-Search query
-Search result
Showing 1 - 50 of 96 items for (author: ueno & l)

EMDB-19067: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factors Balon and RaiA (structure 1).

EMDB-19076: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factor Balon, mRNA and P-site tRNA (structure 2).

EMDB-19077: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factor Balon and EF-Tu(GDP) (structure 3).

PDB-8rd8: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factors Balon and RaiA (structure 1).

PDB-8rdv: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factor Balon, mRNA and P-site tRNA (structure 2).

PDB-8rdw: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factor Balon and EF-Tu(GDP) (structure 3).
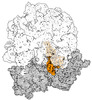
EMDB-43074: 
Cryo-EM structure of the Mycobacterium smegmatis 70S ribosome in complex with hibernation factor Msmeg1130 (Balon) (Structure 4)
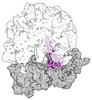
EMDB-43075: 
Cryo-EM structure of the Mycobacterium smegmatis 70S ribosome in complex with hibernation factor Rv2629 (Balon) (Structure 5)
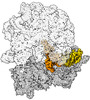
EMDB-43076: 
Cryo-EM structure of the Mycobacterium smegmatis 70S ribosome in complex with hibernation factor Msmeg1130 (Balon) and MsmegEF-Tu(GDP) (Composite structure 6)

EMDB-43077: 
Cryo-EM structure of the Mycobacterium smegmatis 70S ribosome in complex with hibernation factor Msmeg1130 (Balon) and MsmegEF-Tu(GDP) (Structure 6)

EMDB-43078: 
Hibernation factor Msmeg1130 (Balon) and MsmegEF-Tu(GDP) bound to Mycobacterium smegmatis 70S ribosome, from focused 3D classification and refinement (Structure 6)

PDB-8v9j: 
Cryo-EM structure of the Mycobacterium smegmatis 70S ribosome in complex with hibernation factor Msmeg1130 (Balon) (Structure 4)

PDB-8v9k: 
Cryo-EM structure of the Mycobacterium smegmatis 70S ribosome in complex with hibernation factor Rv2629 (Balon) (Structure 5)

PDB-8v9l: 
Cryo-EM structure of the Mycobacterium smegmatis 70S ribosome in complex with hibernation factor Msmeg1130 (Balon) and MsmegEF-Tu(GDP) (Composite structure 6)

EMDB-40687: 
PS3 F1 Rotorless, no ATP

EMDB-40688: 
PS3 F1 Rotorless, low ATP

EMDB-40689: 
PS3 F1 Rotorless, high ATP

PDB-8spv: 
PS3 F1 Rotorless, no ATP

PDB-8spw: 
PS3 F1 Rotorless, low ATP

PDB-8spx: 
PS3 F1 Rotorless, high ATP
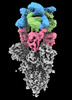
EMDB-16453: 
SARS-CoV-2 Omicron Variant Spike Trimer in complex with three 17T2 Fabs

EMDB-16473: 
SARS-CoV-2 spike in complex with the 17T2 neutralizing antibody Fab fragment (local refinement of RBD and Fab)

PDB-8c89: 
SARS-CoV-2 spike in complex with the 17T2 neutralizing antibody Fab fragment (local refinement of RBD and Fab)

EMDB-15243: 
Mycobacterium tuberculosis ClpC1 hexamer structure bound to the natural product antibiotic ecumicin (class 2)

EMDB-33614: 
Cryo-EM structure of the Mycolicibacterium smegmatis F1-ATPase

EMDB-33615: 
Cryo-EM structure of F-ATP synthase from Mycolicibacterium smegmatis (rotational state 1)

EMDB-33616: 
Cryo-EM structure of F-ATP synthase from Mycolicibacterium smegmatis (rotational state 2)

EMDB-33617: 
Cryo-EM structure of F-ATP synthase from Mycolicibacterium smegmatis (rotational state 3) (backbone)

PDB-7y5a: 
Cryo-EM structure of the Mycolicibacterium smegmatis F1-ATPase

PDB-7y5b: 
Cryo-EM structure of F-ATP synthase from Mycolicibacterium smegmatis (rotational state 1)

PDB-7y5c: 
Cryo-EM structure of F-ATP synthase from Mycolicibacterium smegmatis (rotational state 2)

PDB-7y5d: 
Cryo-EM structure of F-ATP synthase from Mycolicibacterium smegmatis (rotational state 3) (backbone)

EMDB-34221: 
SARS-CoV-2 BA.2.75 spike glycoprotein (closed state 1)

EMDB-34222: 
SARS-CoV-2 BA.2.75 spike glycoprotein (closed state 2)

EMDB-34223: 
SARS-CoV-2 BA.2.75 spike glycoprotein (1-up state)

EMDB-34224: 
SARS-CoV-2 BA.2.75 spike glycoprotein in complex with ACE2

PDB-8gs6: 
Structure of the SARS-CoV-2 BA.2.75 spike glycoprotein (closed state 1)

EMDB-15240: 
Mycobacterium tuberculosis ClpC1 hexamer structure

EMDB-15241: 
Mycobacterium tuberculosis ClpC1 hexamer structure bound to the natural product antibiotic Cyclomarin

EMDB-15242: 
Mycobacterium tuberculosis ClpC1 hexamer structure bound to the natural product antibiotic Ecumycin (class 1)

PDB-8a8u: 
Mycobacterium tuberculosis ClpC1 hexamer structure

PDB-8a8v: 
Mycobacterium tuberculosis ClpC1 hexamer structure bound to the natural product antibiotic Cyclomarin

PDB-8a8w: 
Mycobacterium tuberculosis ClpC1 hexamer structure bound to the natural product antibiotic Ecumycin (class 1)

EMDB-27327: 
PI 3-kinase alpha with nanobody 3-126

EMDB-27330: 
PI 3-kinase alpha with nanobody 3-159

EMDB-27334: 
PI 3-kinase alpha with nanobody 3-142

EMDB-27336: 
PI 3-kinase alpha with nanobody 3-142, crosslinked with DSG

PDB-8dcp: 
PI 3-kinase alpha with nanobody 3-126

PDB-8dcx: 
PI 3-kinase alpha with nanobody 3-159

PDB-8dd4: 
PI 3-kinase alpha with nanobody 3-142
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
