+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Cryo-EM structure of the human 80S ribosome with 4 um Tigecycline | |||||||||
 マップデータ マップデータ | consensus refined map without post processing | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | ribosome / Tigecycline / antibiotic | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報eukaryotic 80S initiation complex / negative regulation of protein neddylation / translation at presynapse / positive regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis / negative regulation of endoplasmic reticulum unfolded protein response / protein tyrosine kinase inhibitor activity / axial mesoderm development / oxidized pyrimidine DNA binding / response to TNF agonist / positive regulation of base-excision repair ...eukaryotic 80S initiation complex / negative regulation of protein neddylation / translation at presynapse / positive regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis / negative regulation of endoplasmic reticulum unfolded protein response / protein tyrosine kinase inhibitor activity / axial mesoderm development / oxidized pyrimidine DNA binding / response to TNF agonist / positive regulation of base-excision repair / ribosomal protein import into nucleus / positive regulation of respiratory burst involved in inflammatory response / embryonic brain development / negative regulation of formation of translation preinitiation complex / nucleolus organization / regulation of adenylate cyclase-activating G protein-coupled receptor signaling pathway / positive regulation of intrinsic apoptotic signaling pathway in response to DNA damage / positive regulation of gastrulation / 90S preribosome assembly / IRE1-RACK1-PP2A complex / positive regulation of Golgi to plasma membrane protein transport / positive regulation of endodeoxyribonuclease activity / TNFR1-mediated ceramide production / negative regulation of DNA repair / negative regulation of RNA splicing / GAIT complex / negative regulation of intrinsic apoptotic signaling pathway in response to hydrogen peroxide / supercoiled DNA binding / neural crest cell differentiation / oxidized purine DNA binding / NF-kappaB complex / middle ear morphogenesis / ubiquitin-like protein conjugating enzyme binding / negative regulation of phagocytosis / regulation of establishment of cell polarity / positive regulation of ubiquitin-protein transferase activity / A band / rRNA modification in the nucleus and cytosol / erythrocyte homeostasis / Formation of the ternary complex, and subsequently, the 43S complex / alpha-beta T cell differentiation / cytoplasmic side of rough endoplasmic reticulum membrane / regulation of G1 to G0 transition / exit from mitosis / laminin receptor activity / pigmentation / positive regulation of intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator / regulation of translation involved in cellular response to UV / protein-DNA complex disassembly / positive regulation of DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator / protein kinase A binding / optic nerve development / negative regulation of ubiquitin protein ligase activity / Ribosomal scanning and start codon recognition / ion channel inhibitor activity / response to aldosterone / Translation initiation complex formation / retinal ganglion cell axon guidance / positive regulation of mitochondrial depolarization / mammalian oogenesis stage / homeostatic process / G1 to G0 transition / activation-induced cell death of T cells / lung morphogenesis / positive regulation of T cell receptor signaling pathway / macrophage chemotaxis / negative regulation of Wnt signaling pathway / fibroblast growth factor binding / iron-sulfur cluster binding / monocyte chemotaxis / positive regulation of activated T cell proliferation / Protein hydroxylation / regulation of cell division / negative regulation of peptidyl-serine phosphorylation / BH3 domain binding / mTORC1-mediated signalling / SARS-CoV-1 modulates host translation machinery / Peptide chain elongation / positive regulation of intrinsic apoptotic signaling pathway by p53 class mediator / cysteine-type endopeptidase activator activity involved in apoptotic process / Selenocysteine synthesis / positive regulation of signal transduction by p53 class mediator / Formation of a pool of free 40S subunits / blastocyst development / phagocytic cup / Eukaryotic Translation Termination / ubiquitin ligase inhibitor activity / negative regulation