+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-6495 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Electron microscopy of apo human XPC complex | |||||||||
 マップデータ マップデータ | EMAN2 reconstruction of apo human XPC complex | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | transcription / DNA repair / stem cells | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報heteroduplex DNA loop binding / pyrimidine dimer repair by nucleotide-excision repair / nucleotide-excision repair factor 2 complex / XPC complex / nucleotide-excision repair complex / 9+2 motile cilium / photoreceptor connecting cilium / DNA damage sensor activity / regulation of proteasomal ubiquitin-dependent protein catabolic process / transcription export complex 2 ...heteroduplex DNA loop binding / pyrimidine dimer repair by nucleotide-excision repair / nucleotide-excision repair factor 2 complex / XPC complex / nucleotide-excision repair complex / 9+2 motile cilium / photoreceptor connecting cilium / DNA damage sensor activity / regulation of proteasomal ubiquitin-dependent protein catabolic process / transcription export complex 2 / heterotrimeric G-protein binding / response to auditory stimulus / nuclear pore nuclear basket / bubble DNA binding / UV-damage excision repair / cellular response to interleukin-7 / response to UV-B / mitotic intra-S DNA damage checkpoint signaling / regulation of mitotic cell cycle phase transition / proteasome binding / site of DNA damage / centriole replication / polyubiquitin modification-dependent protein binding / mRNA transport / embryonic organ development / mismatch repair / SUMOylation of DNA damage response and repair proteins / Loss of Nlp from mitotic centrosomes / Loss of proteins required for interphase microtubule organization from the centrosome / Recruitment of mitotic centrosome proteins and complexes / Recruitment of NuMA to mitotic centrosomes / Anchoring of the basal body to the plasma membrane / centriole / AURKA Activation by TPX2 / proteasome complex / G-protein beta/gamma-subunit complex binding / ciliary basal body / Josephin domain DUBs / regulation of cytokinesis / ubiquitin binding / N-glycan trimming in the ER and Calnexin/Calreticulin cycle / nucleotide-excision repair / DNA Damage Recognition in GG-NER / Formation of Incision Complex in GG-NER / Regulation of PLK1 Activity at G2/M Transition / protein transport / apical part of cell / single-stranded DNA binding / mitotic cell cycle / spermatogenesis / microtubule binding / proteasome-mediated ubiquitin-dependent protein catabolic process / RNA polymerase II-specific DNA-binding transcription factor binding / damaged DNA binding / transcription coactivator activity / response to xenobiotic stimulus / RNA polymerase II cis-regulatory region sequence-specific DNA binding / cell division / intracellular membrane-bounded organelle / DNA repair / centrosome / calcium ion binding / protein-containing complex binding / chromatin / nucleolus / positive regulation of DNA-templated transcription / mitochondrion / nucleoplasm / nucleus / plasma membrane / cytosol / cytoplasm 類似検索 - 分子機能 | |||||||||
| 生物種 |  Homo sapiens (ヒト) Homo sapiens (ヒト) | |||||||||
| 手法 | 単粒子再構成法 / ネガティブ染色法 / 解像度: 25.0 Å | |||||||||
 データ登録者 データ登録者 | Zhang ET / He Y / Grob P / Fong YW / Nogales E / Tjian R | |||||||||
