-Search query
-Search result
Showing 1 - 50 of 118 items for (author: sutherland & h)

EMDB-70721: 
TMPRSS2 (S441A) bound to the HCoV-NL63 S2'region genetically fused to the HCoV-HKU1 RBD
Method: single particle / : McCallum M, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-70722: 
TMPRSS2 S441A in complex with the H1H7 Fab and anti-kappa light chain nanobody
Method: single particle / : McCallum M, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-73656: 
SARS-CoV-2 spike trimer in the early fusion intermediate conformation bound to the VN01H1 Fab (Fab local refinement)
Method: single particle / : Park YJ, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-73657: 
SARS-CoV-2 spike trimer in the early fusion intermediate conformation bound to the VN01H1 Fab (S2 local refinement)
Method: single particle / : Park YJ, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-73786: 
HCoV-NL63 S2' peptide bound to TMPRSS2 S441A (complexed with the H1H7 Fab and an anti-kappa-nanobody)
Method: single particle / : McCallum M, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-73787: 
SARS-CoV-2 S2 trimer stabilized in the early fusion intermediate conformation by circular permutation and clamping by gp41 (E-FICs-v1)
Method: single particle / : McCallum M, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-75233: 
SARS-CoV-2 spike trimer in the early fusion intermediate conformation bound to the VN01H1 Fab (global refinement)
Method: single particle / : Park YJ, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-75694: 
SARS-CoV-2 spike S2 trimer stabilized in the early fusion intermediate conformation (E-FICs-v3) bound to the VN01H1 Fab (Fab local refinement)
Method: single particle / : McCallum M, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-75695: 
SARS-CoV-2 spike S2 trimer stabilized in the early fusion intermediate conformation (E-FICs-v3) bound to the VN01H1 Fab (S2 local refinement)
Method: single particle / : McCallum M, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-75705: 
SARS-CoV-2 spike S2 trimer stabilized in the early fusion intermediate conformation (E-FICs-v3) bound to C77G12 (Fab local refinement)
Method: single particle / : McCallum M, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-75721: 
SARS-CoV-2 spike S2 trimer stabilized in the early fusion intermediate conformation (E-FICs-v3) bound to the VN01H1 Fab
Method: single particle / : McCallum M, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-75722: 
SARS-CoV-2 spike S2 trimer stabilized in the early fusion intermediate conformation (E-FICs-v3) bound to C77G12 (global refinement)
Method: single particle / : McCallum M, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

PDB-11hk: 
SARS-CoV-2 spike S2 trimer stabilized in the early fusion intermediate conformation (E-FICs-v3) bound to the VN01H1 Fab (Fab local refinement)
Method: single particle / : McCallum M, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

PDB-11hl: 
SARS-CoV-2 spike S2 trimer stabilized in the early fusion intermediate conformation (E-FICs-v3) bound to the VN01H1 Fab (S2 local refinement)
Method: single particle / : McCallum M, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

PDB-11hw: 
SARS-CoV-2 spike S2 trimer stabilized in the early fusion intermediate conformation (E-FICs-v3) bound to C77G12 (Fab local refinement)
Method: single particle / : McCallum M, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

PDB-9opq: 
TMPRSS2 (S441A) bound to the HCoV-NL63 S2'region genetically fused to the HCoV-HKU1 RBD
Method: single particle / : McCallum M, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

PDB-9opr: 
TMPRSS2 S441A in complex with the H1H7 Fab and anti-kappa light chain nanobody
Method: single particle / : McCallum M, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

PDB-9yyu: 
SARS-CoV-2 spike trimer in the early fusion intermediate conformation bound to the VN01H1 Fab (Fab local refinement)
Method: single particle / : Park YJ, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

PDB-9yyv: 
SARS-CoV-2 spike trimer in the early fusion intermediate conformation bound to the VN01H1 Fab (S2 local refinement)
Method: single particle / : Park YJ, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

PDB-9z3j: 
HCoV-NL63 S2' peptide bound to TMPRSS2 S441A (complexed with the H1H7 Fab and an anti-kappa-nanobody)
Method: single particle / : McCallum M, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

PDB-9z3k: 
SARS-CoV-2 S2 trimer stabilized in the early fusion intermediate conformation by circular permutation and clamping by gp41 (E-FICs-v1)
Method: single particle / : McCallum M, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-75697: 
SARS-CoV-2 spike S2 trimer stabilized in the early fusion intermediate conformation (E-FICs-v3) bound to C77G12 (S2 local refinement)
Method: single particle / : McCallum M, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

PDB-11hn: 
SARS-CoV-2 spike S2 trimer stabilized in the early fusion intermediate conformation (E-FICs-v3) bound to C77G12 (S2 local refinement)
Method: single particle / : McCallum M, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-49474: 
Honeybee silk hetereotetramer coiled coil
Method: single particle / : Johnston CL, Jobichen C