of respiratory burst involved in inflammatory response / Response of EIF2AK4 (GCN2) to amino acid deficiency / SRP-dependent cotranslational protein targeting to membrane / negative regulation of phosphatidylinositol 3-kinase/protein kinase B signal transduction / protein localization to nucleus / Viral mRNA Translation / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / GTP hydrolysis and joining of the 60S ribosomal subunit / L13a-mediated translational silencing of Ceruloplasmin expression / TOR signaling / endonucleolytic cleavage to generate mature 3'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / T cell proliferation involved in immune response / regulation of translational fidelity 類似検索 - 分子機能 | |||||||||
| 生物種 |  Homo sapiens (ヒト) Homo sapiens (ヒト) | |||||||||
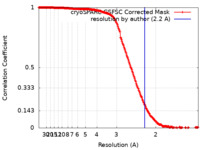
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 2.2 Å | |||||||||
 データ登録者 データ登録者 | Li X / Wang M / Denk T / Cheng J | |||||||||
| 資金援助 | 1件
| |||||||||
 引用 引用 |  ジャーナル: Nat Commun / 年: 2024 ジャーナル: Nat Commun / 年: 2024タイトル: Structural basis for differential inhibition of eukaryotic ribosomes by tigecycline. 著者: Xiang Li / Mengjiao Wang / Timo Denk / Robert Buschauer / Yi Li / Roland Beckmann / Jingdong Cheng /   要旨: Tigecycline is widely used for treating complicated bacterial infections for which there are no effective drugs. It inhibits bacterial protein translation by blocking the ribosomal A-site. However, ...Tigecycline is widely used for treating complicated bacterial infections for which there are no effective drugs. It inhibits bacterial protein translation by blocking the ribosomal A-site. However, even though it is also cytotoxic for human cells, the molecular mechanism of its inhibition remains unclear. Here, we present cryo-EM structures of tigecycline-bound human mitochondrial 55S, 39S, cytoplasmic 80S and yeast cytoplasmic 80S ribosomes. We find that at clinically relevant concentrations, tigecycline effectively targets human 55S mitoribosomes, potentially, by hindering A-site tRNA accommodation and by blocking the peptidyl transfer center. In contrast, tigecycline does not bind to human 80S ribosomes under physiological concentrations. However, at high tigecycline concentrations, in addition to blocking the A-site, both human and yeast 80S ribosomes bind tigecycline at another conserved binding site restricting the movement of the L1 stalk. In conclusion, the observed distinct binding properties of tigecycline may guide new pathways for drug design and therapy. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_39456.map.gz emd_39456.map.gz | 480 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-39456-v30.xml emd-39456-v30.xml emd-39456.xml emd-39456.xml | 104.9 KB 104.9 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_39456_fsc.xml emd_39456_fsc.xml | 20.7 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_39456.png emd_39456.png | 141.2 KB | ||
| Filedesc metadata |  emd-39456.cif.gz emd-39456.cif.gz | 19.9 KB | ||
| その他 |  emd_39456_additional_1.map.gz emd_39456_additional_1.map.gz emd_39456_half_map_1.map.gz emd_39456_half_map_1.map.gz emd_39456_half_map_2.map.gz emd_39456_half_map_2.map.gz | 84.2 MB 885.8 MB 885.8 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-39456 http://ftp.pdbj.org/pub/emdb/structures/EMD-39456 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-39456 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-39456 | HTTPS FTP |
-検証レポート
| 文書・要旨 |  emd_39456_validation.pdf.gz emd_39456_validation.pdf.gz | 1.1 MB | 表示 |  EMDB検証レポート EMDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  emd_39456_full_validation.pdf.gz emd_39456_full_validation.pdf.gz | 1.1 MB | 表示 | |
| XML形式データ |  emd_39456_validation.xml.gz emd_39456_validation.xml.gz | 31.3 KB | 表示 | |
| CIF形式データ |  emd_39456_validation.cif.gz emd_39456_validation.cif.gz | 41.3 KB | 表示 | |
| アーカイブディレクトリ |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-39456 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-39456 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-39456 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-39456 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  8yopMC  8k2aC  8k2bC  8k2cC  8k2dC  8k82C  8xsxC  8xsyC  8xszC  8xt0C  8xt1C  8xt2C  8xt3C  8yooC M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_39456.map.gz / 形式: CCP4 / 大きさ: 953.9 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_39456.map.gz / 形式: CCP4 / 大きさ: 953.9 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | consensus refined map without post processing | ||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 0.727 Å | ||||||||||||||||||||||||||||||||||||