 引用 引用 |  ジャーナル: Proc Natl Acad Sci U S A / 年: 2015 ジャーナル: Proc Natl Acad Sci U S A / 年: 2015タイトル: Architecture of the human XPC DNA repair and stem cell coactivator complex. 著者: Elisa T Zhang / Yuan He / Patricia Grob / Yick W Fong / Eva Nogales / Robert Tjian /  要旨: The Xeroderma pigmentosum complementation group C (XPC) complex is a versatile factor involved in both nucleotide excision repair and transcriptional coactivation as a critical component of the ...The Xeroderma pigmentosum complementation group C (XPC) complex is a versatile factor involved in both nucleotide excision repair and transcriptional coactivation as a critical component of the NANOG, OCT4, and SOX2 pluripotency gene regulatory network. Here we present the structure of the human holo-XPC complex determined by single-particle electron microscopy to reveal a flexible, ear-shaped structure that undergoes localized loss of order upon DNA binding. We also determined the structure of the complete yeast homolog Rad4 holo-complex to find a similar overall architecture to the human complex, consistent with their shared DNA repair functions. Localized differences between these structures reflect an intriguing phylogenetic divergence in transcriptional capabilities that we present here. Having positioned the constituent subunits by tagging and deletion, we propose a model of key interaction interfaces that reveals the structural basis for this difference in functional conservation. Together, our findings establish a framework for understanding the structure-function relationships of the XPC complex in the interplay between transcription and DNA repair. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_6495.map.gz emd_6495.map.gz | 6.4 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-6495-v30.xml emd-6495-v30.xml emd-6495.xml emd-6495.xml | 13.6 KB 13.6 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_6495.tif emd_6495.tif | 755.8 KB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6495 http://ftp.pdbj.org/pub/emdb/structures/EMD-6495 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6495 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6495 | HTTPS FTP |
-検証レポート
| 文書・要旨 |  emd_6495_validation.pdf.gz emd_6495_validation.pdf.gz | 79 KB | 表示 |  EMDB検証レポート EMDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  emd_6495_full_validation.pdf.gz emd_6495_full_validation.pdf.gz | 78 KB | 表示 | |
| XML形式データ |  emd_6495_validation.xml.gz emd_6495_validation.xml.gz | 493 B | 表示 | |
| アーカイブディレクトリ |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6495 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6495 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6495 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6495 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_6495.map.gz / 形式: CCP4 / 大きさ: 7.8 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_6495.map.gz / 形式: CCP4 / 大きさ: 7.8 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
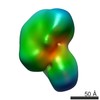
| 注釈 | EMAN2 reconstruction of apo human XPC complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 3.01 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
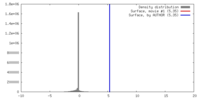
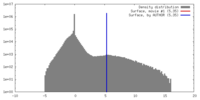
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
- 試料の構成要素
試料の構成要素
-全体 : Apo human XPC complex
| 全体 | 名称: Apo human XPC complex |
|---|---|
| 要素 |
|
-超分子 #1000: Apo human XPC complex
| 超分子 | 名称: Apo human XPC complex / タイプ: sample / ID: 1000 詳細: Monodisperse. Thawed from -80 degrees Celsius and placed on ice immediately prior to grid preparation. 集合状態: heterotrimer / Number unique components: 3 |
|---|---|
