-Search query
-Search result
Showing all 39 items for (author: smits & c)

EMDB-53270: 
DNA polymerase with inhibitor
Method: single particle / : Lamers MH, Urem M, Smits WK

EMDB-53319: 
DNA polymerase with inhibitor #2
Method: single particle / : Lamers MH, Urem M, Smits WK

EMDB-53320: 
DNA polymerase without DNA or inhibitor
Method: single particle / : Lamers MH, Urem M, Smits WK

PDB-9qrn: 
DNA polymerase without DNA or inhibitor
Method: single particle / : Lamers MH, Urem M, Smits WK

EMDB-50910: 
Structure of MadB, a class I dehydrates from Clostridium maddingley in the apo state
Method: single particle / : Knospe CV, Ortiz J, Reiners J, Kedrov A, Gertzen C, Smits SHJ, Schmitt L

EMDB-50911: 
Structure of MadB, a class I dehydrates from Clostridium maddingley, in complex with its substrate
Method: single particle / : Knospe CV, Ortiz J, Reiners J, Kedrov A, Gerten C, Smits SHJ, Schmitt L

PDB-9g04: 
Structure of MadB, a class I dehydrates from Clostridium maddingley in the apo state
Method: single particle / : Knospe CV, Ortiz J, Reiners J, Kedrov A, Gertzen C, Smits SHJ, Schmitt L

PDB-9g05: 
Structure of MadB, a class I dehydrates from Clostridium maddingley, in complex with its substrate
Method: single particle / : Knospe CV, Ortiz J, Reiners J, Kedrov A, Gerten C, Smits SHJ, Schmitt L

EMDB-14250: 
Structure of the SARS-CoV-2 spike glycoprotein in complex with the 87G7 antibody Fab fragment
Method: single particle / : Hurdiss DL, Drulyte I

EMDB-14271: 
Structure of the SARS-CoV-2 spike glycoprotein in complex with the 87G7 antibody Fab fragment (local refinement)
Method: single particle / : Hurdiss DL, Drulyte I

PDB-7r40: 
Structure of the SARS-CoV-2 spike glycoprotein in complex with the 87G7 antibody Fab fragment
Method: single particle / : Hurdiss DL

EMDB-13142: 
Cryo-EM structure of Pdr5 from Saccharomyces cerevisiae in inward-facing conformation without nucleotides
Method: single particle / : Szewczak-Harris A, Wagner M

EMDB-13143: 
Cryo-EM structure of Pdr5 from Saccharomyces cerevisiae in inward-facing conformation with ADP/ATP
Method: single particle / : Szewczak-Harris A, Wagner M

EMDB-13144: 
Cryo-EM structure of Pdr5 from Saccharomyces cerevisiae in inward-facing conformation with ADP/ATP and rhodamine 6G
Method: single particle / : Szewczak-Harris A, Wagner M

EMDB-13145: 
Cryo-EM structure of Pdr5 from Saccharomyces cerevisiae in outward-facing conformation with ADP-orthovanadate/ATP
Method: single particle / : Szewczak-Harris A, Wagner M

PDB-7p03: 
Cryo-EM structure of Pdr5 from Saccharomyces cerevisiae in inward-facing conformation without nucleotides
Method: single particle / : Szewczak-Harris A, Wagner M, Du D, Schmitt L, Luisi BF

PDB-7p04: 
Cryo-EM structure of Pdr5 from Saccharomyces cerevisiae in inward-facing conformation with ADP/ATP
Method: single particle / : Szewczak-Harris A, Wagner M, Du D, Schmitt L, Luisi BF

PDB-7p05: 
Cryo-EM structure of Pdr5 from Saccharomyces cerevisiae in inward-facing conformation with ADP/ATP and rhodamine 6G
Method: single particle / : Szewczak-Harris A, Wagner M, Du D, Schmitt L, Luisi BF

PDB-7p06: 
Cryo-EM structure of Pdr5 from Saccharomyces cerevisiae in outward-facing conformation with ADP-orthovanadate/ATP
Method: single particle / : Szewczak-Harris A, Wagner M, Du D, Schmitt L, Luisi BF