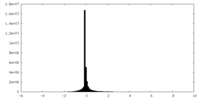
| 密度 |
| ||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
-追加マップ: local resolution filtered map
| ファイル | emd_39456_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | local resolution filtered map | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
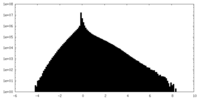
| 密度ヒストグラム |
-ハーフマップ: #1
| ファイル | emd_39456_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
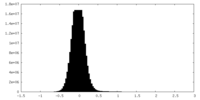
| 密度ヒストグラム |
-ハーフマップ: #2
| ファイル | emd_39456_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
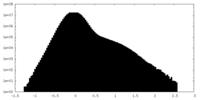
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
+全体 : 80S ribosome with 5um tigecycline
+超分子 #1: 80S ribosome with 5um tigecycline
+分子 #1: 28S rRNA
+分子 #2: 5S rRNA
+分子 #3: 5.8S rRNA
+分子 #49: 18S rRNA
+分子 #4: 60S ribosomal protein L8
+分子 #5: 60S ribosomal protein L3
+分子 #6: 60S ribosomal protein L4
+分子 #7: 60S ribosomal protein L5
+分子 #8: 60S ribosomal protein L6
+分子 #9: 60S ribosomal protein L7
+分子 #10: 60S ribosomal protein L7a
+分子 #11: 60S ribosomal protein L9
+分子 #12: Large ribosomal subunit protein uL16
+分子 #13: 60S ribosomal protein L11
+分子 #14: 60S ribosomal protein L13
+分子 #15: 60S ribosomal protein L14
+分子 #16: 60S ribosomal protein L15
+分子 #17: 60S ribosomal protein L13a
+分子 #18: 60S ribosomal protein L17
+分子 #19: 60S ribosomal protein L18
+分子 #20: 60S ribosomal protein L19
+分子 #21: 60S ribosomal protein L18a
+分子 #22: 60S ribosomal protein L21
+分子 #23: 60S ribosomal protein L22
+分子 #24: 60S ribosomal protein L23
+分子 #25: 60S ribosomal protein L24
+分子 #26: 60S ribosomal protein L23a
+分子 #27: 60S ribosomal protein L26
+分子 #28: 60S ribosomal protein L27
+分子 #29: 60S ribosomal protein L27a
+分子 #30: 60S ribosomal protein L29
+分子 #31: 60S ribosomal protein L30
+分子 #32: 60S ribosomal protein L31
+分子 #33: 60S ribosomal protein L32
+分子 #34: 60S ribosomal protein L35a
+分子 #35: 60S ribosomal protein L34
+分子 #36: 60S ribosomal protein L35
+分子 #37: 60S ribosomal protein L36
+分子 #38: 60S ribosomal protein L37
+分子 #39: 60S ribosomal protein L38
+分子 #40: 60S ribosomal protein L39
+分子 #41: Ubiquitin-60S ribosomal protein L40
+分子 #42: 60S ribosomal protein L41
+分子 #43: 60S ribosomal protein L36a
+分子 #44: 60S ribosomal protein L37a
+分子 #45: 60S ribosomal protein L28
+分子 #46: Large ribosomal subunit protein uL10
+分子 #47: 60S ribosomal protein L12
+分子 #48: 60S ribosomal protein L10a
+分子 #50: 40S ribosomal protein SA
+分子 #51: 40S ribosomal protein S3a
+分子 #52: 40S ribosomal protein S3
+分子 #53: 40S ribosomal protein S4, X isoform
+分子 #54: 40S ribosomal protein S5
+分子 #55: 40S ribosomal protein S7
+分子 #56: 40S ribosomal protein S8
+分子 #57: 40S ribosomal protein S10
+分子 #58: 40S ribosomal protein S11
+分子 #59: 40S ribosomal protein S15
+分子 #60: 40S ribosomal protein S16
+分子 #61: 40S ribosomal protein S17
+分子 #62: 40S ribosomal protein S18
+分子 #63: 40S ribosomal protein S19
+分子 #64: 40S ribosomal protein S20
+分子 #65: 40S ribosomal protein S21
+分子 #66: 40S ribosomal protein S23
+分子 #67: 40S ribosomal protein S26
+分子 #68: 40S ribosomal protein S28
+分子 #69: 40S ribosomal protein S29
+分子 #70: Receptor of activated protein C kinase 1
+分子 #71: 40S ribosomal protein S2
+分子 #72: 40S ribosomal protein S6
+分子 #73: 40S ribosomal protein S9
+分子 #74: 40S ribosomal protein S12
+分子 #75: 40S ribosomal protein S13
+分子 #76: 40S ribosomal protein S14
+分子 #77: 40S ribosomal protein S15a
+分子 #78: 40S ribosomal protein S24
+分子 #79: 40S ribosomal protein S25
+分子 #80: 40S ribosomal protein S27
+分子 #81: 40S ribosomal protein S30
+分子 #82: Ubiquitin-40S ribosomal protein S27a
+分子 #83: MAGNESIUM ION
+分子 #84: ZINC ION
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.4 |
|---|---|
| グリッド | モデル: Quantifoil R3/3 / 材質: COPPER / 支持フィルム - 材質: CARBON / 支持フィルム - トポロジー: CONTINUOUS / 支持フィルム - Film thickness: 2 |
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) 平均電子線量: 44.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / 最大 デフォーカス(公称値): 2.5 µm / 最小 デフォーカス(公称値): 1.0 µm |
| 試料ステージ | ホルダー冷却材: NITROGEN |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー


























































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)