| 分子量 | 実験値: 200 KDa / 理論値: 200 KDa / 手法: SDS-PAGE, size exclusion chromatography |
-分子 #1: Xeroderma pigmentosum group C-complementing protein
| 分子 | 名称: Xeroderma pigmentosum group C-complementing protein / タイプ: protein_or_peptide / ID: 1 Name.synonym: XPC, p125, DNA repair protein complementing XP-C cells 詳細: contains N-terminal 6xHis and TEV cleavage site / コピー数: 1 / 集合状態: monomer / 組換発現: Yes |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) / 別称: human / 細胞中の位置: nucleus, cytoplasm Homo sapiens (ヒト) / 別称: human / 細胞中の位置: nucleus, cytoplasm |
| 分子量 | 実験値: 125 KDa / 理論値: 125 KDa |
| 組換発現 | 生物種:  組換細胞: Sf9 |
| 配列 | UniProtKB: DNA repair protein complementing XP-C cells / GO: XPC complex |
-分子 #2: RAD23 homolog B
| 分子 | 名称: RAD23 homolog B / タイプ: protein_or_peptide / ID: 2 Name.synonym: RAD23B, HR23B, HHR23B, p58, XP-C repair-complementing complex 58 kDa protein 詳細: contains N-terminal 1xFLAG tag / コピー数: 1 / 集合状態: monomer / 組換発現: Yes |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) / 別称: human / 細胞中の位置: nucleus, cytoplasm Homo sapiens (ヒト) / 別称: human / 細胞中の位置: nucleus, cytoplasm |
| 分子量 | 実験値: 58 KDa / 理論値: 58 KDa |
| 組換発現 | 生物種:  組換細胞: Sf9 |
| 配列 | UniProtKB: UV excision repair protein RAD23 homolog B / GO: XPC complex / InterPro: UV excision repair protein Rad23 |
-分子 #3: Centrin-2
| 分子 | 名称: Centrin-2 / タイプ: protein_or_peptide / ID: 3 / Name.synonym: CETN2 / コピー数: 1 / 集合状態: monomer / 組換発現: Yes |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) / 別称: human / 細胞中の位置: nucleus, cytoplasm, centrosomes Homo sapiens (ヒト) / 別称: human / 細胞中の位置: nucleus, cytoplasm, centrosomes |
| 分子量 | 実験値: 18 KDa / 理論値: 18 KDa |
| 組換発現 | 生物種:  組換細胞: Sf9 |
| 配列 | UniProtKB: Centrin-2 / GO: XPC complex / InterPro: INTERPRO: IPR029528 |
-実験情報
-構造解析
| 手法 | ネガティブ染色法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 0.01 mg/mL |
|---|---|
| 緩衝液 | pH: 7.6 詳細: 300 mM KCl, 50 mM HEPES, 0.1% NP-40 alternative, 10% glycerol, 0.1 mM EDTA, 1 mM MgCl2, 1 mM TCEP, 1 mM DTT |
| 染色 | タイプ: NEGATIVE 詳細: Grids with adsorbed protein were floated on 2 successive droplets of buffer G (300 mM KCl, 25 mM HEPES ph 7.6, 3% w/v trehalose, 0.01% NP-40 alternative, 1 mM TCEP, 1 mM DTT, 0.1 mM EDTA, 1 ...詳細: Grids with adsorbed protein were floated on 2 successive droplets of buffer G (300 mM KCl, 25 mM HEPES ph 7.6, 3% w/v trehalose, 0.01% NP-40 alternative, 1 mM TCEP, 1 mM DTT, 0.1 mM EDTA, 1 mM MgCl2) and subsequently floated on 4 successive 1% w/v uranyl droplets for 10 seconds each. |
| グリッド | 詳細: 400 mesh copper grid with thin carbon support |
| 凍結 | 凍結剤: NONE / 装置: OTHER |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TECNAI F20 |
|---|---|
| アライメント法 | Legacy - 非点収差: Astigmatism was corrected at 80,000 times and 280,000 times magnification. |
| 日付 | 2013年8月13日 |
| 撮影 | カテゴリ: CCD フィルム・検出器のモデル: GATAN ULTRASCAN 4000 (4k x 4k) 実像数: 426 / 平均電子線量: 20 e/Å2 / ビット/ピクセル: 32 |
| Tilt angle min | 0 |
| Tilt angle max | 0 |
| 電子線 | 加速電圧: 120 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / Cs: 2.2 mm / 最大 デフォーカス(公称値): 2.25 µm / 最小 デフォーカス(公称値): -0.07 µm / 倍率(公称値): 80000 |
| 試料ステージ | 試料ホルダーモデル: OTHER |
| 実験機器 |  モデル: Tecnai F20 / 画像提供: FEI Company |
- 画像解析
画像解析
| 詳細 | Particles were selected by DoGPicker in the Appion pipeline. |
|---|---|
| CTF補正 | 詳細: whole micrograph |
| 最終 再構成 | アルゴリズム: OTHER / 解像度のタイプ: BY AUTHOR / 解像度: 25.0 Å / 解像度の算出法: OTHER / ソフトウェア - 名称: EMAN2 / 使用した粒子像数: 22777 |
| 最終 2次元分類 | クラス数: 141 |
 ムービー
ムービー コントローラー
コントローラー





















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)