EMDB-47570: 
DH726-1 Fab bound to hemagglutinin from influenza A/Solomon Islands/3/2006
Method: single particle / : Finney J, Harrison SC, Walsh Jr RM, Kelsoe G

EMDB-50049: 
Broad substrate scope C-C oxidation in cyclodipeptides catalysed by a flavin-dependent filament
Method: helical / : Sutherland E, Sundaramoorthy R, Czekster CM

PDB-9exv: 
Broad substrate scope C-C oxidation in cyclodipeptides catalysed by a flavin-dependent filament
Method: helical / : Sutherland E, Sundaramoorthy R, Czekster CM

EMDB-40208: 
Backbone model of de novo-designed chlorophyll-binding nanocage O32-15
Method: single particle / : Redler RL, Ennist NM, Wang S, Baker D, Ekiert DC, Bhabha G

EMDB-40209: 
Chlorophyll-binding region of de novo-designed nanocage O32-15
Method: single particle / : Redler RL, Ennist NM, Wang S, Baker D, Ekiert DC, Bhabha G

PDB-8glt: 
Backbone model of de novo-designed chlorophyll-binding nanocage O32-15
Method: single particle / : Redler RL, Ennist NM, Wang S, Baker D, Ekiert DC, Bhabha G

EMDB-41810: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env Local Refinement
Method: single particle / : Thakur B, Stalls VD, Acharya P

EMDB-41820: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env
Method: single particle / : Thakur B, Stalls VD, Acharya P

EMDB-41823: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env
Method: single particle / : Thakur B, Stalls VD, Acharya P

EMDB-41838: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to partially open HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env
Method: single particle / : Thakur B, Stalls VD, Acharya P

PDB-8u1d: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env Local Refinement
Method: single particle / : Thakur B, Stalls VD, Acharya P

EMDB-27703: 
Structure of RBD directed antibody DH1047 in complex with SARS-CoV-2 spike: Local refinement of RBD-Fab interace
Method: single particle / : May AJ, Manne K, Acharya P

PDB-8dtk: 
Structure of RBD directed antibody DH1047 in complex with SARS-CoV-2 spike: Local refinement of RBD-Fab interace
Method: single particle / : May AJ, Manne K, Acharya P

EMDB-14460: 
P. berghei kinesin-8B motor domain in AMPPNP state bound to tubulin dimer
Method: single particle / : Liu T, Shilliday F, Cook AD, Moores CA

PDB-7z2b: 
P. berghei kinesin-8B motor domain in AMPPNP state bound to tubulin dimer
Method: single particle / : Liu T, Shilliday F, Cook AD, Moores CA
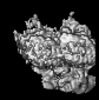
EMDB-14459: 
P. berghei kinesin-8B motor domain in no nucleotide state bound to tubulin dimer
Method: single particle / : Liu T, Shilliday F, Cook AD, Moores CA
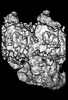
EMDB-14461: 
P. falciparum kinesin-8B motor domain in no nucleotide bound to tubulin dimer
Method: single particle / : Liu T, Shilliday F, Cook AD, Moores CA

PDB-7z2a: 
P. berghei kinesin-8B motor domain in no nucleotide state bound to tubulin dimer
Method: single particle / : Liu T, Shilliday F, Cook AD, Moores CA

PDB-7z2c: 
P. falciparum kinesin-8B motor domain in no nucleotide bound to tubulin dimer
Method: single particle / : Liu T, Shilliday F, Cook AD, Moores CA

EMDB-13149: 
Mammalian orthoreovirus T3SA- in complex with the neural receptor NgR1
Method: single particle / : Strebl M, Sutherland DM, Yu X, Hu L, Wang Z, Prasad BVV, Stehle T, Dermody TS

EMDB-13150: 
Mammalian orthoreovirus T3SA-
Method: single particle / : Strebl M, Sutherland DM, Yu X, Hu L, Wang Z, Prasad BVV, Stehle T, Dermody TS

EMDB-24941: 
Helicobacter Hepaticus CcsBA Open Conformation
Method: single particle / : Mendez DL, Lowder EP

EMDB-24942: 
Helicobacter Hepaticus CcsBA Closed Conformation
Method: single particle / : Mendez DL, Lowder EP

PDB-7s9y: 
Helicobacter Hepaticus CcsBA Open Conformation
Method: single particle / : Mendez DL, Lowder EP, Tillman DE, Sutherland MC, Collier AL, Rau MJ, Fitzpatrick JA, Kranz RG

PDB-7s9z: 
Helicobacter Hepaticus CcsBA Closed Conformation
Method: single particle / : Mendez DL, Lowder EP, Tillman DE, Sutherland MC, Collier AL, Rau MJ, Fitzpatrick JA, Kranz RG

EMDB-24619: 
Negative stain electron microscopy single particle reconstruction of monoclonal antibody DH1024 Fab in complex with HIV-1 Env CH505 SOSIP trimer
Method: single particle / : Edwards RJ, Mansouri K
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