EMDB-21382: 
Negative stain EM map of an MTA-HDAC-MBD complex
Method: single particle / : Silva APG, Low JKK, Tabar MS, Torrado M, Schmidberger JW, Brillault L, Landsberg MJ, Mackay JP

EMDB-22895: 
Low resolution map of the nucleosome remodelling and deacetylase complex from MEL cells.
Method: single particle / : Jackman MJ, Landsberg MJ, Mackay JP
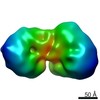
EMDB-22904: 
Low resolution map of the nucleosome deacetylase complex from murine erythroleukemia cells.
Method: single particle / : Jackman MJ, Landsberg MJ, Mackay JP

EMDB-22905: 
Map of the nucleosome deacetylase complex in a twisted conformation
Method: single particle / : Jackman MJ, Landsberg MJ, Mackay JP

EMDB-22906: 
The untwisted conformation of the nucleosome deacetylase complex
Method: single particle / : Jackman MJ, Landsberg MJ, Mackay JP

EMDB-20006: 
Autoinhibited E. coli ATP synthase state 1 - reprocessed
Method: single particle / : Sobti M, Smits C, Wong AS, Ishmukhametov R, Stock D, Sandin S, Stewart AG

EMDB-20007: 
Autoinhibited E. coli ATP synthase state 2 - reprocessed
Method: single particle / : Sobti M, Smits C, Wong AS, Ishmukhametov R, Stock D, Sandin S, Stewart AG

EMDB-20008: 
Autoinhibited E. coli ATP synthase state 3 - reprocessed
Method: single particle / : Sobti M, Smits C, Wong AS, Ishmukhametov R, Stock D, Sandin S, Stewart AG

EMDB-8462: 
Thermus thermophilus V/A-ATPase bound to VH dAbs
Method: single particle / : Davies RB, Smits C, Wong ASW, Stock D, Sandin S, Stewart AG

PDB-5tsj: 
Thermus thermophilus V/A-ATPase bound to VH dAbs
Method: single particle / : Davies RB, Smits C, Wong ASW, Stock D, Sandin S, Stewart AG

EMDB-8359: 
Autoinhibited E. coli ATP synthase state 3
Method: single particle / : Sobti M, Smits C, Wong AS, Ishmukhametov R, Stock D, Sandin S, Stewart AG

PDB-5t4q: 
Autoinhibited E. coli ATP synthase state 3
Method: single particle / : Sobti M, Smits C, Wong ASW, Ishmukhametov R, Stock D, Sandin S, Stewart AG

EMDB-8357: 
Autoinhibited E. coli ATP synthase state 1
Method: single particle / : Sobti M, Smits C, Wong AS, Ishmukhametov R, Stock D, Sandin S, Stewart AG

EMDB-8358: 
Autoinhibited E. coli ATP synthase state 2
Method: single particle / : Sobti M, Smits C, Wong AS, Ishmukhametov R, Stock D, Sandin S, Stewart AG

PDB-5t4o: 
Autoinhibited E. coli ATP synthase state 1
Method: single particle / : Sobti M, Smits C, Wong ASW, Ishmukhametov R, Stock D, Sandin S, Stewart AG

PDB-5t4p: 
Autoinhibited E. coli ATP synthase state 2
Method: single particle / : Sobti M, Smits C, Wong ASW, Ishmukhametov R, Stock D, Sandin S, Stewart AG

EMDB-6291: 
16 Angstrom cryo-EM reconstruction of the alpha, beta, gamma TF55 chaperonin
Method: single particle / : Chaston JJ, Stewart AG, Smits C, Aragao D, Struwe W, Benesch J, Xwong A, Ling M, Ashsan B, Sandin S, Rhodes D, Molugu SK, Bernal RA, Stock D

EMDB-2939: 
Electron tomogram of Pseudomonas moraviensis stanleyae containing selenium nanoparticles
Method: electron tomography / : Ni TW, Staicu L, Nemeth RS, Schwartz C, Crawford D, Seligman JD, Hunter WJ, Pilon-Smits E, Ackerson CJ
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search





 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
