENLYFQGMLE KQSIHLPEGL RAAINKAYRM DELSLITELS EQAALDPQQM MAIKTSATKL VQSVRSERKK STGIDSFLTE YALSSDEGIA LMCLAEALLR VPDNATIDNL IKDKLAGGDW GAHRGQSESF FVNATTWALM LTGKVLTPEK ...Sequence: HHHHHHGKPI PNPLLGLDST ENLYFQGMLE KQSIHLPEGL RAAINKAYRM DELSLITELS EQAALDPQQM MAIKTSATKL VQSVRSERKK STGIDSFLTE YALSSDEGIA LMCLAEALLR VPDNATIDNL IKDKLAGGDW GAHRGQSESF FVNATTWALM LTGKVLTPEK AENTLTKALL KLVNRSSEAV VRKAVDKAMR IMSKQFVMGR TINEALARAK KKEDRGYRYS YDMLGEAALT SADAARYFEA YKEAIISIGE KADKHSDVYR RPGISIKLSA LHPRYSEFQY ERVMAELPPK LLALSRLAKD YGIALTIDAE ESERLDLSLD VIEKVFTDES LQGWNGFGLA VQSYQKRAFY VLDWVAALAR SKQRRIMVRL IKGAYWDSEI KKTQMQGFSE YPVFTRKVFT DVSFQACAKK ILTMTDAIYP QFATHNAYSV AMILNLVGGY RDFEFQCLHG MGNELYEQIV PANCYGIPCR IYAPVGSHED LLPYLVRRLL ENGANSSFVN RIVDDKAPIS ELVEDPVAKS RSLLDKINKN IPLPEDIFLP VRKNSKGFDF TNRLERALLQ QELAKIESKE WQASPMIAGR KLSRDLLQTV MSPQQPAYAI GSVQQATLDD VEVALNQAKL AFELWSKKPV EERASCLNRF ADLLQANMAE LMVLTCREAG KTWSDGIAEV REAIDFCRYY AKKAQELMSS PQRFNGYTGE LNELSLHPRG TILCISPWNF PLAIFTGQVV AGLVTGNCVI AKPAEQTPLI AAYAVKLMHQ AGIPEGVIQL IPGAGETIGA ALVADKRIKA VLFTGSTDTA NLINRTLATR GGEIIPLIAE TGGQNAMIVD SSALLEQVVV DAVTSAFGSA GQRCSALRVL YVQEEVYPRT VELLKGAMAE LVVGDPQWLS TDVGPVIDKE ALSILKNHVE NMRKHHEILY QCTVDDEALS GYFMPPTAIA IDSISALEKE VFGPILHVIQ FKRKDLDKVI NQINQTSYGL TLGIHSRINE TVDYIRQRVH AGNCYVNRNM IGAVVGLQPF GGEGLSGTGP KAGGPNYLIR LCHERTYTVD TTAAGGNASL MSIPEEG |
-Experimental information
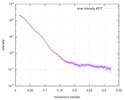
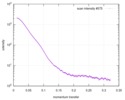
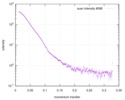
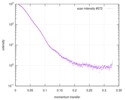
| Beam | Instrument name: Advanced Light Source (ALS) 12.3.1 (SIBYLS) City: Berkeley, CA / 国: USA  / Type of source: X-ray synchrotron / Type of source: X-ray synchrotron | |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Detector | Name: MAR 165 CCD | |||||||||||||||||||||||||||||||||
| Scan | Measurement date: Apr 20, 2010 / Unit: 1/A /
| |||||||||||||||||||||||||||||||||
| Distance distribution function P(R) |
| |||||||||||||||||||||||||||||||||
| Result |
|
 Movie
Movie Controller
Controller