-Search query
-Search result
Showing 1 - 50 of 1,308 items for (author: tong & j)

EMDB-38099: 
Cryo-EM structures of RNF168/UbcH5c-Ub in complex with H2AK13Ub nucleosomes determined by intein-based E2-Ub-NCP conjugation strategy

EMDB-38100: 
Cryo-EM structures of RNF168/UbcH5c-Ub/nucleosomes complex determined by activity-based chemical trapping strategy

EMDB-38101: 
Cryo-EM structures of RNF168/UbcH5c-Ub in complex with H2AK13Ub nucleosomes determined by activity-based chemical trapping strategy (adjacent H2AK13/15 dual-monoubiquitination)

EMDB-38102: 
Cryo-EM map of RNF168/UbcH5c-Ub/nucleosome determined by E2-Ub-NCP conjugation strategy

EMDB-44482: 
Cryo-EM structure of the HIV-1 JR-FL IDL Env trimer in complex with PGT122 Fab
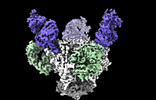
EMDB-44484: 
Cryo-EM structure of the HIV-1 BG505 IDL Env trimer in complex with 3BNC117 and 10-1074 Fabs
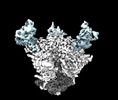
EMDB-44491: 
Cryo-EM structure of the HIV-1 WITO IDL Env trimer in complex with PGT122 Fab

PDB-9ber: 
Cryo-EM structure of the HIV-1 JR-FL IDL Env trimer in complex with PGT122 Fab

PDB-9bew: 
Cryo-EM structure of the HIV-1 BG505 IDL Env trimer in complex with 3BNC117 and 10-1074 Fabs

PDB-9bf6: 
Cryo-EM structure of the HIV-1 WITO IDL Env trimer in complex with PGT122 Fab

EMDB-38873: 
cryo-EM structure of Staphylococcus aureus(ATCC 29213) 50S ribosome in complex with MCX-190.

EMDB-38874: 
Cryo-EM structure of Staphylococcus aureus (15B196) 50S ribosome in complex with MCX-190.

EMDB-38875: 
Cryo-EM structure of Staphylococcus aureus 70S ribosome (strain 15B196) in complex with MCX-190.

EMDB-38876: 
cryo-EM structure of Staphylococcus aureus(ATCC 29213) 70S ribosome in complex with MCX-190.

PDB-8y36: 
cryo-EM structure of Staphylococcus aureus(ATCC 29213) 50S ribosome in complex with MCX-190.

PDB-8y37: 
Cryo-EM structure of Staphylococcus aureus (15B196) 50S ribosome in complex with MCX-190.

PDB-8y38: 
Cryo-EM structure of Staphylococcus aureus 70S ribosome (strain 15B196) in complex with MCX-190.

PDB-8y39: 
cryo-EM structure of Staphylococcus aureus(ATCC 29213) 70S ribosome in complex with MCX-190.

EMDB-43046: 
Full length integrin AlphaIIbBeta3 in pre-active state
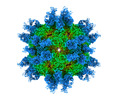

EMDB-32979: 
Cryo-EM structure of Coxsackievirus B1 A-particle in complex with nAb 8A10 (CVB1-A:8A10)

PDB-7x35: 
Cryo-EM structure of Coxsackievirus B1 A-particle in complex with nAb 8A10 (CVB1-A:8A10)

EMDB-38617: 
SARS-CoV-2 RBD + IMCAS-123 + IMCAS-72 Fab

EMDB-38618: 
SARS-CoV-2 RBD + IMCAS-364 + hACE2

EMDB-38619: 
SARS-CoV-2 RBD + IMCAS-364 (Local Refinement)

EMDB-38620: 
SARS-CoV-2 Omicron BA.4 RBD + IMCAS-316 + ACE2

EMDB-38621: 
SARS-CoV-2 spike + IMCAS-123

EMDB-38823: 
Cryo-EM structure of the 123-316 scDb/PT-RBD complex

PDB-8xse: 
SARS-CoV-2 RBD + IMCAS-123 + IMCAS-72 Fab

PDB-8xsf: 
SARS-CoV-2 RBD + IMCAS-364 + hACE2

PDB-8xsi: 
SARS-CoV-2 RBD + IMCAS-364 (Local Refinement)

PDB-8xsj: 
SARS-CoV-2 Omicron BA.4 RBD + IMCAS-316 + ACE2

PDB-8xsl: 
SARS-CoV-2 spike + IMCAS-123

PDB-8y0y: 
Cryo-EM structure of the 123-316 scDb/PT-RBD complex

EMDB-38860: 
structure of RSF-147bp NCP complex Class 0
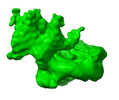

EMDB-38861: 
Structure of RSF-147bpNCP complex class 2

EMDB-38865: 
RSF-38N38NCP complex Class 2

EMDB-37895: 
PNPase mutant of Mycobacterium tuberculosis

EMDB-37896: 
PNPase of M.tuberculosis with its RNA substrate

EMDB-37905: 
PNPase of Mycobacterium tuberculosis

PDB-8wwp: 
PNPase mutant of Mycobacterium tuberculosis

PDB-8wx0: 
PNPase of M.tuberculosis with its RNA substrate

PDB-8wxf: 
PNPase of Mycobacterium tuberculosis

EMDB-35116: 
Cryo-EM structure of 6-subunit Smc5/6

EMDB-35128: 
Cryo-EM structure of 6-subunit Smc5/6 arm region

EMDB-35184: 
Cryo-EM structure of 5-subunit Smc5/6 hinge region

EMDB-35185: 
Cryo-EM structure of 5-subunit Smc5/6 arm region

EMDB-35186: 
Cryo-EM structure of 5-subunit Smc5/6 head region

EMDB-35187: 
Cryo-EM structure of 5-subunit Smc5/6
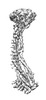
EMDB-37584: 
Cryo-EM structure of 6-subunit Smc5/6 hinge region

EMDB-37586: 
Cryo-EM structure of 6-subunit Smc5/6 head region
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
